Gene Page: DHRS7B
Summary ?
| GeneID | 25979 |
| Symbol | DHRS7B |
| Synonyms | CGI-93|SDR32C1 |
| Description | dehydrogenase/reductase (SDR family) member 7B |
| Reference | MIM:616160|HGNC:HGNC:24547|Ensembl:ENSG00000109016|HPRD:13215|Vega:OTTHUMG00000178944 |
| Gene type | protein-coding |
| Map location | 17p12 |
| Pascal p-value | 0.348 |
| Sherlock p-value | 3.539E-5 |
| Fetal beta | -1.117 |
| Support | G2Cdb.human_BAYES-COLLINS-MOUSE-PSD-CONSENSUS G2Cdb.humanPSD G2Cdb.humanPSP |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
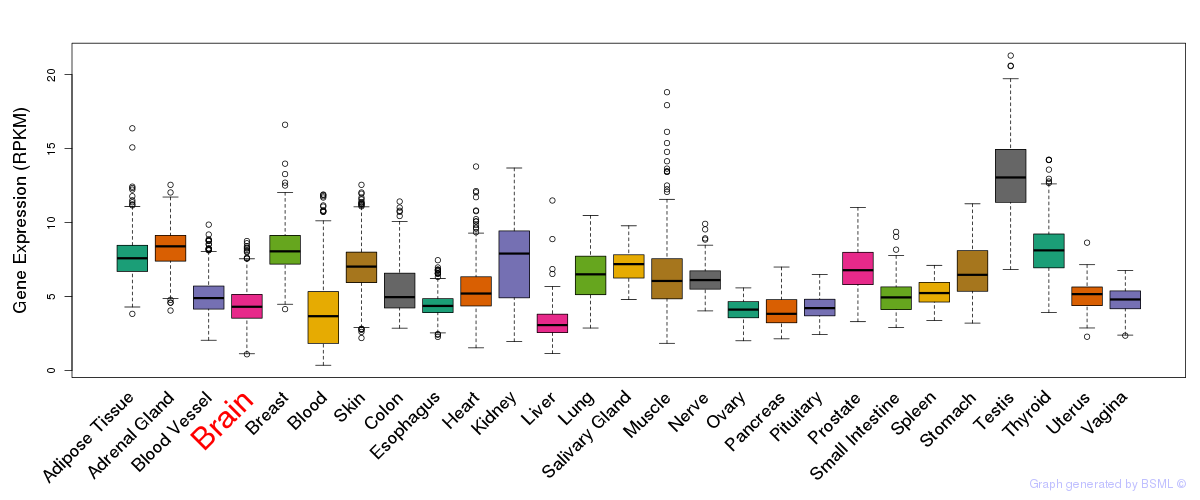
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| EML1 | 0.95 | 0.91 |
| AFF3 | 0.93 | 0.93 |
| NEUROD2 | 0.93 | 0.86 |
| ZNF238 | 0.92 | 0.92 |
| DAB1 | 0.92 | 0.88 |
| SSBP2 | 0.92 | 0.91 |
| MED13L | 0.92 | 0.90 |
| LZTS1 | 0.92 | 0.88 |
| KCTD12 | 0.92 | 0.82 |
| BACH2 | 0.91 | 0.87 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SERPINB6 | -0.69 | -0.80 |
| HSD17B14 | -0.64 | -0.84 |
| HEPN1 | -0.63 | -0.79 |
| AIFM3 | -0.63 | -0.81 |
| TSC22D4 | -0.62 | -0.81 |
| BDH2 | -0.62 | -0.81 |
| HLA-F | -0.61 | -0.75 |
| RAMP1 | -0.61 | -0.79 |
| FAH | -0.61 | -0.76 |
| S100B | -0.61 | -0.85 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PICCALUGA ANGIOIMMUNOBLASTIC LYMPHOMA UP | 205 | 140 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN MLL SIGNATURE 1 UP | 380 | 236 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN MLL SIGNATURE 2 UP | 418 | 263 | All SZGR 2.0 genes in this pathway |
| BERENJENO TRANSFORMED BY RHOA UP | 536 | 340 | All SZGR 2.0 genes in this pathway |
| LASTOWSKA NEUROBLASTOMA COPY NUMBER UP | 181 | 108 | All SZGR 2.0 genes in this pathway |
| NUYTTEN EZH2 TARGETS DN | 1024 | 594 | All SZGR 2.0 genes in this pathway |
| BURTON ADIPOGENESIS 6 | 189 | 112 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY E2F4 UNSTIMULATED | 728 | 415 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| MILI PSEUDOPODIA HAPTOTAXIS DN | 668 | 419 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE LATE | 1137 | 655 | All SZGR 2.0 genes in this pathway |