Gene Page: SLC26A8
Summary ?
| GeneID | 116369 |
| Symbol | SLC26A8 |
| Synonyms | SPGF3|TAT1 |
| Description | solute carrier family 26 member 8 |
| Reference | MIM:608480|HGNC:HGNC:14468|Ensembl:ENSG00000112053|HPRD:12241|Vega:OTTHUMG00000014586 |
| Gene type | protein-coding |
| Map location | 6p21 |
| Pascal p-value | 0.48 |
| Sherlock p-value | 0.206 |
| Fetal beta | -0.493 |
| eGene | Cerebellar Hemisphere Cerebellum Myers' cis & trans Meta |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| DNM:Xu_2012 | Whole Exome Sequencing analysis | De novo mutations of 4 genes were identified by exome sequencing of 795 samples in this study | |
| GSMA_I | Genome scan meta-analysis | Psr: 0.033 | |
| GSMA_IIE | Genome scan meta-analysis (European-ancestry samples) | Psr: 0.04433 |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| SLC26A8 | chr6 | 35927251 | C | T | NM_001193476 NM_052961 NM_138718 | p.617E>K p.617E>K p.512E>K | missense missense missense | Schizophrenia | DNM:Xu_2012 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7370564 | 0 | SLC26A8 | 116369 | 0.18 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
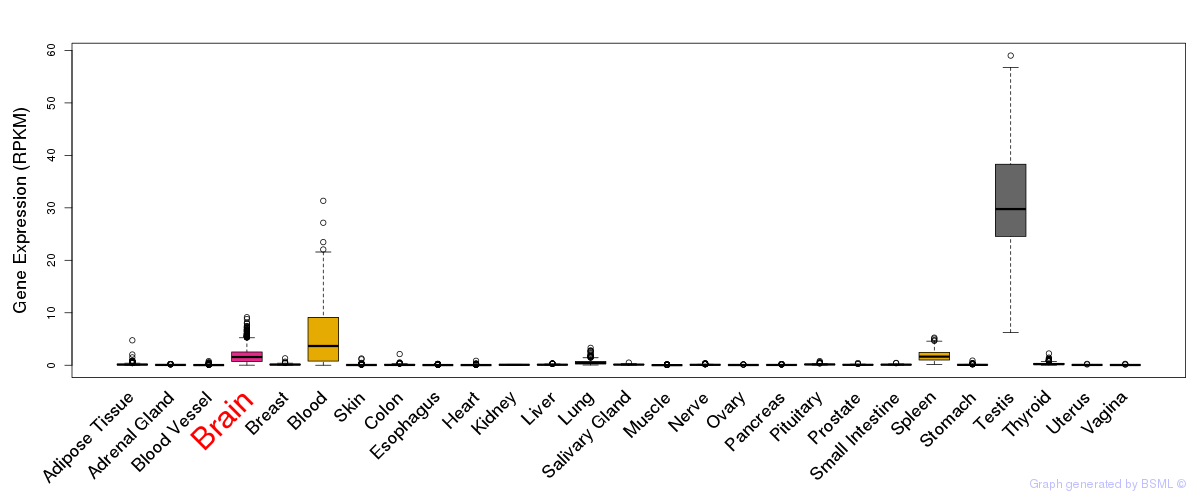
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005254 | chloride channel activity | IDA | 1183472 | |
| GO:0005215 | transporter activity | IEA | - | |
| GO:0015380 | anion exchanger activity | IEA | - | |
| GO:0015116 | sulfate transmembrane transporter activity | IDA | 1183472 | |
| GO:0019531 | oxalate transmembrane transporter activity | IDA | 1183472 | |
| GO:0047485 | protein N-terminus binding | IPI | 11278976 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0007283 | spermatogenesis | IEA | - | |
| GO:0007126 | meiosis | IEA | - | |
| GO:0008272 | sulfate transport | IDA | 1183472 | |
| GO:0006821 | chloride transport | IDA | 1183472 | |
| GO:0006811 | ion transport | IEA | - | |
| GO:0007275 | multicellular organismal development | IEA | - | |
| GO:0030154 | cell differentiation | IEA | - | |
| GO:0019532 | oxalate transport | IDA | 1183472 | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0016021 | integral to membrane | IEA | - | |
| GO:0005886 | plasma membrane | IDA | 11278976 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER WITH H3K9ME3 DN | 120 | 71 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3 UNMETHYLATED | 536 | 296 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3 UNMETHYLATED | 228 | 119 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |