Gene Page: CLEC4G
Summary ?
| GeneID | 339390 |
| Symbol | CLEC4G |
| Synonyms | DTTR431|LP2698|LSECtin|UNQ431 |
| Description | C-type lectin domain family 4 member G |
| Reference | MIM:616256|HGNC:HGNC:24591|Ensembl:ENSG00000182566|HPRD:16721|Vega:OTTHUMG00000182519 |
| Gene type | protein-coding |
| Map location | 19p13.2 |
| Pascal p-value | 0.941 |
| Fetal beta | -1.328 |
| DMG | 2 (# studies) |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 2 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg09772661 | 19 | 7794952 | CLEC4G | 2.03E-5 | -0.785 | 0.016 | DMG:Wockner_2014 |
| cg12768640 | 19 | 7795428 | CLEC4G | 3.91E-10 | -0.018 | 7.56E-7 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs4653163 | chr1 | 36627444 | CLEC4G | 339390 | 0.04 | trans | ||
| rs1954436 | chr1 | 62248194 | CLEC4G | 339390 | 0.18 | trans | ||
| rs2816169 | chr1 | 179210675 | CLEC4G | 339390 | 0.1 | trans | ||
| rs17022378 | chr1 | 206654057 | CLEC4G | 339390 | 0.09 | trans | ||
| rs6753473 | chr2 | 26526418 | CLEC4G | 339390 | 0.01 | trans | ||
| rs17021846 | chr2 | 38243146 | CLEC4G | 339390 | 0.09 | trans | ||
| rs17028363 | chr2 | 41913161 | CLEC4G | 339390 | 0.01 | trans | ||
| rs16829545 | chr2 | 151977407 | CLEC4G | 339390 | 3.322E-9 | trans | ||
| rs16841750 | chr2 | 158288461 | CLEC4G | 339390 | 0.11 | trans | ||
| rs10183246 | chr2 | 237187882 | CLEC4G | 339390 | 0.01 | trans | ||
| rs10195546 | chr2 | 237187933 | CLEC4G | 339390 | 0.05 | trans | ||
| rs9810143 | chr3 | 5060209 | CLEC4G | 339390 | 0 | trans | ||
| rs6797307 | chr3 | 8601563 | CLEC4G | 339390 | 5.444E-4 | trans | ||
| rs662007 | chr4 | 125253160 | CLEC4G | 339390 | 0.03 | trans | ||
| rs17008119 | chr4 | 125321425 | CLEC4G | 339390 | 0.01 | trans | ||
| rs7692715 | chr4 | 125335209 | CLEC4G | 339390 | 0 | trans | ||
| rs17762315 | chr5 | 76807576 | CLEC4G | 339390 | 0 | trans | ||
| rs7729096 | chr5 | 76835927 | CLEC4G | 339390 | 0 | trans | ||
| rs13360496 | 0 | CLEC4G | 339390 | 0.01 | trans | |||
| rs16890367 | chr6 | 38078448 | CLEC4G | 339390 | 0 | trans | ||
| rs1511745 | chr6 | 165954685 | CLEC4G | 339390 | 0.01 | trans | ||
| rs11965579 | chr6 | 165969504 | CLEC4G | 339390 | 0.11 | trans | ||
| rs12706918 | chr7 | 80266147 | CLEC4G | 339390 | 0.07 | trans | ||
| rs6996695 | chr8 | 77540580 | CLEC4G | 339390 | 0.04 | trans | ||
| rs17124813 | chr10 | 110347464 | CLEC4G | 339390 | 0.01 | trans | ||
| rs4980232 | chr10 | 124577756 | CLEC4G | 339390 | 0.15 | trans | ||
| rs11025709 | chr11 | 20745072 | CLEC4G | 339390 | 0.1 | trans | ||
| rs506698 | chr11 | 79651244 | CLEC4G | 339390 | 0.03 | trans | ||
| rs7972699 | chr12 | 105155290 | CLEC4G | 339390 | 0.02 | trans | ||
| rs7488439 | chr12 | 105155814 | CLEC4G | 339390 | 0.01 | trans | ||
| rs17036489 | chr12 | 105292283 | CLEC4G | 339390 | 0.02 | trans | ||
| rs457696 | chr13 | 32581313 | CLEC4G | 339390 | 0.19 | trans | ||
| rs9566986 | chr13 | 32595026 | CLEC4G | 339390 | 0.13 | trans | ||
| rs7337967 | chr13 | 53519044 | CLEC4G | 339390 | 0.03 | trans | ||
| rs7337531 | chr13 | 109611085 | CLEC4G | 339390 | 0.08 | trans | ||
| rs16955618 | chr15 | 29937543 | CLEC4G | 339390 | 4.013E-11 | trans | ||
| rs17711752 | chr16 | 34436184 | CLEC4G | 339390 | 0.15 | trans | ||
| rs16968252 | chr17 | 31692447 | CLEC4G | 339390 | 0.02 | trans | ||
| rs11873184 | chr18 | 1584081 | CLEC4G | 339390 | 0.08 | trans | ||
| rs1041786 | chr21 | 22617710 | CLEC4G | 339390 | 0 | trans | ||
| rs16993189 | chrX | 144666166 | CLEC4G | 339390 | 0.1 | trans |
Section II. Transcriptome annotation
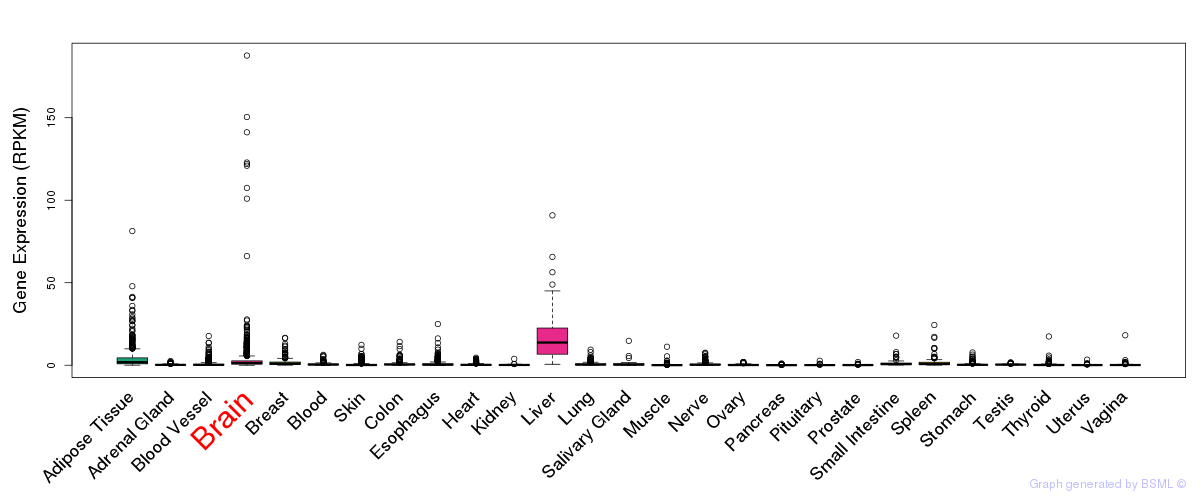
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| FULCHER INFLAMMATORY RESPONSE LECTIN VS LPS UP | 579 | 346 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN UP | 1142 | 669 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN UP | 612 | 367 | All SZGR 2.0 genes in this pathway |
| SHETH LIVER CANCER VS TXNIP LOSS PAM4 | 261 | 153 | All SZGR 2.0 genes in this pathway |
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| BENPORATH ES WITH H3K27ME3 | 1118 | 744 | All SZGR 2.0 genes in this pathway |
| KUMAR PATHOGEN LOAD BY MACROPHAGES | 275 | 155 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |
| NABA ECM AFFILIATED | 171 | 89 | All SZGR 2.0 genes in this pathway |
| NABA MATRISOME ASSOCIATED | 753 | 411 | All SZGR 2.0 genes in this pathway |
| NABA MATRISOME | 1028 | 559 | All SZGR 2.0 genes in this pathway |