Gene Page: PRMT7
Summary ?
| GeneID | 54496 |
| Symbol | PRMT7 |
| Synonyms | - |
| Description | protein arginine methyltransferase 7 |
| Reference | MIM:610087|HGNC:HGNC:25557|Ensembl:ENSG00000132600|HPRD:15174|Vega:OTTHUMG00000137556 |
| Gene type | protein-coding |
| Map location | 16q22.1 |
| Pascal p-value | 3.168E-4 |
| Sherlock p-value | 0.653 |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus Nucleus accumbens basal ganglia Putamen basal ganglia |
| Support | Ascano FMRP targets Chromatin Remodeling Genes |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs78572164 | 16 | 67931112 | PRMT7 | ENSG00000132600.12 | 7.39353E-6 | 0.04 | -413765 | gtex_brain_ba24 |
| rs10719112 | 16 | 68187782 | PRMT7 | ENSG00000132600.12 | 6.45841E-6 | 0.01 | -157095 | gtex_brain_putamen_basal |
| rs7206885 | 16 | 68323088 | PRMT7 | ENSG00000132600.12 | 9.72539E-6 | 0.01 | -21789 | gtex_brain_putamen_basal |
| rs10775303 | 16 | 68360914 | PRMT7 | ENSG00000132600.12 | 3.69743E-6 | 0.01 | 16037 | gtex_brain_putamen_basal |
| rs118081518 | 16 | 68385425 | PRMT7 | ENSG00000132600.12 | 1.09764E-5 | 0.01 | 40548 | gtex_brain_putamen_basal |
| rs1465471 | 16 | 68396377 | PRMT7 | ENSG00000132600.12 | 1.06534E-5 | 0.01 | 51500 | gtex_brain_putamen_basal |
| rs11646700 | 16 | 68421668 | PRMT7 | ENSG00000132600.12 | 8.16803E-7 | 0.01 | 76791 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
General gene expression (GTEx)
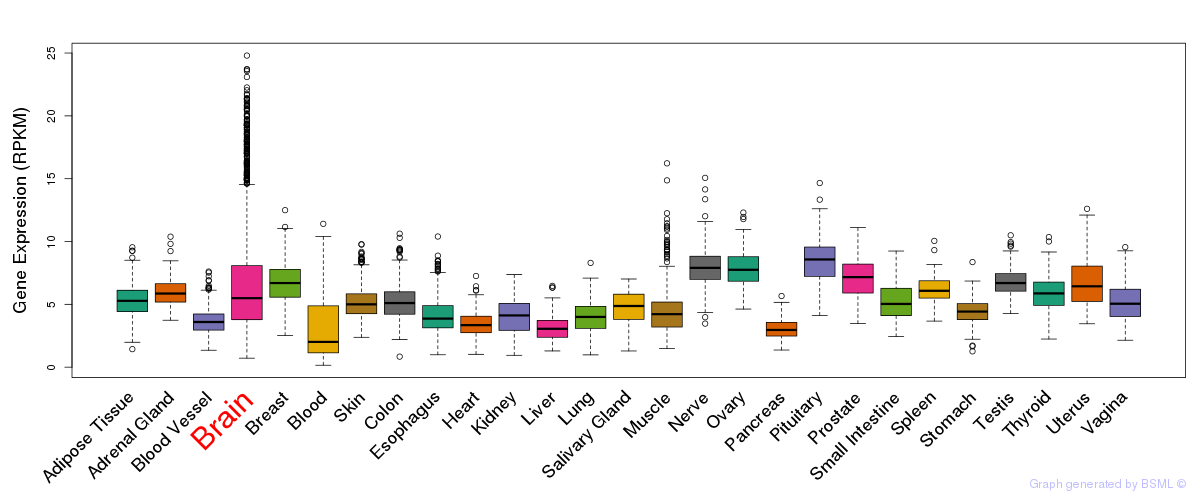
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| CASORELLI ACUTE PROMYELOCYTIC LEUKEMIA DN | 663 | 425 | All SZGR 2.0 genes in this pathway |
| GAL LEUKEMIC STEM CELL DN | 244 | 153 | All SZGR 2.0 genes in this pathway |
| GARGALOVIC RESPONSE TO OXIDIZED PHOSPHOLIPIDS TURQUOISE DN | 53 | 25 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS DN | 459 | 276 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GAUSSMANN MLL AF4 FUSION TARGETS G UP | 238 | 135 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| MANALO HYPOXIA DN | 289 | 166 | All SZGR 2.0 genes in this pathway |
| CUI TCF21 TARGETS 2 UP | 428 | 266 | All SZGR 2.0 genes in this pathway |
| SATO SILENCED BY METHYLATION IN PANCREATIC CANCER 1 | 419 | 273 | All SZGR 2.0 genes in this pathway |
| AMUNDSON POOR SURVIVAL AFTER GAMMA RADIATION 2G | 171 | 96 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER ERBB2 UP | 147 | 83 | All SZGR 2.0 genes in this pathway |
| QI PLASMACYTOMA DN | 100 | 63 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS DN | 882 | 538 | All SZGR 2.0 genes in this pathway |