Gene Page: CR1L
Summary ?
| GeneID | 1379 |
| Symbol | CR1L |
| Synonyms | - |
| Description | complement component 3b/4b receptor 1-like |
| Reference | MIM:605886|HGNC:HGNC:2335|Ensembl:ENSG00000197721|Vega:OTTHUMG00000036354 |
| Gene type | protein-coding |
| Map location | 1q32.1 |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| CR1L | chr1 | 207891009 | C | G | NM_175710 | p.539P>A | missense | Schizophrenia | DNM:Fromer_2014 |
Section II. Transcriptome annotation
General gene expression (GTEx)
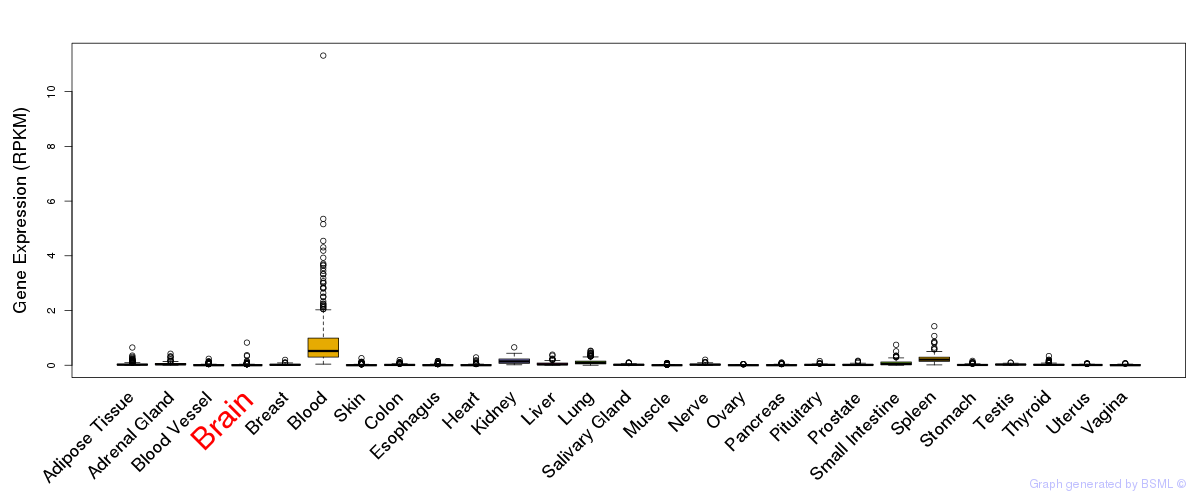
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZNF384 | 0.96 | 0.96 |
| GTF3C2 | 0.95 | 0.95 |
| SMPD4 | 0.95 | 0.96 |
| MOGS | 0.95 | 0.96 |
| NXF1 | 0.94 | 0.93 |
| PPIL2 | 0.94 | 0.95 |
| PCIF1 | 0.94 | 0.95 |
| SF1 | 0.94 | 0.93 |
| YY1AP1 | 0.94 | 0.94 |
| CNOT3 | 0.94 | 0.94 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.31 | -0.79 | -0.89 |
| AF347015.27 | -0.77 | -0.88 |
| MT-CO2 | -0.77 | -0.88 |
| C5orf53 | -0.75 | -0.78 |
| MT-CYB | -0.74 | -0.86 |
| AF347015.33 | -0.74 | -0.84 |
| AF347015.8 | -0.74 | -0.87 |
| AF347015.21 | -0.71 | -0.91 |
| S100B | -0.71 | -0.78 |
| AF347015.15 | -0.70 | -0.83 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHENG BOUND BY FOXP3 | 491 | 310 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 UNSTIMULATED | 1229 | 713 | All SZGR 2.0 genes in this pathway |
| IWANAGA CARCINOGENESIS BY KRAS PTEN UP | 181 | 112 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER WITH H3K27ME3 UP | 295 | 149 | All SZGR 2.0 genes in this pathway |
| COLINA TARGETS OF 4EBP1 AND 4EBP2 | 356 | 214 | All SZGR 2.0 genes in this pathway |
| RAO BOUND BY SALL4 ISOFORM B | 517 | 302 | All SZGR 2.0 genes in this pathway |