Gene Page: PRDM4
Summary ?
| GeneID | 11108 |
| Symbol | PRDM4 |
| Synonyms | PFM1 |
| Description | PR domain 4 |
| Reference | MIM:605780|HGNC:HGNC:9348|Ensembl:ENSG00000110851|HPRD:10425|Vega:OTTHUMG00000169914 |
| Gene type | protein-coding |
| Map location | 12q23.3 |
| Pascal p-value | 0.877 |
| Sherlock p-value | 0.688 |
| Fetal beta | -0.358 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
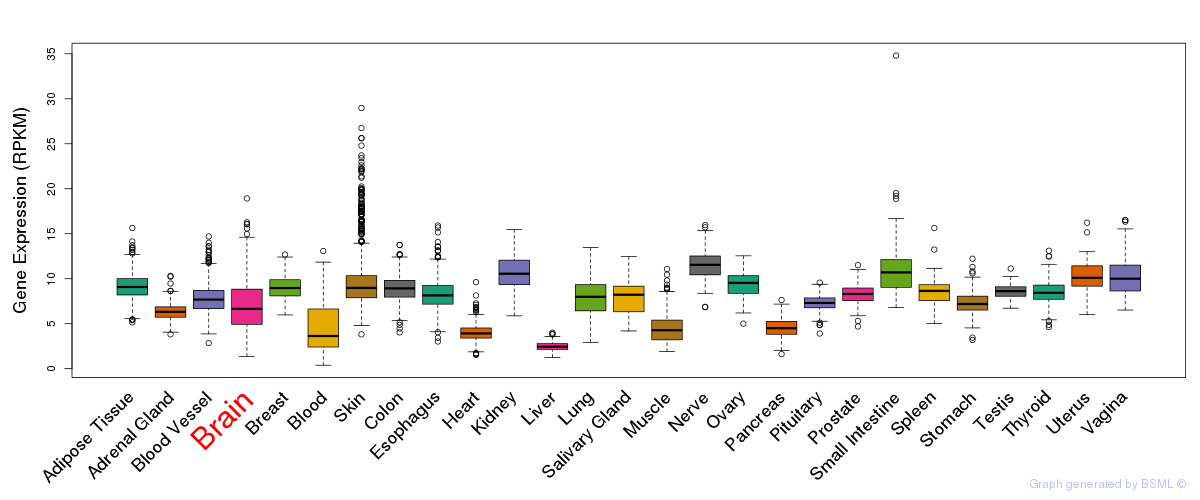
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| EIF4A1 | 0.95 | 0.94 |
| SF3B3 | 0.95 | 0.91 |
| RAF1 | 0.94 | 0.91 |
| SSRP1 | 0.94 | 0.92 |
| COG4 | 0.94 | 0.94 |
| TRAFD1 | 0.94 | 0.89 |
| ZNF16 | 0.94 | 0.92 |
| DEDD | 0.94 | 0.93 |
| MTA2 | 0.94 | 0.88 |
| SMARCB1 | 0.94 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.27 | -0.77 | -0.89 |
| AF347015.31 | -0.76 | -0.88 |
| MT-CO2 | -0.75 | -0.88 |
| HLA-F | -0.75 | -0.79 |
| AF347015.33 | -0.75 | -0.88 |
| MT-CYB | -0.74 | -0.87 |
| C5orf53 | -0.74 | -0.75 |
| AF347015.8 | -0.74 | -0.88 |
| AIFM3 | -0.73 | -0.77 |
| FXYD1 | -0.72 | -0.84 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NEUROTROPHIN SIGNALING PATHWAY | 126 | 103 | All SZGR 2.0 genes in this pathway |
| PID P75 NTR PATHWAY | 69 | 51 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALLING BY NGF | 217 | 167 | All SZGR 2.0 genes in this pathway |
| REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | 81 | 61 | All SZGR 2.0 genes in this pathway |
| GARGALOVIC RESPONSE TO OXIDIZED PHOSPHOLIPIDS MAGENTA UP | 28 | 18 | All SZGR 2.0 genes in this pathway |
| UDAYAKUMAR MED1 TARGETS DN | 240 | 171 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS UP | 811 | 508 | All SZGR 2.0 genes in this pathway |
| GEORGES TARGETS OF MIR192 AND MIR215 | 893 | 528 | All SZGR 2.0 genes in this pathway |
| MORI EMU MYC LYMPHOMA BY ONSET TIME UP | 110 | 69 | All SZGR 2.0 genes in this pathway |
| MASSARWEH TAMOXIFEN RESISTANCE UP | 578 | 341 | All SZGR 2.0 genes in this pathway |
| BONCI TARGETS OF MIR15A AND MIR16 1 | 91 | 75 | All SZGR 2.0 genes in this pathway |
| WEST ADRENOCORTICAL TUMOR UP | 294 | 199 | All SZGR 2.0 genes in this pathway |
| PODAR RESPONSE TO ADAPHOSTIN UP | 147 | 98 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP D | 280 | 158 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |