Gene Page: ZFR2
Summary ?
| GeneID | 23217 |
| Symbol | ZFR2 |
| Synonyms | KIAA1086 |
| Description | zinc finger RNA binding protein 2 |
| Reference | HGNC:HGNC:29189|Ensembl:ENSG00000105278|Vega:OTTHUMG00000180918 |
| Gene type | protein-coding |
| Map location | 19p13.3 |
| Pascal p-value | 0.116 |
| Sherlock p-value | 0.59 |
| Fetal beta | -0.394 |
| eGene | Cerebellar Hemisphere |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
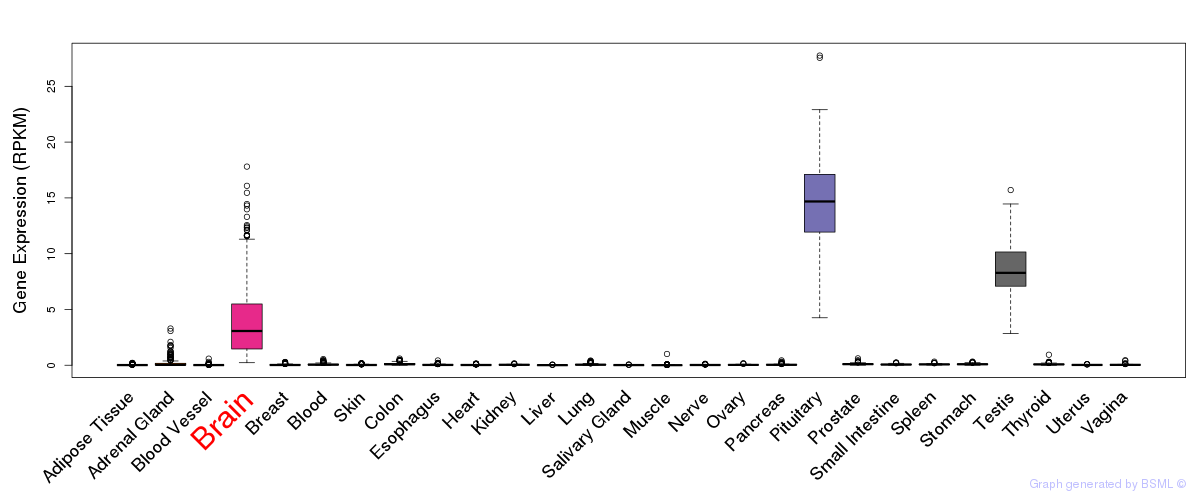
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MTHFD1 | 0.74 | 0.68 |
| TMCO7 | 0.72 | 0.74 |
| NBR1 | 0.72 | 0.69 |
| SEC23B | 0.71 | 0.73 |
| SND1 | 0.71 | 0.68 |
| TERF2IP | 0.70 | 0.72 |
| ZNF207 | 0.70 | 0.67 |
| RPA1 | 0.70 | 0.66 |
| FBXO18 | 0.70 | 0.67 |
| ZFR | 0.69 | 0.71 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.21 | -0.55 | -0.54 |
| MT-CO2 | -0.55 | -0.56 |
| AF347015.8 | -0.52 | -0.54 |
| AF347015.33 | -0.52 | -0.53 |
| AF347015.31 | -0.52 | -0.53 |
| MT-CYB | -0.50 | -0.52 |
| AF347015.2 | -0.50 | -0.50 |
| AF347015.27 | -0.49 | -0.53 |
| AF347015.26 | -0.49 | -0.52 |
| AF347015.18 | -0.49 | -0.58 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PYEON HPV POSITIVE TUMORS UP | 98 | 47 | All SZGR 2.0 genes in this pathway |
| SLEBOS HEAD AND NECK CANCER WITH HPV UP | 84 | 43 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 1 UP | 276 | 165 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 2 UP | 139 | 83 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 SIGNATURE 3 UP | 341 | 197 | All SZGR 2.0 genes in this pathway |
| KOYAMA SEMA3B TARGETS DN | 411 | 249 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL AND BRAIN QTL TRANS | 185 | 114 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 18HR DN | 178 | 121 | All SZGR 2.0 genes in this pathway |
| TSENG IRS1 TARGETS DN | 135 | 88 | All SZGR 2.0 genes in this pathway |
| NAKAYAMA FGF2 TARGETS | 29 | 17 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP A | 898 | 516 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |