Gene Page: VILL
Summary ?
| GeneID | 50853 |
| Symbol | VILL |
| Synonyms | - |
| Description | villin-like |
| Reference | HGNC:HGNC:30906|Ensembl:ENSG00000136059|HPRD:18283|Vega:OTTHUMG00000130814 |
| Gene type | protein-coding |
| Map location | 3p21.3 |
| Pascal p-value | 0.158 |
| Sherlock p-value | 0.202 |
| Fetal beta | -0.417 |
| DMG | 1 (# studies) |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellum Cortex Frontal Cortex BA9 Hippocampus Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
| GSMA_I | Genome scan meta-analysis | Psr: 0.006 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg25663755 | 3 | 38035463 | VILL | 1.062E-4 | 0.508 | 0.028 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs1967256 | chr5 | 90299222 | VILL | 50853 | 0.14 | trans | ||
| rs11954387 | chr5 | 90302116 | VILL | 50853 | 0.14 | trans | ||
| rs17182597 | chr7 | 12393594 | VILL | 50853 | 0.06 | trans | ||
| rs3011362 | chr10 | 122553193 | VILL | 50853 | 0.02 | trans | ||
| rs2997307 | chr10 | 122561073 | VILL | 50853 | 0.03 | trans | ||
| rs897740 | chr19 | 6901558 | VILL | 50853 | 0.2 | trans | ||
| rs9826470 | 3 | 38019428 | VILL | ENSG00000136059.10 | 7.745E-7 | 0.02 | -10122 | gtex_brain_ba24 |
| rs7632911 | 3 | 38030899 | VILL | ENSG00000136059.10 | 2.407E-6 | 0.02 | 1349 | gtex_brain_ba24 |
| rs7644531 | 3 | 38030933 | VILL | ENSG00000136059.10 | 2.411E-6 | 0.02 | 1383 | gtex_brain_ba24 |
| rs9872804 | 3 | 38033462 | VILL | ENSG00000136059.10 | 2.606E-6 | 0.02 | 3912 | gtex_brain_ba24 |
| rs9823482 | 3 | 38036807 | VILL | ENSG00000136059.10 | 2.44E-6 | 0.02 | 7257 | gtex_brain_ba24 |
| rs9829569 | 3 | 38038329 | VILL | ENSG00000136059.10 | 2.45E-6 | 0.02 | 8779 | gtex_brain_ba24 |
| rs928807 | 3 | 38042182 | VILL | ENSG00000136059.10 | 2.467E-6 | 0.02 | 12632 | gtex_brain_ba24 |
| rs9311172 | 3 | 38044143 | VILL | ENSG00000136059.10 | 2.481E-6 | 0.02 | 14593 | gtex_brain_ba24 |
| rs9875248 | 3 | 38044592 | VILL | ENSG00000136059.10 | 2.484E-6 | 0.02 | 15042 | gtex_brain_ba24 |
| rs6793254 | 3 | 38044951 | VILL | ENSG00000136059.10 | 2.496E-6 | 0.02 | 15401 | gtex_brain_ba24 |
| rs6769106 | 3 | 38044969 | VILL | ENSG00000136059.10 | 2.496E-6 | 0.02 | 15419 | gtex_brain_ba24 |
| rs9311173 | 3 | 38045528 | VILL | ENSG00000136059.10 | 2.501E-6 | 0.02 | 15978 | gtex_brain_ba24 |
| rs9843506 | 3 | 38046023 | VILL | ENSG00000136059.10 | 2.505E-6 | 0.02 | 16473 | gtex_brain_ba24 |
| rs9311176 | 3 | 38047607 | VILL | ENSG00000136059.10 | 2.516E-6 | 0.02 | 18057 | gtex_brain_ba24 |
| rs6776745 | 3 | 38004661 | VILL | ENSG00000136059.10 | 3.742E-7 | 0 | -24889 | gtex_brain_putamen_basal |
| rs6801985 | 3 | 38005183 | VILL | ENSG00000136059.10 | 3.742E-7 | 0 | -24367 | gtex_brain_putamen_basal |
| rs1573291 | 3 | 38009749 | VILL | ENSG00000136059.10 | 3.713E-7 | 0 | -19801 | gtex_brain_putamen_basal |
| rs9826470 | 3 | 38019428 | VILL | ENSG00000136059.10 | 9.025E-10 | 0 | -10122 | gtex_brain_putamen_basal |
| rs11717282 | 3 | 38019924 | VILL | ENSG00000136059.10 | 5.642E-8 | 0 | -9626 | gtex_brain_putamen_basal |
| rs10095 | 3 | 38023553 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -5997 | gtex_brain_putamen_basal |
| rs11129775 | 3 | 38024408 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -5142 | gtex_brain_putamen_basal |
| rs11709029 | 3 | 38024420 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -5130 | gtex_brain_putamen_basal |
| rs60500371 | 3 | 38025750 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -3800 | gtex_brain_putamen_basal |
| rs73825449 | 3 | 38025991 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -3559 | gtex_brain_putamen_basal |
| rs17037286 | 3 | 38026387 | VILL | ENSG00000136059.10 | 5.178E-8 | 0 | -3163 | gtex_brain_putamen_basal |
| rs11711286 | 3 | 38026652 | VILL | ENSG00000136059.10 | 9.725E-9 | 0 | -2898 | gtex_brain_putamen_basal |
| rs11715511 | 3 | 38026914 | VILL | ENSG00000136059.10 | 9.144E-9 | 0 | -2636 | gtex_brain_putamen_basal |
| rs11719260 | 3 | 38027101 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -2449 | gtex_brain_putamen_basal |
| rs11712625 | 3 | 38028345 | VILL | ENSG00000136059.10 | 3.693E-9 | 0 | -1205 | gtex_brain_putamen_basal |
| rs11713558 | 3 | 38028614 | VILL | ENSG00000136059.10 | 2.941E-7 | 0 | -936 | gtex_brain_putamen_basal |
| rs11129776 | 3 | 38028675 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -875 | gtex_brain_putamen_basal |
| rs11922163 | 3 | 38028851 | VILL | ENSG00000136059.10 | 1.281E-8 | 0 | -699 | gtex_brain_putamen_basal |
| rs56264852 | 3 | 38030567 | VILL | ENSG00000136059.10 | 3.826E-7 | 0 | 1017 | gtex_brain_putamen_basal |
| rs11715770 | 3 | 38030747 | VILL | ENSG00000136059.10 | 3.127E-13 | 0 | 1197 | gtex_brain_putamen_basal |
| rs7632911 | 3 | 38030899 | VILL | ENSG00000136059.10 | 5.274E-15 | 0 | 1349 | gtex_brain_putamen_basal |
| rs7644531 | 3 | 38030933 | VILL | ENSG00000136059.10 | 5.256E-15 | 0 | 1383 | gtex_brain_putamen_basal |
| rs11715803 | 3 | 38031020 | VILL | ENSG00000136059.10 | 2.169E-8 | 0 | 1470 | gtex_brain_putamen_basal |
| rs7611147 | 3 | 38031372 | VILL | ENSG00000136059.10 | 1.215E-9 | 0 | 1822 | gtex_brain_putamen_basal |
| rs9872804 | 3 | 38033462 | VILL | ENSG00000136059.10 | 5.312E-15 | 0 | 3912 | gtex_brain_putamen_basal |
| rs7648182 | 3 | 38035508 | VILL | ENSG00000136059.10 | 1.195E-10 | 0 | 5958 | gtex_brain_putamen_basal |
| rs9823482 | 3 | 38036807 | VILL | ENSG00000136059.10 | 5.087E-15 | 0 | 7257 | gtex_brain_putamen_basal |
| rs9829569 | 3 | 38038329 | VILL | ENSG00000136059.10 | 4.972E-15 | 0 | 8779 | gtex_brain_putamen_basal |
| rs6809649 | 3 | 38038982 | VILL | ENSG00000136059.10 | 1.152E-10 | 0 | 9432 | gtex_brain_putamen_basal |
| rs6550514 | 3 | 38039187 | VILL | ENSG00000136059.10 | 1.149E-10 | 0 | 9637 | gtex_brain_putamen_basal |
| rs7641519 | 3 | 38039302 | VILL | ENSG00000136059.10 | 1.428E-10 | 0 | 9752 | gtex_brain_putamen_basal |
| rs928807 | 3 | 38042182 | VILL | ENSG00000136059.10 | 5.453E-15 | 0 | 12632 | gtex_brain_putamen_basal |
| rs9843296 | 3 | 38043904 | VILL | ENSG00000136059.10 | 2.45E-10 | 0 | 14354 | gtex_brain_putamen_basal |
| rs9311172 | 3 | 38044143 | VILL | ENSG00000136059.10 | 4.781E-15 | 0 | 14593 | gtex_brain_putamen_basal |
| rs146852558 | 3 | 38044327 | VILL | ENSG00000136059.10 | 8.138E-11 | 0 | 14777 | gtex_brain_putamen_basal |
| rs9875248 | 3 | 38044592 | VILL | ENSG00000136059.10 | 4.765E-15 | 0 | 15042 | gtex_brain_putamen_basal |
| rs6793254 | 3 | 38044951 | VILL | ENSG00000136059.10 | 4.75E-15 | 0 | 15401 | gtex_brain_putamen_basal |
| rs6769106 | 3 | 38044969 | VILL | ENSG00000136059.10 | 4.75E-15 | 0 | 15419 | gtex_brain_putamen_basal |
| rs35261592 | 3 | 38044996 | VILL | ENSG00000136059.10 | 7.151E-11 | 0 | 15446 | gtex_brain_putamen_basal |
| rs9311173 | 3 | 38045528 | VILL | ENSG00000136059.10 | 4.717E-15 | 0 | 15978 | gtex_brain_putamen_basal |
| rs9843506 | 3 | 38046023 | VILL | ENSG00000136059.10 | 4.703E-15 | 0 | 16473 | gtex_brain_putamen_basal |
| rs9311175 | 3 | 38047211 | VILL | ENSG00000136059.10 | 7.142E-11 | 0 | 17661 | gtex_brain_putamen_basal |
| rs9311176 | 3 | 38047607 | VILL | ENSG00000136059.10 | 4.657E-15 | 0 | 18057 | gtex_brain_putamen_basal |
| rs9816693 | 3 | 38047954 | VILL | ENSG00000136059.10 | 1.145E-10 | 0 | 18404 | gtex_brain_putamen_basal |
| rs57208122 | 3 | 38049029 | VILL | ENSG00000136059.10 | 2.397E-9 | 0 | 19479 | gtex_brain_putamen_basal |
| rs113213588 | 3 | 38050159 | VILL | ENSG00000136059.10 | 5.855E-8 | 0 | 20609 | gtex_brain_putamen_basal |
| rs9857730 | 3 | 38051941 | VILL | ENSG00000136059.10 | 8.71E-14 | 0 | 22391 | gtex_brain_putamen_basal |
| rs9842880 | 3 | 38052613 | VILL | ENSG00000136059.10 | 9.087E-10 | 0 | 23063 | gtex_brain_putamen_basal |
| rs9758003 | 3 | 38053206 | VILL | ENSG00000136059.10 | 8.926E-10 | 0 | 23656 | gtex_brain_putamen_basal |
| rs57948844 | 3 | 38053322 | VILL | ENSG00000136059.10 | 1.574E-9 | 0 | 23772 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
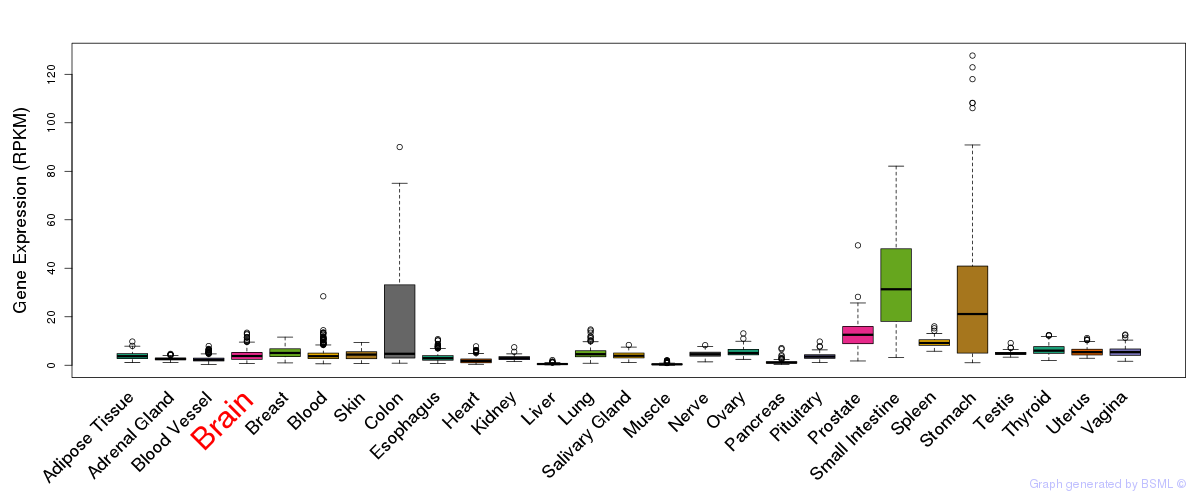
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0003779 | actin binding | IEA | - | |
| GO:0005509 | calcium ion binding | IEA | - | |
| GO:0005200 | structural constituent of cytoskeleton | TAS | 9179494 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0007010 | cytoskeleton organization | IEA | - | |
| GO:0051016 | barbed-end actin filament capping | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0015629 | actin cytoskeleton | TAS | 9179494 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| SENGUPTA NASOPHARYNGEAL CARCINOMA DN | 349 | 157 | All SZGR 2.0 genes in this pathway |
| BORCZUK MALIGNANT MESOTHELIOMA DN | 104 | 59 | All SZGR 2.0 genes in this pathway |
| VECCHI GASTRIC CANCER ADVANCED VS EARLY DN | 138 | 70 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY SERUM DEPRIVATION UP | 552 | 347 | All SZGR 2.0 genes in this pathway |
| HEIDENBLAD AMPLICON 12P11 12 DN | 30 | 16 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER NORMAL LIKE UP | 476 | 285 | All SZGR 2.0 genes in this pathway |
| GRADE COLON CANCER UP | 871 | 505 | All SZGR 2.0 genes in this pathway |
| CHYLA CBFA2T3 TARGETS UP | 387 | 225 | All SZGR 2.0 genes in this pathway |
| ACEVEDO FGFR1 TARGETS IN PROSTATE CANCER MODEL UP | 289 | 184 | All SZGR 2.0 genes in this pathway |
| LIM MAMMARY STEM CELL DN | 428 | 246 | All SZGR 2.0 genes in this pathway |