Gene Page: VPS39
Summary ?
| GeneID | 23339 |
| Symbol | VPS39 |
| Synonyms | TLP|VAM6|hVam6p |
| Description | VPS39, HOPS complex subunit |
| Reference | MIM:612188|HGNC:HGNC:20593|Ensembl:ENSG00000166887|HPRD:10300|Vega:OTTHUMG00000130467 |
| Gene type | protein-coding |
| Map location | 15q15.1 |
| Pascal p-value | 3.627E-4 |
| Sherlock p-value | 0.064 |
| eGene | Hippocampus Myers' cis & trans |
| Support | CompositeSet Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Xu_2012 | Whole Exome Sequencing analysis | De novo mutations of 4 genes were identified by exome sequencing of 795 samples in this study | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| VPS39 | chr15 | 42454322 | G | A | NM_015289 | p.790R>C | missense | Schizophrenia | DNM:Xu_2012 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs4864466 | chr4 | 54403975 | VPS39 | 23339 | 0.2 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
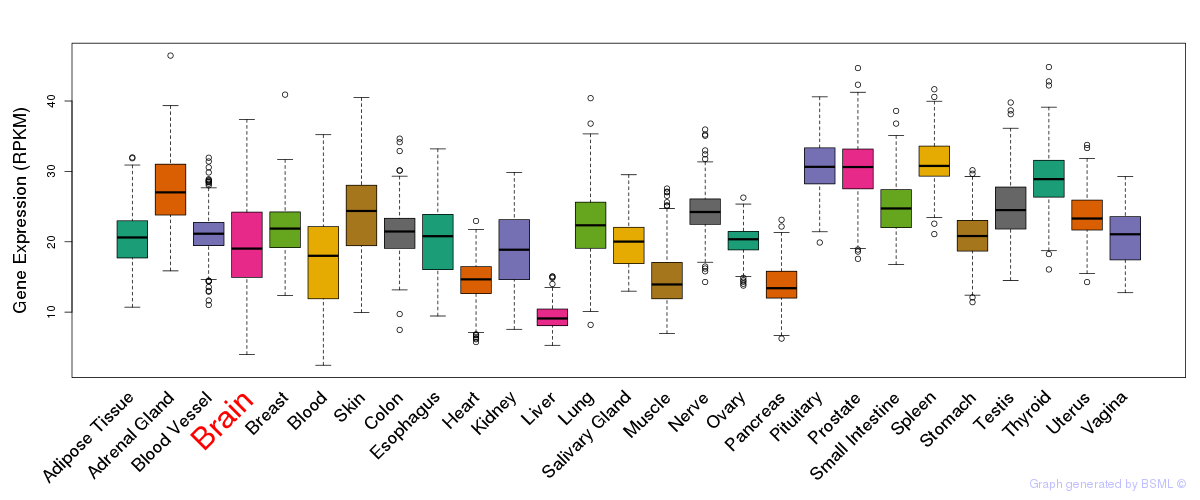
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| LRRC40 | 0.89 | 0.89 |
| ASNSD1 | 0.89 | 0.90 |
| PWP1 | 0.88 | 0.90 |
| FAM164A | 0.88 | 0.89 |
| ARMCX1 | 0.88 | 0.89 |
| COPB1 | 0.88 | 0.91 |
| CDC42 | 0.87 | 0.87 |
| NCUBE1 | 0.87 | 0.87 |
| CAMLG | 0.87 | 0.89 |
| RALA | 0.87 | 0.87 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.74 | -0.81 |
| AF347015.33 | -0.72 | -0.80 |
| AF347015.8 | -0.71 | -0.82 |
| MT-CYB | -0.71 | -0.81 |
| AF347015.26 | -0.71 | -0.84 |
| AF347015.27 | -0.70 | -0.78 |
| AF347015.31 | -0.70 | -0.79 |
| AF347015.2 | -0.70 | -0.82 |
| AF347015.15 | -0.70 | -0.81 |
| AIFM3 | -0.69 | -0.77 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PENG LEUCINE DEPRIVATION UP | 142 | 93 | All SZGR 2.0 genes in this pathway |
| NOUZOVA TRETINOIN AND H4 ACETYLATION | 143 | 85 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 48HR DN | 504 | 323 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR UP | 783 | 442 | All SZGR 2.0 genes in this pathway |
| IWANAGA CARCINOGENESIS BY KRAS PTEN DN | 353 | 226 | All SZGR 2.0 genes in this pathway |
| CADWELL ATG16L1 TARGETS DN | 70 | 43 | All SZGR 2.0 genes in this pathway |
| KIM ALL DISORDERS OLIGODENDROCYTE NUMBER CORR UP | 756 | 494 | All SZGR 2.0 genes in this pathway |