Gene Page: NDUFS5
Summary ?
| GeneID | 4725 |
| Symbol | NDUFS5 |
| Synonyms | CI-15k|CI15K |
| Description | NADH:ubiquinone oxidoreductase subunit S5 |
| Reference | MIM:603847|HGNC:HGNC:7712|Ensembl:ENSG00000168653|HPRD:11961|Vega:OTTHUMG00000007497 |
| Gene type | protein-coding |
| Map location | 1p34.2-p33 |
| Pascal p-value | 0.014 |
| Sherlock p-value | 0.031 |
| Fetal beta | -0.509 |
| eGene | Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
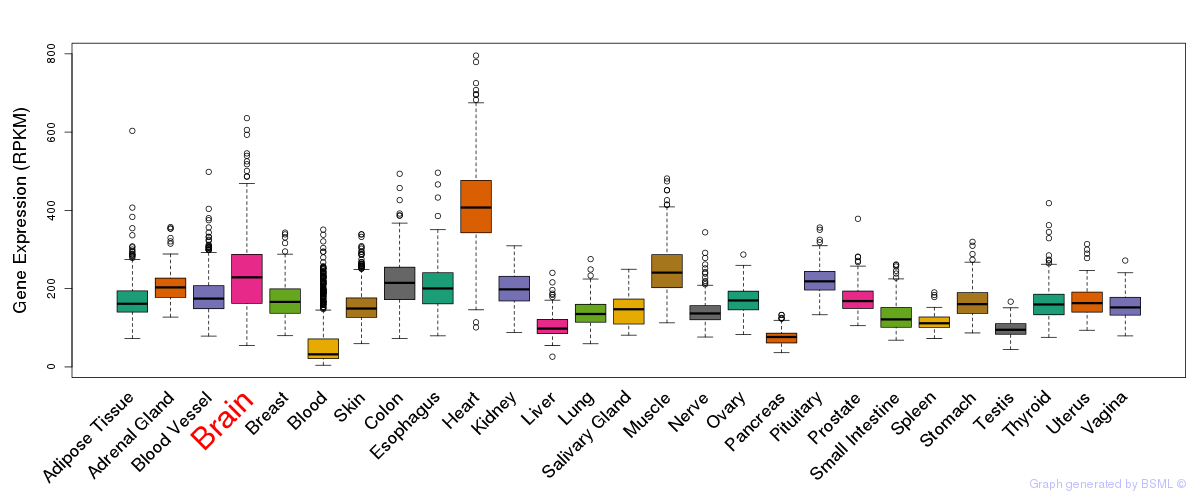
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MAD2L1 | 0.93 | 0.77 |
| CDC2 | 0.93 | 0.58 |
| CCNA2 | 0.92 | 0.45 |
| EXO1 | 0.92 | 0.56 |
| DLGAP5 | 0.92 | 0.38 |
| DEPDC1 | 0.92 | 0.57 |
| RRM2 | 0.91 | 0.42 |
| PBK | 0.91 | 0.42 |
| AURKA | 0.91 | 0.56 |
| MELK | 0.91 | 0.44 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SLC9A3R2 | -0.33 | -0.31 |
| AF347015.26 | -0.31 | -0.43 |
| AF347015.33 | -0.31 | -0.41 |
| MT-CYB | -0.30 | -0.40 |
| AF347015.15 | -0.30 | -0.40 |
| MT-CO2 | -0.30 | -0.39 |
| AF347015.27 | -0.30 | -0.40 |
| AF347015.8 | -0.30 | -0.40 |
| LGI4 | -0.29 | -0.22 |
| AIFM3 | -0.29 | -0.32 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG OXIDATIVE PHOSPHORYLATION | 135 | 73 | All SZGR 2.0 genes in this pathway |
| KEGG ALZHEIMERS DISEASE | 169 | 110 | All SZGR 2.0 genes in this pathway |
| KEGG PARKINSONS DISEASE | 133 | 78 | All SZGR 2.0 genes in this pathway |
| KEGG HUNTINGTONS DISEASE | 185 | 109 | All SZGR 2.0 genes in this pathway |
| REACTOME TCA CYCLE AND RESPIRATORY ELECTRON TRANSPORT | 141 | 85 | All SZGR 2.0 genes in this pathway |
| REACTOME RESPIRATORY ELECTRON TRANSPORT | 79 | 44 | All SZGR 2.0 genes in this pathway |
| REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | 98 | 56 | All SZGR 2.0 genes in this pathway |
| LANDIS ERBB2 BREAST TUMORS 324 DN | 149 | 93 | All SZGR 2.0 genes in this pathway |
| SCHLOSSER MYC AND SERUM RESPONSE SYNERGY | 32 | 22 | All SZGR 2.0 genes in this pathway |
| WONG MITOCHONDRIA GENE MODULE | 217 | 122 | All SZGR 2.0 genes in this pathway |
| MOOTHA PGC | 420 | 269 | All SZGR 2.0 genes in this pathway |
| SETLUR PROSTATE CANCER TMPRSS2 ERG FUSION UP | 67 | 48 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS DN | 1972 | 1213 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS UP | 682 | 433 | All SZGR 2.0 genes in this pathway |