Gene Page: SUGT1
Summary ?
| GeneID | 10910 |
| Symbol | SUGT1 |
| Synonyms | SGT1 |
| Description | SGT1 homolog, MIS12 kinetochore complex assembly cochaperone |
| Reference | MIM:604098|HGNC:HGNC:16987| |
| Gene type | protein-coding |
| Map location | 13q14.3 |
| Pascal p-value | 0.003 |
| Sherlock p-value | 0.977 |
| Fetal beta | 0.214 |
| DMG | 1 (# studies) |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 1 |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg26168557 | 13 | 53226688 | SUGT1 | 1.52E-12 | -0.014 | 1.43E-7 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs10507575 | chr13 | 53791403 | SUGT1 | 10910 | 0.13 | cis | ||
| rs8002469 | chr13 | 53936312 | SUGT1 | 10910 | 0.16 | cis | ||
| rs1155107 | chr3 | 26843376 | SUGT1 | 10910 | 0.08 | trans | ||
| rs17029291 | chr3 | 32402138 | SUGT1 | 10910 | 0.02 | trans | ||
| rs17165234 | chr5 | 126403634 | SUGT1 | 10910 | 0.03 | trans | ||
| rs3779536 | chr7 | 127233976 | SUGT1 | 10910 | 0.09 | trans | ||
| rs7894597 | chr10 | 9384581 | SUGT1 | 10910 | 0.06 | trans | ||
| rs7895623 | chr10 | 57180539 | SUGT1 | 10910 | 0.19 | trans | ||
| rs7076439 | chr10 | 93286841 | SUGT1 | 10910 | 0.12 | trans | ||
| rs11226214 | chr11 | 104046824 | SUGT1 | 10910 | 0.15 | trans | ||
| rs1324669 | chr13 | 107872446 | SUGT1 | 10910 | 8.182E-7 | trans | ||
| rs12441507 | chr15 | 61484028 | SUGT1 | 10910 | 0.07 | trans | ||
| rs17145698 | chrX | 40218345 | SUGT1 | 10910 | 1.118E-5 | trans | ||
| rs17328938 | chrX | 99519447 | SUGT1 | 10910 | 0.09 | trans | ||
| rs1335267 | chrX | 99570223 | SUGT1 | 10910 | 0.09 | trans |
Section II. Transcriptome annotation
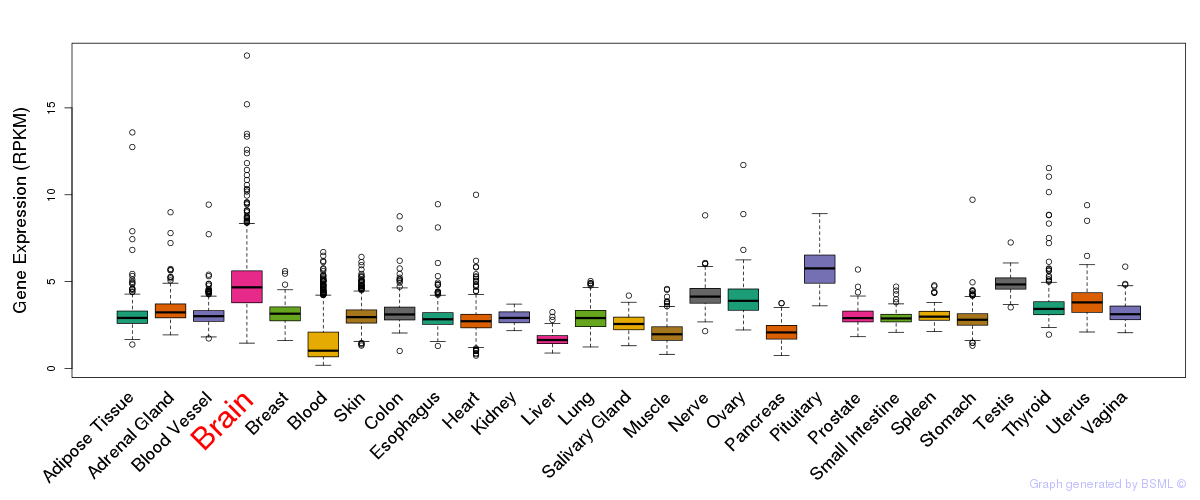
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SLC25A11 | 0.82 | 0.76 |
| MRPL4 | 0.82 | 0.77 |
| C11orf59 | 0.82 | 0.80 |
| MRPL38 | 0.81 | 0.79 |
| WDR45 | 0.81 | 0.81 |
| TMEM222 | 0.80 | 0.81 |
| NIT1 | 0.80 | 0.77 |
| YIPF2 | 0.80 | 0.75 |
| HDHD3 | 0.80 | 0.77 |
| ACY1 | 0.79 | 0.76 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SEC62 | -0.48 | -0.54 |
| EIF5B | -0.47 | -0.52 |
| AC010300.1 | -0.42 | -0.56 |
| AF347015.18 | -0.41 | -0.43 |
| MT-ATP8 | -0.39 | -0.38 |
| NSBP1 | -0.38 | -0.43 |
| AC005921.3 | -0.37 | -0.47 |
| AC016705.1 | -0.36 | -0.34 |
| AC073534.1 | -0.34 | -0.48 |
| AC100783.1 | -0.33 | -0.30 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NOD LIKE RECEPTOR SIGNALING PATHWAY | 62 | 47 | All SZGR 2.0 genes in this pathway |
| NAKAMURA TUMOR ZONE PERIPHERAL VS CENTRAL UP | 285 | 181 | All SZGR 2.0 genes in this pathway |
| WANG LMO4 TARGETS DN | 352 | 225 | All SZGR 2.0 genes in this pathway |
| OSMAN BLADDER CANCER UP | 404 | 246 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED UP | 633 | 376 | All SZGR 2.0 genes in this pathway |
| ROVERSI GLIOMA COPY NUMBER UP | 100 | 75 | All SZGR 2.0 genes in this pathway |
| RASHI RESPONSE TO IONIZING RADIATION 6 | 84 | 54 | All SZGR 2.0 genes in this pathway |
| BENPORATH MYC MAX TARGETS | 775 | 494 | All SZGR 2.0 genes in this pathway |
| ACEVEDO METHYLATED IN LIVER CANCER DN | 940 | 425 | All SZGR 2.0 genes in this pathway |
| DANG BOUND BY MYC | 1103 | 714 | All SZGR 2.0 genes in this pathway |
| YAO TEMPORAL RESPONSE TO PROGESTERONE CLUSTER 9 | 76 | 45 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 3 DN | 918 | 550 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE LATE | 1137 | 655 | All SZGR 2.0 genes in this pathway |