Gene Page: CRP
Summary ?
| GeneID | 1401 |
| Symbol | CRP |
| Synonyms | PTX1 |
| Description | C-reactive protein, pentraxin-related |
| Reference | MIM:123260|HGNC:HGNC:2367|Ensembl:ENSG00000132693|HPRD:00422|Vega:OTTHUMG00000035344 |
| Gene type | protein-coding |
| Map location | 1q23.2 |
| Fetal beta | 0.609 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| GSMA_I | Genome scan meta-analysis | Psr: 0.0235 | |
| GSMA_IIA | Genome scan meta-analysis (All samples) | Psr: 0.00814 | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias,schizotypal | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
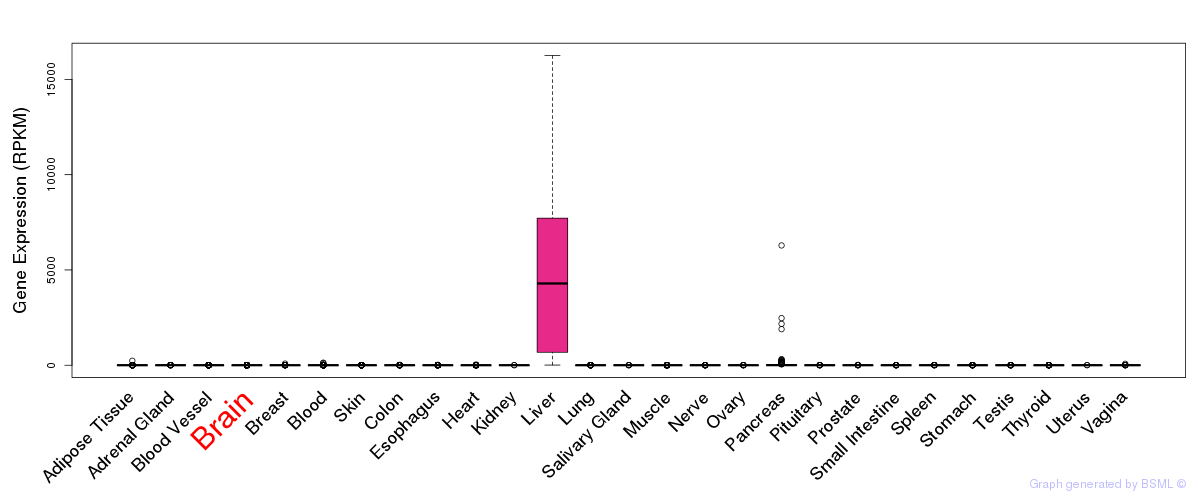
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AP000907.5 | 0.97 | 0.99 |
| DBNDD2 | 0.91 | 0.93 |
| CMTM5 | 0.90 | 0.94 |
| MAL | 0.90 | 0.93 |
| RTKN | 0.89 | 0.93 |
| APOD | 0.89 | 0.94 |
| SEPT4 | 0.88 | 0.90 |
| RNASE1 | 0.88 | 0.92 |
| PLA2G16 | 0.88 | 0.93 |
| NINJ2 | 0.87 | 0.92 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| NKIRAS2 | -0.65 | -0.76 |
| CRMP1 | -0.63 | -0.74 |
| FGD1 | -0.61 | -0.73 |
| PCDHB18 | -0.61 | -0.76 |
| PURG | -0.61 | -0.72 |
| KIAA1949 | -0.61 | -0.74 |
| HMGB3 | -0.61 | -0.78 |
| FARP1 | -0.61 | -0.72 |
| ZNF599 | -0.61 | -0.74 |
| ZNF821 | -0.61 | -0.73 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005509 | calcium ion binding | IEA | - | |
| GO:0005515 | protein binding | IPI | 17785206 | |
| GO:0033265 | choline binding | TAS | 2477488 | |
| GO:0051637 | Gram-positive bacterial cell surface binding | TAS | 2477488 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0006954 | inflammatory response | TAS | 10408362 | |
| GO:0006953 | acute-phase response | IEA | - | |
| GO:0008228 | opsonization | TAS | 2477488 | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0005576 | extracellular region | NAS | 14718574 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID IL6 7 PATHWAY | 47 | 40 | All SZGR 2.0 genes in this pathway |
| REACTOME INNATE IMMUNE SYSTEM | 279 | 178 | All SZGR 2.0 genes in this pathway |
| REACTOME IMMUNE SYSTEM | 933 | 616 | All SZGR 2.0 genes in this pathway |
| REACTOME COMPLEMENT CASCADE | 32 | 22 | All SZGR 2.0 genes in this pathway |
| REACTOME CREATION OF C4 AND C2 ACTIVATORS | 10 | 5 | All SZGR 2.0 genes in this pathway |
| REACTOME INITIAL TRIGGERING OF COMPLEMENT | 16 | 10 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 1 UP | 276 | 165 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 2 UP | 139 | 83 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 SIGNATURE 3 UP | 341 | 197 | All SZGR 2.0 genes in this pathway |
| HSIAO LIVER SPECIFIC GENES | 244 | 154 | All SZGR 2.0 genes in this pathway |
| SU LIVER | 55 | 32 | All SZGR 2.0 genes in this pathway |
| TUOMISTO TUMOR SUPPRESSION BY COL13A1 DN | 17 | 11 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| TAVAZOIE METASTASIS | 108 | 68 | All SZGR 2.0 genes in this pathway |
| CHIANG LIVER CANCER SUBCLASS CTNNB1 DN | 170 | 105 | All SZGR 2.0 genes in this pathway |
| CAIRO HEPATOBLASTOMA DN | 267 | 160 | All SZGR 2.0 genes in this pathway |
| CAIRO LIVER DEVELOPMENT DN | 222 | 141 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS UP | 504 | 321 | All SZGR 2.0 genes in this pathway |
| MAEKAWA ATF2 TARGETS | 24 | 19 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS DN | 553 | 343 | All SZGR 2.0 genes in this pathway |
| SERVITJA LIVER HNF1A TARGETS DN | 157 | 105 | All SZGR 2.0 genes in this pathway |