Gene Page: PRPF40B
Summary ?
| GeneID | 25766 |
| Symbol | PRPF40B |
| Synonyms | HYPC |
| Description | pre-mRNA processing factor 40 homolog B |
| Reference | HGNC:HGNC:25031|Ensembl:ENSG00000110844|HPRD:11044|Vega:OTTHUMG00000169568 |
| Gene type | protein-coding |
| Map location | 12q13.12 |
| Pascal p-value | 0.053 |
| DMG | 2 (# studies) |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Nishioka_2013 | Genome-wide DNA methylation analysis | The authors investigated the methylation profiles of DNA in peripheral blood cells from 18 patients with first-episode schizophrenia (FESZ) and from 15 normal controls. | 3 |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 3 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg10727759 | 12 | 50017361 | PRPF40B | 4.73E-6 | -0.274 | 0.01 | DMG:Wockner_2014 |
| cg05165862 | 12 | 50030609 | PRPF40B | 4.043E-4 | 0.335 | 0.044 | DMG:Wockner_2014 |
| cg08512940 | 12 | 50016787 | PRPF40B | -0.018 | 0.6 | DMG:Nishioka_2013 |
Section II. Transcriptome annotation
General gene expression (GTEx)
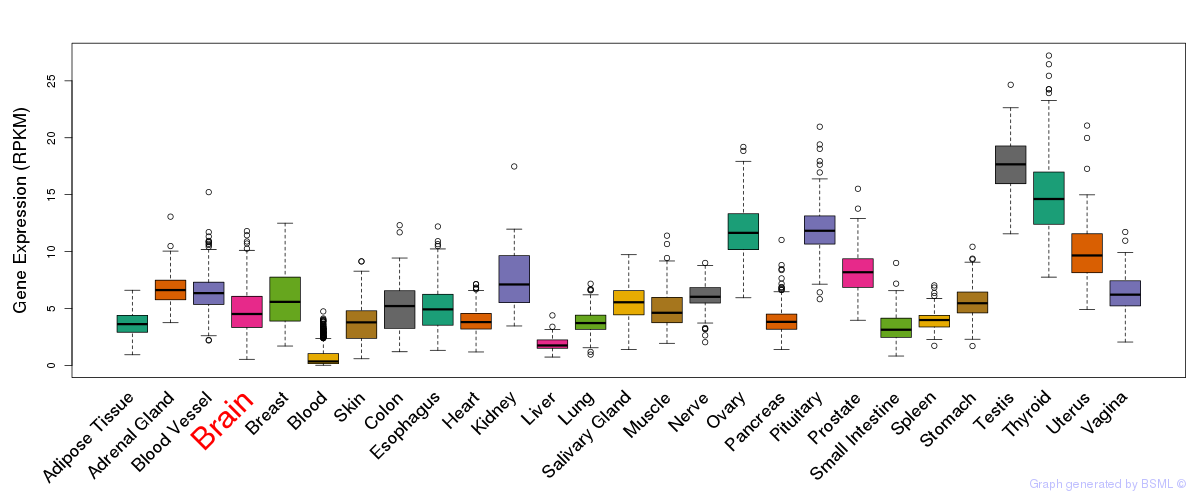
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| GLOD4 | 0.92 | 0.93 |
| C9orf30 | 0.91 | 0.91 |
| SUMO2 | 0.91 | 0.94 |
| PPM1G | 0.91 | 0.91 |
| ARMCX1 | 0.90 | 0.92 |
| BOP1 | 0.90 | 0.92 |
| CYTH2 | 0.90 | 0.93 |
| ZNF830 | 0.90 | 0.92 |
| ZNF259 | 0.90 | 0.90 |
| ARPC5 | 0.90 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| HLA-F | -0.76 | -0.81 |
| TSC22D4 | -0.75 | -0.82 |
| HEPN1 | -0.75 | -0.80 |
| MT-CO2 | -0.74 | -0.84 |
| AF347015.31 | -0.74 | -0.84 |
| AF347015.33 | -0.74 | -0.85 |
| TINAGL1 | -0.72 | -0.79 |
| MT-CYB | -0.72 | -0.84 |
| AIFM3 | -0.72 | -0.77 |
| FXYD1 | -0.71 | -0.82 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG SPLICEOSOME | 128 | 72 | All SZGR 2.0 genes in this pathway |
| GAUSSMANN MLL AF4 FUSION TARGETS C UP | 170 | 114 | All SZGR 2.0 genes in this pathway |