Gene Page: CSMD2
Summary ?
| GeneID | 114784 |
| Symbol | CSMD2 |
| Synonyms | dJ1007G16.1|dJ1007G16.2|dJ947L8.1 |
| Description | CUB and Sushi multiple domains 2 |
| Reference | MIM:608398|HGNC:HGNC:19290|Ensembl:ENSG00000121904|HPRD:13094|Vega:OTTHUMG00000011135 |
| Gene type | protein-coding |
| Map location | 1p34.3 |
| Pascal p-value | 0.115 |
| Sherlock p-value | 0.671 |
| Fetal beta | 0.896 |
| DMG | 1 (# studies) |
| eGene | Cerebellar Hemisphere Myers' cis & trans Meta |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 2 |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| CSMD2 | chr1 | 34071483 | T | C | NM_052896 | p.2110D>G | missense | Schizophrenia | DNM:Fromer_2014 |
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg09270225 | 1 | 34632658 | C1orf94;CSMD2 | 2.98E-4 | -0.511 | 0.039 | DMG:Wockner_2014 |
| cg02187889 | 1 | 34629366 | CSMD2 | 8.29E-8 | -0.009 | 1.93E-5 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs6738420 | chr2 | 159785939 | CSMD2 | 114784 | 0.2 | trans | ||
| rs1522698 | chr2 | 159788917 | CSMD2 | 114784 | 0.16 | trans | ||
| rs4759017 | chr12 | 56691599 | CSMD2 | 114784 | 0.16 | trans | ||
| rs6573406 | chr14 | 62397444 | CSMD2 | 114784 | 0.04 | trans |
Section II. Transcriptome annotation
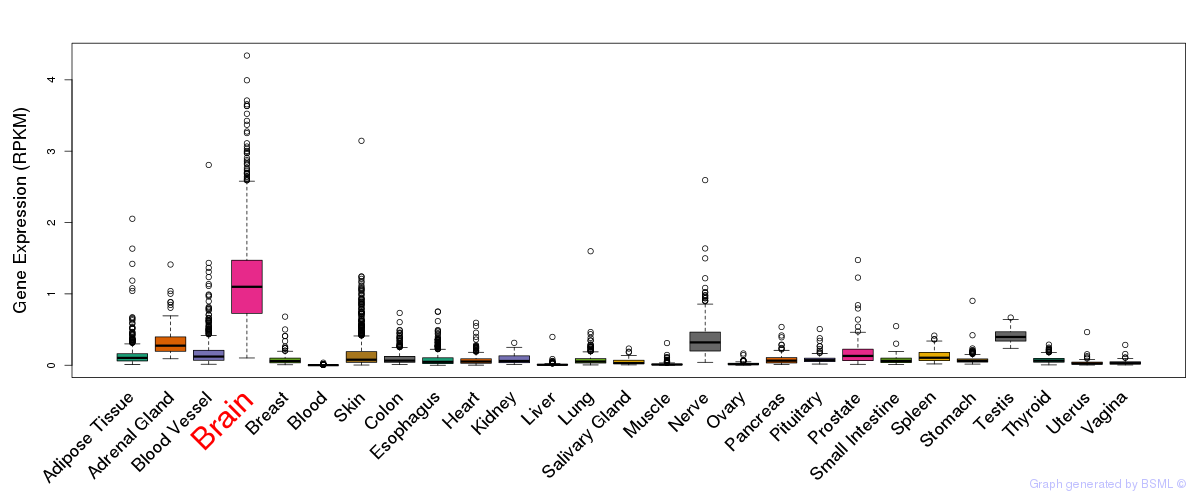
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE FIMA DN | 284 | 156 | All SZGR 2.0 genes in this pathway |
| BENPORATH ES WITH H3K27ME3 | 1118 | 744 | All SZGR 2.0 genes in this pathway |