Gene Page: KCNH8
Summary ?
| GeneID | 131096 |
| Symbol | KCNH8 |
| Synonyms | ELK|ELK1|Kv12.1|elk3 |
| Description | potassium voltage-gated channel subfamily H member 8 |
| Reference | MIM:608260|HGNC:HGNC:18864|Ensembl:ENSG00000183960|HPRD:10006|Vega:OTTHUMG00000129891 |
| Gene type | protein-coding |
| Map location | 3p24.3 |
| Pascal p-value | 0.705 |
| Sherlock p-value | 0.945 |
| Fetal beta | 0.109 |
| DMG | 2 (# studies) |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 3 |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 3 |
| GSMA_I | Genome scan meta-analysis | Psr: 0.006 | |
| Expression | Meta-analysis of gene expression | P value: 1.804 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg18500192 | 3 | 19189930 | KCNH8 | 1.169E-4 | -0.329 | 0.029 | DMG:Wockner_2014 |
| cg24997944 | 3 | 19189709 | KCNH8 | 8.94E-8 | -0.008 | 2.05E-5 | DMG:Jaffe_2016 |
| cg07657236 | 3 | 19190202 | KCNH8 | 1.04E-7 | -0.016 | 2.29E-5 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs651185 | chr1 | 233871029 | KCNH8 | 131096 | 0.02 | trans | ||
| snp_a-2241490 | 0 | KCNH8 | 131096 | 0.12 | trans | |||
| rs2322319 | chr3 | 3637199 | KCNH8 | 131096 | 0.15 | trans | ||
| rs17256926 | chr5 | 9399927 | KCNH8 | 131096 | 0.1 | trans | ||
| rs5003961 | chr7 | 78072443 | KCNH8 | 131096 | 0.01 | trans | ||
| rs4515471 | chr7 | 78072585 | KCNH8 | 131096 | 0.01 | trans | ||
| rs10259327 | chr7 | 78074924 | KCNH8 | 131096 | 0.01 | trans | ||
| rs16924550 | chr12 | 22174951 | KCNH8 | 131096 | 0.16 | trans | ||
| rs17137051 | chr16 | 4521748 | KCNH8 | 131096 | 0.14 | trans | ||
| rs184583 | chr19 | 30371589 | KCNH8 | 131096 | 0.15 | trans |
Section II. Transcriptome annotation
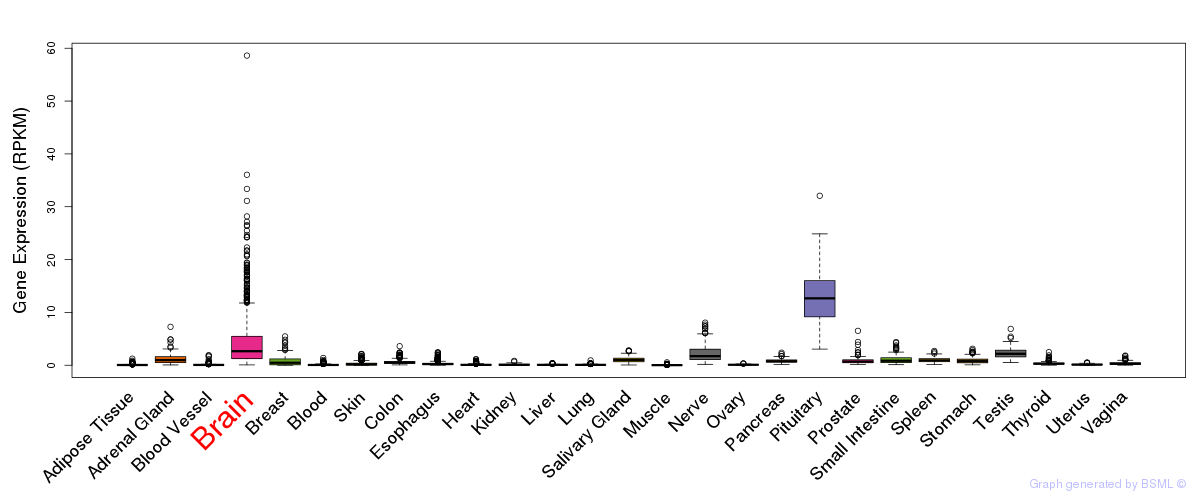
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0000155 | two-component sensor activity | IEA | - | |
| GO:0005244 | voltage-gated ion channel activity | IEA | - | |
| GO:0005249 | voltage-gated potassium channel activity | IEA | - | |
| GO:0030955 | potassium ion binding | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0000160 | two-component signal transduction system (phosphorelay) | IEA | - | |
| GO:0006355 | regulation of transcription, DNA-dependent | IEA | - | |
| GO:0006811 | ion transport | IEA | - | |
| GO:0006813 | potassium ion transport | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0016020 | membrane | IEA | - | |
| GO:0016021 | integral to membrane | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME NEURONAL SYSTEM | 279 | 221 | All SZGR 2.0 genes in this pathway |
| REACTOME VOLTAGE GATED POTASSIUM CHANNELS | 43 | 32 | All SZGR 2.0 genes in this pathway |
| REACTOME POTASSIUM CHANNELS | 98 | 68 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| MEISSNER BRAIN HCP WITH H3K4ME3 AND H3K27ME3 | 1069 | 729 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3K4ME2 AND H3K27ME3 | 349 | 234 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3K27ME3 | 590 | 403 | All SZGR 2.0 genes in this pathway |
| HOELZEL NF1 TARGETS UP | 139 | 93 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |
Section VI. microRNA annotation
| miRNA family | Target position | miRNA ID | miRNA seq | ||
|---|---|---|---|---|---|
| UTR start | UTR end | Match method | |||
| miR-143 | 475 | 481 | m8 | hsa-miR-143brain | UGAGAUGAAGCACUGUAGCUCA |
| miR-193 | 509 | 515 | m8 | hsa-miR-193a | AACUGGCCUACAAAGUCCCAG |
| hsa-miR-193b | AACUGGCCCUCAAAGUCCCGCUUU | ||||
| miR-219 | 485 | 492 | 1A,m8 | hsa-miR-219brain | UGAUUGUCCAAACGCAAUUCU |
| miR-221/222 | 159 | 165 | 1A | hsa-miR-221brain | AGCUACAUUGUCUGCUGGGUUUC |
| hsa-miR-222brain | AGCUACAUCUGGCUACUGGGUCUC | ||||
| miR-485-3p | 601 | 607 | 1A | hsa-miR-485-3p | GUCAUACACGGCUCUCCUCUCU |
| miR-496 | 541 | 548 | 1A,m8 | hsa-miR-496 | AUUACAUGGCCAAUCUC |
- SZ: miRNAs which differentially expressed in brain cortex of schizophrenia patients comparing with control samples using microarray. Click here to see the list of SZ related miRNAs.
- Brain: miRNAs which are expressed in brain based on miRNA microarray expression studies. Click here to see the list of brain related miRNAs.