Gene Page: SMYD5
Summary ?
| GeneID | 10322 |
| Symbol | SMYD5 |
| Synonyms | NN8-4AG|RAI15|RRG1|ZMYND23 |
| Description | SMYD family member 5 |
| Reference | HGNC:HGNC:16258|Ensembl:ENSG00000135632|HPRD:18080|Vega:OTTHUMG00000150482 |
| Gene type | protein-coding |
| Map location | 2p13.2 |
| Pascal p-value | 1.564E-4 |
| Sherlock p-value | 0.013 |
| Fetal beta | 0.059 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
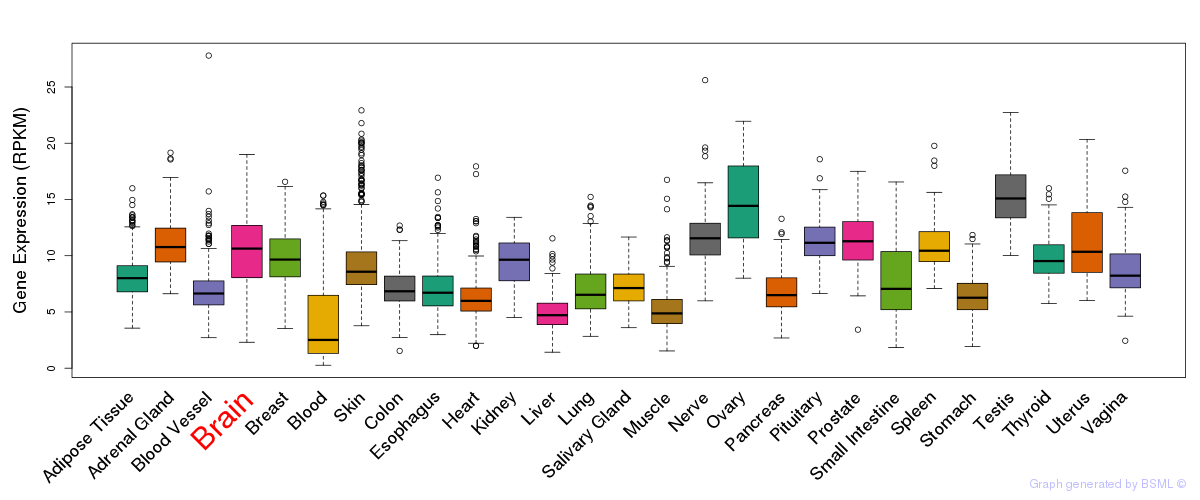
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| WIBG | 0.89 | 0.90 |
| SF4 | 0.88 | 0.88 |
| CYTH2 | 0.88 | 0.88 |
| EXOSC7 | 0.87 | 0.88 |
| MUS81 | 0.87 | 0.91 |
| PRPF31 | 0.87 | 0.88 |
| CNPY3 | 0.87 | 0.88 |
| WDR46 | 0.87 | 0.87 |
| TMED9 | 0.87 | 0.87 |
| CXXC1 | 0.87 | 0.90 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.31 | -0.80 | -0.81 |
| AF347015.27 | -0.80 | -0.83 |
| MT-CO2 | -0.79 | -0.79 |
| MT-CYB | -0.78 | -0.81 |
| AF347015.33 | -0.78 | -0.79 |
| AF347015.8 | -0.77 | -0.80 |
| AF347015.2 | -0.75 | -0.79 |
| AF347015.15 | -0.75 | -0.81 |
| AF347015.26 | -0.72 | -0.77 |
| AF347015.21 | -0.70 | -0.74 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DOANE RESPONSE TO ANDROGEN DN | 241 | 146 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN DN | 770 | 415 | All SZGR 2.0 genes in this pathway |
| OXFORD RALA AND RALB TARGETS UP | 10 | 7 | All SZGR 2.0 genes in this pathway |
| NGUYEN NOTCH1 TARGETS DN | 86 | 67 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS STEM CELL AND PROGENITOR | 681 | 420 | All SZGR 2.0 genes in this pathway |
| MONNIER POSTRADIATION TUMOR ESCAPE UP | 393 | 244 | All SZGR 2.0 genes in this pathway |
| SANSOM APC TARGETS | 212 | 121 | All SZGR 2.0 genes in this pathway |
| SANSOM APC TARGETS REQUIRE MYC | 210 | 123 | All SZGR 2.0 genes in this pathway |
| HELLER HDAC TARGETS DN | 292 | 189 | All SZGR 2.0 genes in this pathway |
| HELLER HDAC TARGETS SILENCED BY METHYLATION DN | 281 | 179 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS DN | 882 | 538 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS DN | 553 | 343 | All SZGR 2.0 genes in this pathway |