Gene Page: ZNF365
Summary ?
| GeneID | 22891 |
| Symbol | ZNF365 |
| Synonyms | Su48|UAN|ZNF365D |
| Description | zinc finger protein 365 |
| Reference | MIM:607818|HGNC:HGNC:18194|Ensembl:ENSG00000138311|HPRD:06380|Vega:OTTHUMG00000018302 |
| Gene type | protein-coding |
| Map location | 10q21.2 |
| Pascal p-value | 3.719E-6 |
| Fetal beta | -0.427 |
| DMG | 1 (# studies) |
| eGene | Putamen basal ganglia Myers' cis & trans |
| Support | Darnell FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg01137532 | 10 | 64133888 | ZNF365 | 5.93E-9 | -0.008 | 3.18E-6 | DMG:Jaffe_2016 |
| cg13482925 | 10 | 64028750 | ZNF365 | 9.68E-8 | -0.013 | 2.16E-5 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs16869851 | chr4 | 20690056 | ZNF365 | 22891 | 0.17 | trans | ||
| rs4636539 | 10 | 64182727 | ZNF365 | ENSG00000138311.11 | 3.10461E-6 | 0.04 | 48776 | gtex_brain_putamen_basal |
| rs2138559 | 10 | 64188532 | ZNF365 | ENSG00000138311.11 | 3.10489E-6 | 0.04 | 54581 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
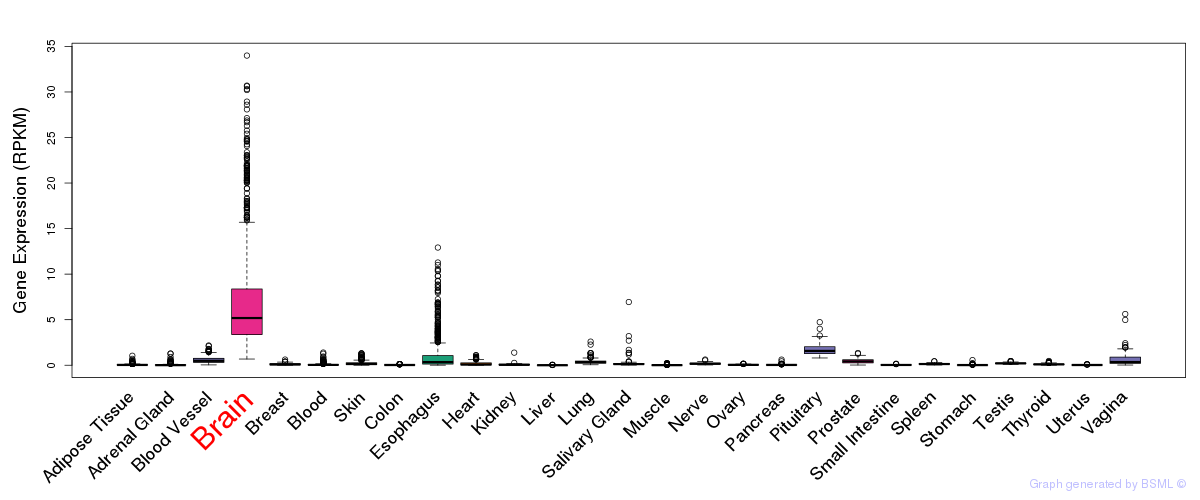
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PRICKLE1 | 0.80 | 0.83 |
| NRCAM | 0.80 | 0.84 |
| SH3GL2 | 0.79 | 0.84 |
| HYOU1 | 0.78 | 0.86 |
| CYFIP2 | 0.78 | 0.85 |
| CDH18 | 0.77 | 0.81 |
| AMPH | 0.76 | 0.81 |
| DIRAS1 | 0.76 | 0.82 |
| PFKM | 0.76 | 0.78 |
| NDUFS1 | 0.76 | 0.82 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| RAB34 | -0.50 | -0.58 |
| RHOC | -0.48 | -0.57 |
| EFEMP2 | -0.46 | -0.50 |
| PSME2 | -0.46 | -0.48 |
| S100A16 | -0.46 | -0.50 |
| C1orf61 | -0.46 | -0.58 |
| C1orf54 | -0.44 | -0.52 |
| AF347015.21 | -0.44 | -0.47 |
| DBI | -0.43 | -0.54 |
| IGFBP7 | -0.43 | -0.49 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| NAGASHIMA NRG1 SIGNALING UP | 176 | 123 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS UP | 214 | 155 | All SZGR 2.0 genes in this pathway |
| MARTORIATI MDM4 TARGETS NEUROEPITHELIUM UP | 176 | 111 | All SZGR 2.0 genes in this pathway |
| MARTORIATI MDM4 TARGETS FETAL LIVER UP | 227 | 137 | All SZGR 2.0 genes in this pathway |
| NUYTTEN EZH2 TARGETS DN | 1024 | 594 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| BENPORATH EED TARGETS | 1062 | 725 | All SZGR 2.0 genes in this pathway |
| ONDER CDH1 TARGETS 1 DN | 169 | 102 | All SZGR 2.0 genes in this pathway |
| KAAB HEART ATRIUM VS VENTRICLE UP | 249 | 170 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 48HR UP | 180 | 125 | All SZGR 2.0 genes in this pathway |
| KIM ALL DISORDERS OLIGODENDROCYTE NUMBER CORR UP | 756 | 494 | All SZGR 2.0 genes in this pathway |
| KIM ALL DISORDERS CALB1 CORR UP | 548 | 370 | All SZGR 2.0 genes in this pathway |
| MIYAGAWA TARGETS OF EWSR1 ETS FUSIONS DN | 229 | 135 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |
| PEDERSEN METASTASIS BY ERBB2 ISOFORM 4 | 110 | 66 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |