Gene Page: CHRNB4
Summary ?
| GeneID | 1143 |
| Symbol | CHRNB4 |
| Synonyms | - |
| Description | cholinergic receptor nicotinic beta 4 subunit |
| Reference | MIM:118509|HGNC:HGNC:1964|Ensembl:ENSG00000117971|HPRD:00337|Vega:OTTHUMG00000143860 |
| Gene type | protein-coding |
| Map location | 15q24 |
| Pascal p-value | 1.2E-9 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
| GO_Annotation | Mapping neuro-related keywords to Gene Ontology annotations | Hits with neuro-related keywords: 7 |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
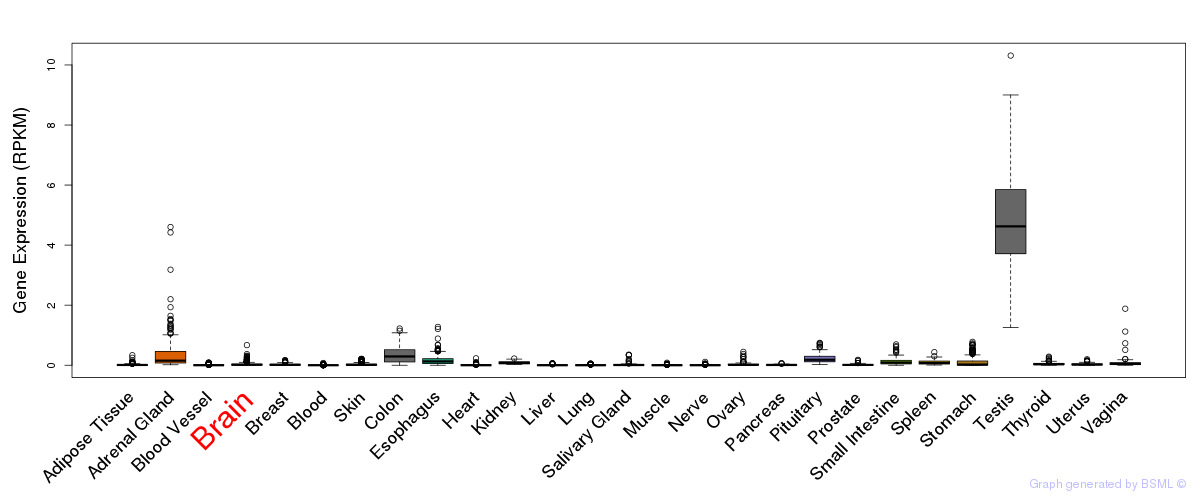
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PC | 0.79 | 0.81 |
| NR1H2 | 0.79 | 0.80 |
| THOP1 | 0.72 | 0.72 |
| NDUFAF3 | 0.70 | 0.73 |
| PNPLA2 | 0.70 | 0.65 |
| NAT8L | 0.70 | 0.72 |
| JOSD2 | 0.70 | 0.76 |
| GHDC | 0.70 | 0.67 |
| PDK2 | 0.70 | 0.74 |
| SPHK2 | 0.68 | 0.71 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SCN9A | -0.42 | -0.30 |
| C17orf57 | -0.41 | -0.34 |
| AC109829.1 | -0.40 | -0.36 |
| CCDC148 | -0.39 | -0.40 |
| AC073995.1 | -0.39 | -0.37 |
| GNRHR | -0.38 | -0.32 |
| AC087071.1 | -0.38 | -0.30 |
| NOX1 | -0.37 | -0.33 |
| ANKRD36B | -0.37 | -0.35 |
| CCDC122 | -0.36 | -0.29 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0015464 | acetylcholine receptor activity | IDA | Neurotransmitter (GO term level: 6) | 8906617 |
| GO:0004889 | nicotinic acetylcholine-activated cation-selective channel activity | TAS | 8906617 |11742001 | |
| GO:0005216 | ion channel activity | IEA | - | |
| GO:0005230 | extracellular ligand-gated ion channel activity | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0060084 | synaptic transmission involved in micturition | IMP | neuron, Synap (GO term level: 8) | 11450844 |
| GO:0046928 | regulation of neurotransmitter secretion | NAS | Synap, Neurotransmitter (GO term level: 9) | 11742001 |
| GO:0007271 | synaptic transmission, cholinergic | TAS | neuron, Cholinergic, Synap, Neurotransmitter (GO term level: 7) | 1330682 |
| GO:0001508 | regulation of action potential | IEA | - | |
| GO:0007165 | signal transduction | IDA | 8906617 | |
| GO:0007626 | locomotory behavior | IEA | - | |
| GO:0006811 | ion transport | TAS | 8906617 | |
| GO:0006940 | regulation of smooth muscle contraction | IEA | - | |
| GO:0035095 | behavioral response to nicotine | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0005892 | nicotinic acetylcholine-gated receptor-channel complex | IDA | Neurotransmitter (GO term level: 9) | 8906617 |
| GO:0005892 | nicotinic acetylcholine-gated receptor-channel complex | TAS | Neurotransmitter (GO term level: 9) | 11742001 |
| GO:0045211 | postsynaptic membrane | IEA | Synap, Neurotransmitter (GO term level: 5) | - |
| GO:0045202 | synapse | IEA | neuron, Synap, Neurotransmitter, Glial (GO term level: 2) | - |
| GO:0005886 | plasma membrane | IEA | - | |
| GO:0030054 | cell junction | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NEUROACTIVE LIGAND RECEPTOR INTERACTION | 272 | 195 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | 186 | 155 | All SZGR 2.0 genes in this pathway |
| REACTOME NEURONAL SYSTEM | 279 | 221 | All SZGR 2.0 genes in this pathway |
| REACTOME NEUROTRANSMITTER RECEPTOR BINDING AND DOWNSTREAM TRANSMISSION IN THE POSTSYNAPTIC CELL | 137 | 110 | All SZGR 2.0 genes in this pathway |
| REACTOME ACETYLCHOLINE BINDING AND DOWNSTREAM EVENTS | 16 | 13 | All SZGR 2.0 genes in this pathway |
| REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | 13 | 12 | All SZGR 2.0 genes in this pathway |
| REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | 12 | 10 | All SZGR 2.0 genes in this pathway |
| MORI PLASMA CELL UP | 51 | 29 | All SZGR 2.0 genes in this pathway |
| SANSOM APC TARGETS UP | 126 | 84 | All SZGR 2.0 genes in this pathway |
| ASTIER INTEGRIN SIGNALING | 59 | 44 | All SZGR 2.0 genes in this pathway |
| DAZARD RESPONSE TO UV NHEK UP | 244 | 151 | All SZGR 2.0 genes in this pathway |
| DAZARD UV RESPONSE CLUSTER G1 | 67 | 41 | All SZGR 2.0 genes in this pathway |
| LEIN PONS MARKERS | 89 | 59 | All SZGR 2.0 genes in this pathway |
| LEIN MEDULLA MARKERS | 81 | 48 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MCV6 ICP WITH H3K27ME3 | 74 | 46 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF ICP WITH H3K27ME3 | 206 | 108 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN ES ICP WITH H3K4ME3 | 718 | 401 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN NPC ICP WITH H3K4ME3 | 445 | 257 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |