Gene Page: TOLLIP
Summary ?
| GeneID | 54472 |
| Symbol | TOLLIP |
| Synonyms | IL-1RAcPIP |
| Description | toll interacting protein |
| Reference | MIM:606277|HGNC:HGNC:16476|Ensembl:ENSG00000078902|HPRD:05887|Vega:OTTHUMG00000133333 |
| Gene type | protein-coding |
| Map location | 11p15.5 |
| Pascal p-value | 0.09 |
| Sherlock p-value | 0.443 |
| Fetal beta | -2.343 |
| DMG | 2 (# studies) |
| Support | G2Cdb.human_BAYES-COLLINS-HUMAN-PSD-CONSENSUS CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Nishioka_2013 | Genome-wide DNA methylation analysis | The authors investigated the methylation profiles of DNA in peripheral blood cells from 18 patients with first-episode schizophrenia (FESZ) and from 15 normal controls. | 5 |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 5 |
| Expression | Meta-analysis of gene expression | P value: 1.416 | |
| Network | Shortest path distance of core genes in the Human protein-protein interaction network | Contribution to shortest path in PPI network: 0.1454 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg17676428 | 11 | 1314042 | TOLLIP | 9.13E-6 | 0.474 | 0.013 | DMG:Wockner_2014 |
| cg11259409 | 11 | 1331124 | LOC255512;TOLLIP | 9.04E-5 | -0.225 | 0.027 | DMG:Wockner_2014 |
| cg03101580 | 11 | 1302893 | TOLLIP | 9.43E-5 | 0.608 | 0.027 | DMG:Wockner_2014 |
| cg26080154 | 11 | 1314228 | TOLLIP | 2.4E-4 | 0.38 | 0.037 | DMG:Wockner_2014 |
| cg13547299 | 11 | 1330611 | TOLLIP | -0.028 | 0.49 | DMG:Nishioka_2013 |
Section II. Transcriptome annotation
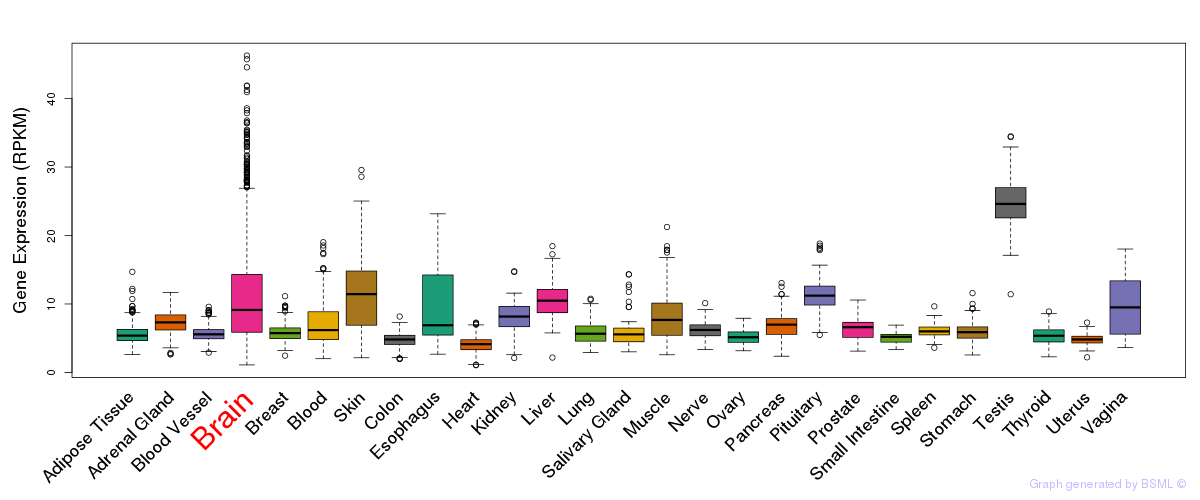
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0004871 | signal transducer activity | NAS | 11441107 | |
| GO:0005121 | Toll binding | NAS | 11441107 | |
| GO:0005515 | protein binding | IPI | 10854325 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0007267 | cell-cell signaling | TAS | 10854325 | |
| GO:0007242 | intracellular signaling cascade | NAS | 10854325 | |
| GO:0006955 | immune response | IEA | - | |
| GO:0006954 | inflammatory response | NAS | 10854325 | |
| GO:0016310 | phosphorylation | IDA | 1085432 | |
| GO:0045321 | leukocyte activation | NAS | 11441107 | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0005737 | cytoplasm | IC | 11441107 | |
| GO:0045323 | interleukin-1 receptor complex | NAS | 10854325 | |
| GO:0045092 | interleukin-18 receptor complex | NAS | 10854325 |
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| DAZAP2 | KIAA0058 | MGC14319 | MGC766 | PRTB | DAZ associated protein 2 | Two-hybrid | BioGRID | 16189514 |
| IL1R1 | CD121A | D2S1473 | IL-1R-alpha | IL1R | IL1RA | P80 | interleukin 1 receptor, type I | Affinity Capture-Western | BioGRID | 10854325 |
| IL1RAP | C3orf13 | FLJ37788 | IL-1RAcP | IL1R3 | interleukin 1 receptor accessory protein | - | HPRD,BioGRID | 10854325 |
| IRAK1 | IRAK | pelle | interleukin-1 receptor-associated kinase 1 | - | HPRD,BioGRID | 10854325 |
| LDB1 | CLIM2 | NLI | LIM domain binding 1 | Two-hybrid | BioGRID | 16189514 |
| RBPMS | HERMES | RNA binding protein with multiple splicing | Two-hybrid | BioGRID | 16189514 |
| RHOXF2 | PEPP-2 | PEPP2 | THG1 | Rhox homeobox family, member 2 | Two-hybrid | BioGRID | 16189514 |
| SEPT1 | DIFF6 | LARP | MGC20394 | PNUTL3 | SEP1 | septin 1 | Two-hybrid | BioGRID | 16189514 |
| SETDB1 | ESET | KG1T | KIAA0067 | KMT1E | SET domain, bifurcated 1 | Two-hybrid | BioGRID | 16169070 |
| TLR2 | CD282 | TIL4 | toll-like receptor 2 | - | HPRD,BioGRID | 11751856 |
| TLR4 | ARMD10 | CD284 | TOLL | hToll | toll-like receptor 4 | - | HPRD,BioGRID | 11751856 |
| TOLLIP | FLJ33531 | IL-1RAcPIP | toll interacting protein | Reconstituted Complex | BioGRID | 11751856 |
| TOM1 | FLJ33404 | target of myb1 (chicken) | - | HPRD,BioGRID | 14563850 |
| XRN2 | - | 5'-3' exoribonuclease 2 | Two-hybrid | BioGRID | 15231747 |
| ZBTB16 | PLZF | ZNF145 | zinc finger and BTB domain containing 16 | Two-hybrid | BioGRID | 16169070 |
| ZHX1 | - | zinc fingers and homeoboxes 1 | Two-hybrid | BioGRID | 16169070 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG TOLL LIKE RECEPTOR SIGNALING PATHWAY | 102 | 88 | All SZGR 2.0 genes in this pathway |
| BIOCARTA GSK3 PATHWAY | 27 | 26 | All SZGR 2.0 genes in this pathway |
| BIOCARTA IL1R PATHWAY | 33 | 29 | All SZGR 2.0 genes in this pathway |
| BIOCARTA TOLL PATHWAY | 37 | 31 | All SZGR 2.0 genes in this pathway |
| PID IL1 PATHWAY | 34 | 28 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALING BY ILS | 107 | 86 | All SZGR 2.0 genes in this pathway |
| REACTOME IL1 SIGNALING | 39 | 30 | All SZGR 2.0 genes in this pathway |
| REACTOME IMMUNE SYSTEM | 933 | 616 | All SZGR 2.0 genes in this pathway |
| REACTOME CYTOKINE SIGNALING IN IMMUNE SYSTEM | 270 | 204 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN MLL SIGNATURE 1 UP | 380 | 236 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN MLL SIGNATURE 2 UP | 418 | 263 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| ENK UV RESPONSE EPIDERMIS UP | 293 | 179 | All SZGR 2.0 genes in this pathway |
| CREIGHTON AKT1 SIGNALING VIA MTOR UP | 34 | 22 | All SZGR 2.0 genes in this pathway |
| PEREZ TP53 TARGETS | 1174 | 695 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| LABBE WNT3A TARGETS DN | 97 | 53 | All SZGR 2.0 genes in this pathway |
| LABBE TARGETS OF TGFB1 AND WNT3A DN | 108 | 68 | All SZGR 2.0 genes in this pathway |
| NAKAMURA METASTASIS MODEL UP | 45 | 26 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE LATE | 1137 | 655 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS UP | 682 | 433 | All SZGR 2.0 genes in this pathway |
| KUMAR PATHOGEN LOAD BY MACROPHAGES | 275 | 155 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |