| KEGG FATTY ACID METABOLISM
| 42 | 29 | All SZGR 2.0 genes in this pathway |
| KEGG VALINE LEUCINE AND ISOLEUCINE DEGRADATION
| 44 | 26 | All SZGR 2.0 genes in this pathway |
| KEGG LYSINE DEGRADATION
| 44 | 29 | All SZGR 2.0 genes in this pathway |
| KEGG TRYPTOPHAN METABOLISM
| 40 | 33 | All SZGR 2.0 genes in this pathway |
| KEGG BETA ALANINE METABOLISM
| 22 | 16 | All SZGR 2.0 genes in this pathway |
| KEGG PROPANOATE METABOLISM
| 33 | 22 | All SZGR 2.0 genes in this pathway |
| KEGG BUTANOATE METABOLISM
| 34 | 20 | All SZGR 2.0 genes in this pathway |
| KEGG LIMONENE AND PINENE DEGRADATION
| 10 | 8 | All SZGR 2.0 genes in this pathway |
| REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION
| 14 | 10 | All SZGR 2.0 genes in this pathway |
| REACTOME METABOLISM OF LIPIDS AND LIPOPROTEINS
| 478 | 302 | All SZGR 2.0 genes in this pathway |
| REACTOME FATTY ACID TRIACYLGLYCEROL AND KETONE BODY METABOLISM
| 168 | 115 | All SZGR 2.0 genes in this pathway |
| DAVICIONI MOLECULAR ARMS VS ERMS UP
| 332 | 228 | All SZGR 2.0 genes in this pathway |
| DIAZ CHRONIC MEYLOGENOUS LEUKEMIA UP
| 1382 | 904 | All SZGR 2.0 genes in this pathway |
| TONKS TARGETS OF RUNX1 RUNX1T1 FUSION MONOCYTE UP
| 204 | 140 | All SZGR 2.0 genes in this pathway |
| TIEN INTESTINE PROBIOTICS 24HR UP
| 557 | 331 | All SZGR 2.0 genes in this pathway |
| ENK UV RESPONSE KERATINOCYTE UP
| 530 | 342 | All SZGR 2.0 genes in this pathway |
| LASTOWSKA NEUROBLASTOMA COPY NUMBER DN
| 800 | 473 | All SZGR 2.0 genes in this pathway |
| HUMMERICH SKIN CANCER PROGRESSION DN
| 100 | 64 | All SZGR 2.0 genes in this pathway |
| DAIRKEE TERT TARGETS UP
| 380 | 213 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS
| 882 | 572 | All SZGR 2.0 genes in this pathway |
| FLECHNER BIOPSY KIDNEY TRANSPLANT REJECTED VS OK DN
| 546 | 351 | All SZGR 2.0 genes in this pathway |
| NADLER OBESITY DN
| 48 | 34 | All SZGR 2.0 genes in this pathway |
| GERY CEBP TARGETS
| 126 | 90 | All SZGR 2.0 genes in this pathway |
| PAL PRMT5 TARGETS UP
| 203 | 135 | All SZGR 2.0 genes in this pathway |
| BURTON ADIPOGENESIS 5
| 126 | 78 | All SZGR 2.0 genes in this pathway |
| DAZARD RESPONSE TO UV NHEK UP
| 244 | 151 | All SZGR 2.0 genes in this pathway |
| TAKAO RESPONSE TO UVB RADIATION DN
| 98 | 67 | All SZGR 2.0 genes in this pathway |
| KAAB HEART ATRIUM VS VENTRICLE DN
| 261 | 183 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 3
| 720 | 440 | All SZGR 2.0 genes in this pathway |
| WONG MITOCHONDRIA GENE MODULE
| 217 | 122 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN
| 540 | 340 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER TUMOR VS NORMAL ADJACENT TISSUE DN
| 274 | 165 | All SZGR 2.0 genes in this pathway |
| MOOTHA HUMAN MITODB 6 2002
| 429 | 260 | All SZGR 2.0 genes in this pathway |
| LEE LIVER CANCER SURVIVAL UP
| 185 | 112 | All SZGR 2.0 genes in this pathway |
| MOOTHA MITOCHONDRIA
| 447 | 277 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP
| 1305 | 895 | All SZGR 2.0 genes in this pathway |
| HSIAO HOUSEKEEPING GENES
| 389 | 245 | All SZGR 2.0 genes in this pathway |
| BOYAULT LIVER CANCER SUBCLASS G3 DN
| 51 | 32 | All SZGR 2.0 genes in this pathway |
| WOO LIVER CANCER RECURRENCE DN
| 80 | 54 | All SZGR 2.0 genes in this pathway |
| NGO MALIGNANT GLIOMA 1P LOH
| 17 | 13 | All SZGR 2.0 genes in this pathway |
| HOSHIDA LIVER CANCER SUBCLASS S3
| 266 | 180 | All SZGR 2.0 genes in this pathway |
| WONG EMBRYONIC STEM CELL CORE
| 335 | 193 | All SZGR 2.0 genes in this pathway |
| MOOTHA FFA OXYDATION
| 22 | 13 | All SZGR 2.0 genes in this pathway |
| KIM ALL DISORDERS OLIGODENDROCYTE NUMBER CORR UP
| 756 | 494 | All SZGR 2.0 genes in this pathway |
| KIM BIPOLAR DISORDER OLIGODENDROCYTE DENSITY CORR UP
| 682 | 440 | All SZGR 2.0 genes in this pathway |
| WAKABAYASHI ADIPOGENESIS PPARG RXRA BOUND 8D
| 882 | 506 | All SZGR 2.0 genes in this pathway |
| FOSTER KDM1A TARGETS DN
| 211 | 119 | All SZGR 2.0 genes in this pathway |
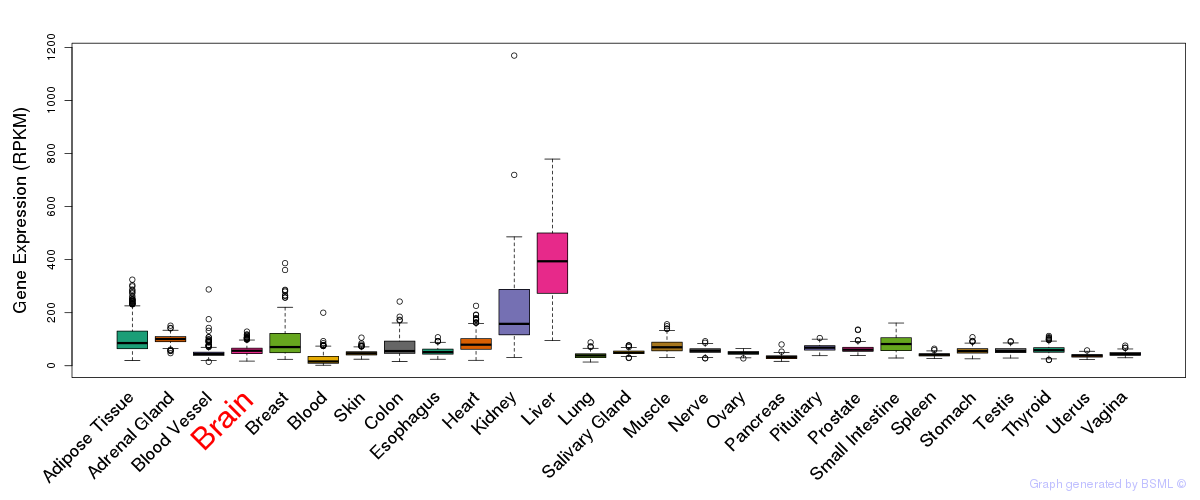
 eQTL annotation
eQTL annotation