Gene Page: TIMM10
Summary ?
| GeneID | 26519 |
| Symbol | TIMM10 |
| Synonyms | TIM10|TIM10A |
| Description | translocase of inner mitochondrial membrane 10 homolog (yeast) |
| Reference | MIM:602251|HGNC:HGNC:11814|Ensembl:ENSG00000134809|HPRD:03768|Vega:OTTHUMG00000167039 |
| Gene type | protein-coding |
| Map location | 11q12.1 |
| Pascal p-value | 0.005 |
| Sherlock p-value | 0.958 |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs2584858 | chr11 | 57207076 | TIMM10 | 26519 | 0.14 | cis | ||
| rs2584859 | chr11 | 57207446 | TIMM10 | 26519 | 0.14 | cis | ||
| rs2581924 | chr11 | 57208467 | TIMM10 | 26519 | 0.07 | cis | ||
| rs2649667 | chr11 | 57270508 | TIMM10 | 26519 | 3.224E-8 | cis | ||
| rs4939148 | chr11 | 57290960 | TIMM10 | 26519 | 0.08 | cis | ||
| rs10896623 | chr11 | 57295863 | TIMM10 | 26519 | 0.02 | cis | ||
| rs2729371 | chr11 | 57321321 | TIMM10 | 26519 | 9.197E-8 | cis | ||
| rs4938871 | chr11 | 57329401 | TIMM10 | 26519 | 5.418E-5 | cis | ||
| rs7950126 | chr11 | 57349186 | TIMM10 | 26519 | 0 | cis | ||
| rs2649662 | chr11 | 57352561 | TIMM10 | 26519 | 5.708E-8 | cis | ||
| rs2649663 | chr11 | 57357304 | TIMM10 | 26519 | 2.342E-7 | cis | ||
| rs2649667 | chr11 | 57270508 | TIMM10 | 26519 | 7.515E-6 | trans | ||
| rs2729371 | chr11 | 57321321 | TIMM10 | 26519 | 2.001E-5 | trans | ||
| rs4938871 | chr11 | 57329401 | TIMM10 | 26519 | 0.01 | trans | ||
| rs7950126 | chr11 | 57349186 | TIMM10 | 26519 | 0.19 | trans | ||
| rs2649662 | chr11 | 57352561 | TIMM10 | 26519 | 1.288E-5 | trans | ||
| rs2649663 | chr11 | 57357304 | TIMM10 | 26519 | 4.884E-5 | trans | ||
| rs11229035 | 11 | 57216746 | TIMM10 | ENSG00000134809.4 | 1.325E-6 | 0 | 81530 | gtex_brain_ba24 |
| rs2581921 | 11 | 57222607 | TIMM10 | ENSG00000134809.4 | 4.058E-6 | 0 | 75669 | gtex_brain_ba24 |
| rs2729394 | 11 | 57233567 | TIMM10 | ENSG00000134809.4 | 2.834E-6 | 0 | 64709 | gtex_brain_ba24 |
| rs2258835 | 11 | 57243588 | TIMM10 | ENSG00000134809.4 | 2.794E-6 | 0 | 54688 | gtex_brain_ba24 |
| rs2729353 | 11 | 57248003 | TIMM10 | ENSG00000134809.4 | 2.508E-6 | 0 | 50273 | gtex_brain_ba24 |
| rs2581923 | 11 | 57249991 | TIMM10 | ENSG00000134809.4 | 1.597E-6 | 0 | 48285 | gtex_brain_ba24 |
| rs2848634 | 11 | 57252328 | TIMM10 | ENSG00000134809.4 | 1.597E-6 | 0 | 45948 | gtex_brain_ba24 |
| rs2848635 | 11 | 57253931 | TIMM10 | ENSG00000134809.4 | 1.597E-6 | 0 | 44345 | gtex_brain_ba24 |
| rs2729384 | 11 | 57258617 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 39659 | gtex_brain_ba24 |
| rs10750864 | 11 | 57259694 | TIMM10 | ENSG00000134809.4 | 2.22E-6 | 0 | 38582 | gtex_brain_ba24 |
| rs2509899 | 11 | 57262596 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 35680 | gtex_brain_ba24 |
| rs2848640 | 11 | 57262680 | TIMM10 | ENSG00000134809.4 | 1.433E-6 | 0 | 35596 | gtex_brain_ba24 |
| rs2729387 | 11 | 57262699 | TIMM10 | ENSG00000134809.4 | 1.535E-6 | 0 | 35577 | gtex_brain_ba24 |
| rs2581922 | 11 | 57265400 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 32876 | gtex_brain_ba24 |
| rs1001956 | 11 | 57266949 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 31327 | gtex_brain_ba24 |
| rs2584856 | 11 | 57267312 | TIMM10 | ENSG00000134809.4 | 8.143E-7 | 0 | 30964 | gtex_brain_ba24 |
| rs2250673 | 11 | 57268170 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 30106 | gtex_brain_ba24 |
| rs56032818 | 11 | 57269429 | TIMM10 | ENSG00000134809.4 | 1.971E-6 | 0 | 28847 | gtex_brain_ba24 |
| rs2649667 | 11 | 57270509 | TIMM10 | ENSG00000134809.4 | 1.435E-6 | 0 | 27767 | gtex_brain_ba24 |
| rs397802949 | 11 | 57274179 | TIMM10 | ENSG00000134809.4 | 3.125E-6 | 0 | 24097 | gtex_brain_ba24 |
| rs2511984 | 11 | 57278733 | TIMM10 | ENSG00000134809.4 | 8.921E-8 | 0 | 19543 | gtex_brain_ba24 |
| rs2467327 | 11 | 57279228 | TIMM10 | ENSG00000134809.4 | 3.046E-8 | 0 | 19048 | gtex_brain_ba24 |
| rs2848626 | 11 | 57283988 | TIMM10 | ENSG00000134809.4 | 6.497E-10 | 0 | 14288 | gtex_brain_ba24 |
| rs2729371 | 11 | 57321322 | TIMM10 | ENSG00000134809.4 | 1.848E-10 | 0 | -23046 | gtex_brain_ba24 |
| rs2649652 | 11 | 57324428 | TIMM10 | ENSG00000134809.4 | 4.759E-10 | 0 | -26152 | gtex_brain_ba24 |
| rs2729374 | 11 | 57326927 | TIMM10 | ENSG00000134809.4 | 4.784E-10 | 0 | -28651 | gtex_brain_ba24 |
| rs2649650 | 11 | 57327567 | TIMM10 | ENSG00000134809.4 | 3.985E-10 | 0 | -29291 | gtex_brain_ba24 |
| rs2848625 | 11 | 57330650 | TIMM10 | ENSG00000134809.4 | 3.985E-10 | 0 | -32374 | gtex_brain_ba24 |
| rs2649648 | 11 | 57331337 | TIMM10 | ENSG00000134809.4 | 1.334E-7 | 0 | -33061 | gtex_brain_ba24 |
| rs2511983 | 11 | 57346874 | TIMM10 | ENSG00000134809.4 | 3.985E-10 | 0 | -48598 | gtex_brain_ba24 |
| rs489858 | 11 | 57347525 | TIMM10 | ENSG00000134809.4 | 2.073E-10 | 0 | -49249 | gtex_brain_ba24 |
| rs1967179 | 11 | 57348483 | TIMM10 | ENSG00000134809.4 | 1.797E-7 | 0 | -50207 | gtex_brain_ba24 |
| rs2649662 | 11 | 57352562 | TIMM10 | ENSG00000134809.4 | 3.26E-10 | 0 | -54286 | gtex_brain_ba24 |
| rs2729356 | 11 | 57353397 | TIMM10 | ENSG00000134809.4 | 6.893E-8 | 0 | -55121 | gtex_brain_ba24 |
| rs2729355 | 11 | 57353983 | TIMM10 | ENSG00000134809.4 | 4.784E-10 | 0 | -55707 | gtex_brain_ba24 |
| rs2729354 | 11 | 57355970 | TIMM10 | ENSG00000134809.4 | 3.844E-10 | 0 | -57694 | gtex_brain_ba24 |
| rs2848630 | 11 | 57356907 | TIMM10 | ENSG00000134809.4 | 4.784E-10 | 0 | -58631 | gtex_brain_ba24 |
| rs2649663 | 11 | 57357305 | TIMM10 | ENSG00000134809.4 | 4.787E-10 | 0 | -59029 | gtex_brain_ba24 |
| rs1846567 | 11 | 57357593 | TIMM10 | ENSG00000134809.4 | 7.978E-10 | 0 | -59317 | gtex_brain_ba24 |
| rs1846568 | 11 | 57357714 | TIMM10 | ENSG00000134809.4 | 7.782E-10 | 0 | -59438 | gtex_brain_ba24 |
| rs2454663 | 11 | 57358083 | TIMM10 | ENSG00000134809.4 | 1.53E-9 | 0 | -59807 | gtex_brain_ba24 |
| rs2511992 | 11 | 57358343 | TIMM10 | ENSG00000134809.4 | 2.455E-7 | 0 | -60067 | gtex_brain_ba24 |
| rs2581921 | 11 | 57222607 | TIMM10 | ENSG00000134809.4 | 4.816E-9 | 0 | 75669 | gtex_brain_putamen_basal |
| rs2729363 | 11 | 57226854 | TIMM10 | ENSG00000134809.4 | 2.011E-10 | 0 | 71422 | gtex_brain_putamen_basal |
| rs2511986 | 11 | 57231559 | TIMM10 | ENSG00000134809.4 | 1.146E-10 | 0 | 66717 | gtex_brain_putamen_basal |
| rs2729394 | 11 | 57233567 | TIMM10 | ENSG00000134809.4 | 1.212E-11 | 0 | 64709 | gtex_brain_putamen_basal |
| rs3851117 | 11 | 57237113 | TIMM10 | ENSG00000134809.4 | 5.08E-10 | 0 | 61163 | gtex_brain_putamen_basal |
| rs2584862 | 11 | 57241956 | TIMM10 | ENSG00000134809.4 | 2.22E-11 | 0 | 56320 | gtex_brain_putamen_basal |
| rs2258835 | 11 | 57243588 | TIMM10 | ENSG00000134809.4 | 1.939E-11 | 0 | 54688 | gtex_brain_putamen_basal |
| rs2729353 | 11 | 57248003 | TIMM10 | ENSG00000134809.4 | 4.024E-10 | 0 | 50273 | gtex_brain_putamen_basal |
| rs2581923 | 11 | 57249991 | TIMM10 | ENSG00000134809.4 | 1.111E-10 | 0 | 48285 | gtex_brain_putamen_basal |
| rs2729377 | 11 | 57250084 | TIMM10 | ENSG00000134809.4 | 6.643E-10 | 0 | 48192 | gtex_brain_putamen_basal |
| rs2848634 | 11 | 57252328 | TIMM10 | ENSG00000134809.4 | 1.111E-10 | 0 | 45948 | gtex_brain_putamen_basal |
| rs2848635 | 11 | 57253931 | TIMM10 | ENSG00000134809.4 | 1.111E-10 | 0 | 44345 | gtex_brain_putamen_basal |
| rs2439890 | 11 | 57254088 | TIMM10 | ENSG00000134809.4 | 1.753E-8 | 0 | 44188 | gtex_brain_putamen_basal |
| rs2729384 | 11 | 57258617 | TIMM10 | ENSG00000134809.4 | 3.704E-12 | 0 | 39659 | gtex_brain_putamen_basal |
| rs10750864 | 11 | 57259694 | TIMM10 | ENSG00000134809.4 | 1.044E-11 | 0 | 38582 | gtex_brain_putamen_basal |
| rs2509899 | 11 | 57262596 | TIMM10 | ENSG00000134809.4 | 3.704E-12 | 0 | 35680 | gtex_brain_putamen_basal |
| rs2848640 | 11 | 57262680 | TIMM10 | ENSG00000134809.4 | 4.854E-12 | 0 | 35596 | gtex_brain_putamen_basal |
| rs2729387 | 11 | 57262699 | TIMM10 | ENSG00000134809.4 | 1.196E-10 | 0 | 35577 | gtex_brain_putamen_basal |
| rs2848641 | 11 | 57263068 | TIMM10 | ENSG00000134809.4 | 2.149E-9 | 0 | 35208 | gtex_brain_putamen_basal |
| rs60293750 | 11 | 57264674 | TIMM10 | ENSG00000134809.4 | 1.442E-9 | 0 | 33602 | gtex_brain_putamen_basal |
| rs2581922 | 11 | 57265400 | TIMM10 | ENSG00000134809.4 | 6.573E-11 | 0 | 32876 | gtex_brain_putamen_basal |
| rs1001956 | 11 | 57266949 | TIMM10 | ENSG00000134809.4 | 4.576E-11 | 0 | 31327 | gtex_brain_putamen_basal |
| rs2584856 | 11 | 57267312 | TIMM10 | ENSG00000134809.4 | 5.764E-11 | 0 | 30964 | gtex_brain_putamen_basal |
| rs2250673 | 11 | 57268170 | TIMM10 | ENSG00000134809.4 | 6.573E-11 | 0 | 30106 | gtex_brain_putamen_basal |
| rs56032818 | 11 | 57269429 | TIMM10 | ENSG00000134809.4 | 2.261E-9 | 0 | 28847 | gtex_brain_putamen_basal |
| rs2649667 | 11 | 57270509 | TIMM10 | ENSG00000134809.4 | 6.573E-11 | 0 | 27767 | gtex_brain_putamen_basal |
| rs397802949 | 11 | 57274179 | TIMM10 | ENSG00000134809.4 | 1.71E-9 | 0 | 24097 | gtex_brain_putamen_basal |
| rs2511984 | 11 | 57278733 | TIMM10 | ENSG00000134809.4 | 9.844E-13 | 0 | 19543 | gtex_brain_putamen_basal |
| rs2467327 | 11 | 57279228 | TIMM10 | ENSG00000134809.4 | 9.628E-13 | 0 | 19048 | gtex_brain_putamen_basal |
| rs2848626 | 11 | 57283988 | TIMM10 | ENSG00000134809.4 | 9.088E-13 | 0 | 14288 | gtex_brain_putamen_basal |
| rs2649665 | 11 | 57289328 | TIMM10 | ENSG00000134809.4 | 1.792E-6 | 0 | 8948 | gtex_brain_putamen_basal |
| rs2729371 | 11 | 57321322 | TIMM10 | ENSG00000134809.4 | 6.481E-12 | 0 | -23046 | gtex_brain_putamen_basal |
| rs2649652 | 11 | 57324428 | TIMM10 | ENSG00000134809.4 | 4.874E-10 | 0 | -26152 | gtex_brain_putamen_basal |
| rs2729374 | 11 | 57326927 | TIMM10 | ENSG00000134809.4 | 1.198E-10 | 0 | -28651 | gtex_brain_putamen_basal |
| rs2649650 | 11 | 57327567 | TIMM10 | ENSG00000134809.4 | 6.332E-10 | 0 | -29291 | gtex_brain_putamen_basal |
| rs2848625 | 11 | 57330650 | TIMM10 | ENSG00000134809.4 | 6.326E-10 | 0 | -32374 | gtex_brain_putamen_basal |
| rs2511983 | 11 | 57346874 | TIMM10 | ENSG00000134809.4 | 9.684E-10 | 0 | -48598 | gtex_brain_putamen_basal |
| rs489858 | 11 | 57347525 | TIMM10 | ENSG00000134809.4 | 3.101E-9 | 0 | -49249 | gtex_brain_putamen_basal |
| rs2649662 | 11 | 57352562 | TIMM10 | ENSG00000134809.4 | 4.403E-9 | 0 | -54286 | gtex_brain_putamen_basal |
| rs2729355 | 11 | 57353983 | TIMM10 | ENSG00000134809.4 | 1.422E-10 | 0 | -55707 | gtex_brain_putamen_basal |
| rs2729354 | 11 | 57355970 | TIMM10 | ENSG00000134809.4 | 8.111E-10 | 0 | -57694 | gtex_brain_putamen_basal |
| rs2848630 | 11 | 57356907 | TIMM10 | ENSG00000134809.4 | 1.422E-10 | 0 | -58631 | gtex_brain_putamen_basal |
| rs2649663 | 11 | 57357305 | TIMM10 | ENSG00000134809.4 | 7.346E-10 | 0 | -59029 | gtex_brain_putamen_basal |
| rs1846567 | 11 | 57357593 | TIMM10 | ENSG00000134809.4 | 1.466E-10 | 0 | -59317 | gtex_brain_putamen_basal |
| rs1846568 | 11 | 57357714 | TIMM10 | ENSG00000134809.4 | 8.832E-10 | 0 | -59438 | gtex_brain_putamen_basal |
| rs2454663 | 11 | 57358083 | TIMM10 | ENSG00000134809.4 | 1.398E-9 | 0 | -59807 | gtex_brain_putamen_basal |
| rs2511992 | 11 | 57358343 | TIMM10 | ENSG00000134809.4 | 2.635E-7 | 0 | -60067 | gtex_brain_putamen_basal |
| rs11229169 | 11 | 57703725 | TIMM10 | ENSG00000134809.4 | 2.657E-6 | 0 | -405449 | gtex_brain_putamen_basal |
| rs1596992 | 11 | 57866752 | TIMM10 | ENSG00000134809.4 | 3.364E-6 | 0 | -568476 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
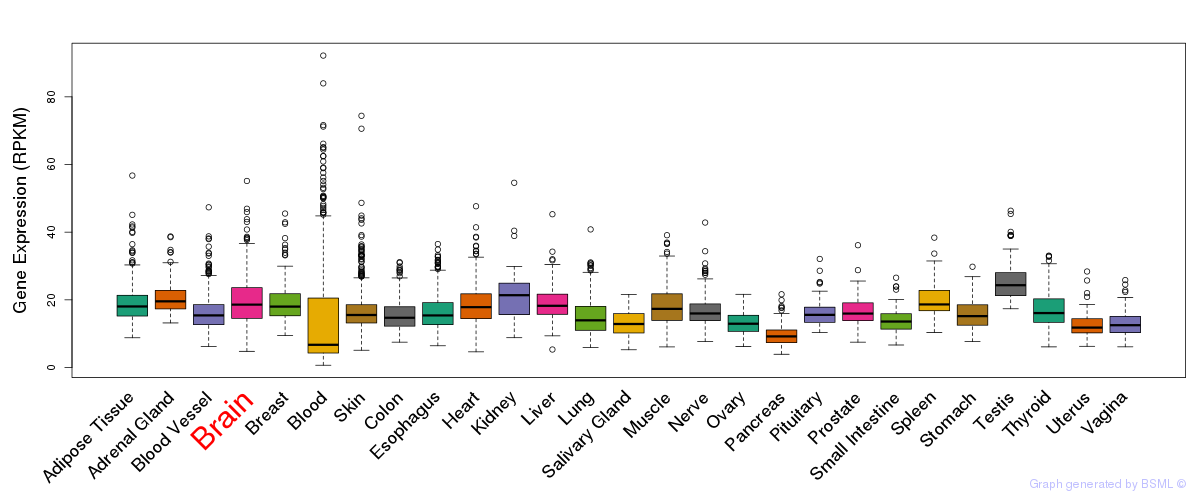
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME MITOCHONDRIAL PROTEIN IMPORT | 58 | 24 | All SZGR 2.0 genes in this pathway |
| REACTOME METABOLISM OF PROTEINS | 518 | 242 | All SZGR 2.0 genes in this pathway |
| SOTIRIOU BREAST CANCER GRADE 1 VS 3 UP | 151 | 84 | All SZGR 2.0 genes in this pathway |
| FULCHER INFLAMMATORY RESPONSE LECTIN VS LPS UP | 579 | 346 | All SZGR 2.0 genes in this pathway |
| CASORELLI ACUTE PROMYELOCYTIC LEUKEMIA DN | 663 | 425 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA DN | 1375 | 806 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN DN | 770 | 415 | All SZGR 2.0 genes in this pathway |
| GOERING BLOOD HDL CHOLESTEROL QTL CIS | 13 | 7 | All SZGR 2.0 genes in this pathway |
| NUYTTEN EZH2 TARGETS DN | 1024 | 594 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| WEI MYCN TARGETS WITH E BOX | 795 | 478 | All SZGR 2.0 genes in this pathway |
| GARCIA TARGETS OF FLI1 AND DAX1 DN | 176 | 104 | All SZGR 2.0 genes in this pathway |
| AMIT EGF RESPONSE 480 MCF10A | 43 | 26 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR DN | 911 | 527 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR DN | 1011 | 592 | All SZGR 2.0 genes in this pathway |
| GAVIN FOXP3 TARGETS CLUSTER T4 | 94 | 69 | All SZGR 2.0 genes in this pathway |
| WONG MITOCHONDRIA GENE MODULE | 217 | 122 | All SZGR 2.0 genes in this pathway |
| SARRIO EPITHELIAL MESENCHYMAL TRANSITION UP | 180 | 114 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER UP | 973 | 570 | All SZGR 2.0 genes in this pathway |
| ALONSO METASTASIS UP | 198 | 128 | All SZGR 2.0 genes in this pathway |
| MOOTHA HUMAN MITODB 6 2002 | 429 | 260 | All SZGR 2.0 genes in this pathway |
| MOOTHA MITOCHONDRIA | 447 | 277 | All SZGR 2.0 genes in this pathway |
| COLINA TARGETS OF 4EBP1 AND 4EBP2 | 356 | 214 | All SZGR 2.0 genes in this pathway |
| DANG REGULATED BY MYC UP | 72 | 53 | All SZGR 2.0 genes in this pathway |
| DANG MYC TARGETS UP | 143 | 100 | All SZGR 2.0 genes in this pathway |
| DANG BOUND BY MYC | 1103 | 714 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE DN | 841 | 431 | All SZGR 2.0 genes in this pathway |
| BAKKER FOXO3 TARGETS DN | 187 | 109 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS DN | 882 | 538 | All SZGR 2.0 genes in this pathway |