Gene Page: ATR
Summary ?
| GeneID | 545 |
| Symbol | ATR |
| Synonyms | FCTCS|FRP1|MEC1|SCKL|SCKL1 |
| Description | ATR serine/threonine kinase |
| Reference | MIM:601215|HGNC:HGNC:882|Ensembl:ENSG00000175054|HPRD:08369|Vega:OTTHUMG00000159234 |
| Gene type | protein-coding |
| Map location | 3q23 |
| Pascal p-value | 0.799 |
| Fetal beta | -0.454 |
| eGene | Myers' cis & trans |
| Support | Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs4491879 | chr3 | 189373908 | ATR | 545 | 0.05 | trans | ||
| rs4505678 | chr3 | 189374356 | ATR | 545 | 0.06 | trans | ||
| rs16894557 | chr6 | 28999825 | ATR | 545 | 0 | trans |
Section II. Transcriptome annotation
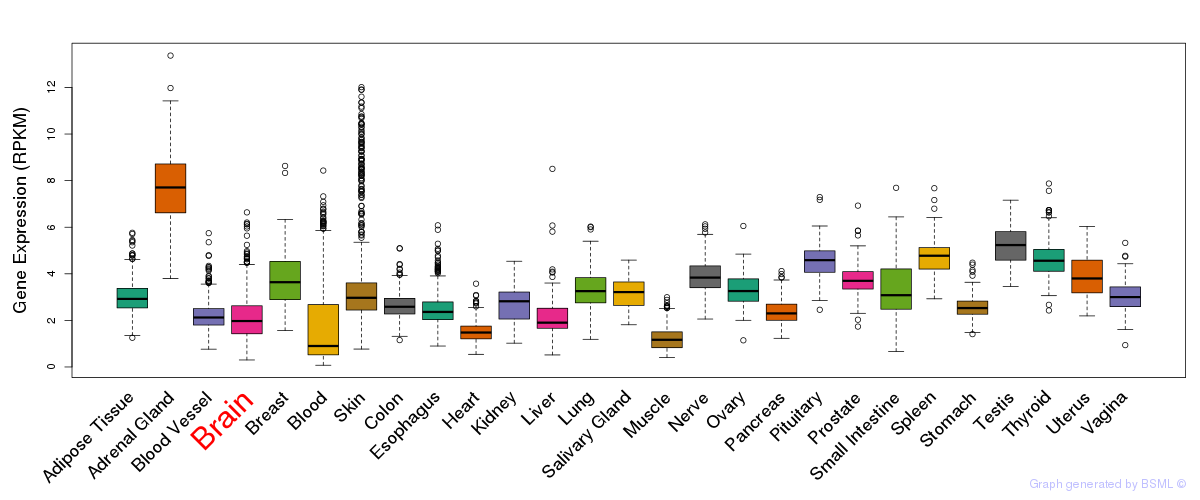
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| ABL1 | ABL | JTK7 | bcr/abl | c-ABL | p150 | v-abl | c-abl oncogene 1, receptor tyrosine kinase | Protein-peptide | BioGRID | 10608806 |
| AP1B1 | ADTB1 | AP105A | BAM22 | CLAPB2 | adaptor-related protein complex 1, beta 1 subunit | Protein-peptide | BioGRID | 10608806 |
| AP2A2 | ADTAB | CLAPA2 | HIP9 | HYPJ | adaptor-related protein complex 2, alpha 2 subunit | Protein-peptide | BioGRID | 10608806 |
| ARHGEF1 | GEF1 | LBCL2 | LSC | P115-RHOGEF | SUB1.5 | Rho guanine nucleotide exchange factor (GEF) 1 | Protein-peptide | BioGRID | 10608806 |
| ATM | AT1 | ATA | ATC | ATD | ATDC | ATE | DKFZp781A0353 | MGC74674 | TEL1 | TELO1 | ataxia telangiectasia mutated | Protein-peptide | BioGRID | 10608806 |
| ATMIN | ASCIZ | DKFZp779K1455 | FLJ76795 | KIAA0431 | ZNF822 | ATM interactor | ATR interacts with ASCIZ. This interaction was modeled on a demonstrated interaction between yeast ATR and human ASCIZ. | BIND | 15933716 |
| ATR | FRP1 | MEC1 | SCKL | SCKL1 | ataxia telangiectasia and Rad3 related | Protein-peptide | BioGRID | 10608806 |
| ATRIP | DKFZp762J2115 | FLJ12343 | MGC20625 | MGC21482 | MGC26740 | ATR interacting protein | Affinity Capture-MS Affinity Capture-Western Biochemical Activity | BioGRID | 11721054 |14657349 |15210935 |
| BRCA1 | BRCAI | BRCC1 | IRIS | PSCP | RNF53 | breast cancer 1, early onset | BRCA1 physically and functionally interacts with ATR during genotoxic stress | BIND | 11016625 |11278964 |
| BRCA1 | BRCAI | BRCC1 | IRIS | PSCP | RNF53 | breast cancer 1, early onset | Affinity Capture-Western Protein-peptide Reconstituted Complex | BioGRID | 10608806 |11016625 |11114888 |11278964 |
| BRCA2 | BRCC2 | FACD | FAD | FAD1 | FANCB | FANCD | FANCD1 | breast cancer 2, early onset | Protein-peptide | BioGRID | 10608806 |
| CHD4 | DKFZp686E06161 | Mi-2b | Mi2-BETA | chromodomain helicase DNA binding protein 4 | - | HPRD,BioGRID | 10545197 |
| CHEK1 | CHK1 | CHK1 checkpoint homolog (S. pombe) | Protein-peptide | BioGRID | 10608806 |
| CHEK1 | CHK1 | CHK1 checkpoint homolog (S. pombe) | ATR interacts with CHEK1 (CHK1). | BIND | 15775976 |
| CLSPN | CLASPIN | MGC131612 | MGC131613 | MGC131615 | claspin homolog (Xenopus laevis) | - | HPRD,BioGRID | 12766152 |
| E4F1 | E4F | MGC99614 | E4F transcription factor 1 | Protein-peptide | BioGRID | 10608806 |
| EEF1E1 | AIMP3 | P18 | eukaryotic translation elongation factor 1 epsilon 1 | p18 interacts with ATR. | BIND | 15680327 |
| HDAC1 | DKFZp686H12203 | GON-10 | HD1 | RPD3 | RPD3L1 | histone deacetylase 1 | Affinity Capture-Western | BioGRID | 10545197 |
| HDAC2 | RPD3 | YAF1 | histone deacetylase 2 | Affinity Capture-Western Co-purification | BioGRID | 10545197 |
| LIG4 | - | ligase IV, DNA, ATP-dependent | Protein-peptide | BioGRID | 10608806 |
| MCM2 | BM28 | CCNL1 | CDCL1 | D3S3194 | KIAA0030 | MGC10606 | MITOTIN | cdc19 | minichromosome maintenance complex component 2 | Biochemical Activity | BioGRID | 15210935 |
| MCM7 | CDABP0042 | CDC47 | MCM2 | P1.1-MCM3 | P1CDC47 | P85MCM | PNAS-146 | minichromosome maintenance complex component 7 | Affinity Capture-Western | BioGRID | 15210935 |
| MRE11A | ATLD | HNGS1 | MRE11 | MRE11B | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | Protein-peptide | BioGRID | 10608806 |
| MSH2 | COCA1 | FCC1 | HNPCC | HNPCC1 | LCFS2 | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | - | HPRD,BioGRID | 14657349 |
| MTA1 | - | metastasis associated 1 | Affinity Capture-Western | BioGRID | 10545197 |
| MTA2 | DKFZp686F2281 | MTA1L1 | PID | metastasis associated 1 family, member 2 | Affinity Capture-Western | BioGRID | 10545197 |
| NBN | AT-V1 | AT-V2 | ATV | FLJ10155 | MGC87362 | NBS | NBS1 | P95 | nibrin | Protein-peptide | BioGRID | 10608806 |
| NBN | AT-V1 | AT-V2 | ATV | FLJ10155 | MGC87362 | NBS | NBS1 | P95 | nibrin | NBS1 interacts with ATR. | BIND | 15758953 |
| PA2G4 | EBP1 | HG4-1 | p38-2G4 | proliferation-associated 2G4, 38kDa | Protein-peptide | BioGRID | 10608806 |
| PIK3CA | MGC142161 | MGC142163 | PI3K | p110-alpha | phosphoinositide-3-kinase, catalytic, alpha polypeptide | Protein-peptide | BioGRID | 10608806 |
| PML | MYL | PP8675 | RNF71 | TRIM19 | promyelocytic leukemia | ATR phosphorylates PML. | BIND | 15195100 |
| POLD1 | CDC2 | POLD | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | Protein-peptide | BioGRID | 10608806 |
| PPP2R2A | B55-ALPHA | B55A | FLJ26613 | MGC52248 | PR52A | PR55A | protein phosphatase 2 (formerly 2A), regulatory subunit B, alpha isoform | Protein-peptide | BioGRID | 10608806 |
| PTS | FLJ97081 | PTPS | 6-pyruvoyltetrahydropterin synthase | Protein-peptide | BioGRID | 10608806 |
| RAD17 | CCYC | FLJ41520 | HRAD17 | R24L | RAD17SP | RAD24 | RAD17 homolog (S. pombe) | Affinity Capture-Western Biochemical Activity Protein-peptide | BioGRID | 10608806 |11418864 |
| RHEB | MGC111559 | RHEB2 | Ras homolog enriched in brain | ATR interacts with Rheb. | BIND | 15854902 |
| RHEB | MGC111559 | RHEB2 | Ras homolog enriched in brain | Affinity Capture-Western Reconstituted Complex | BioGRID | 15854902 |
| RPA1 | HSSB | REPA1 | RF-A | RP-A | RPA70 | replication protein A1, 70kDa | Protein-peptide | BioGRID | 10608806 |
| TP53 | FLJ92943 | LFS1 | TRP53 | p53 | tumor protein p53 | ATR interacts with TP53 (p53). | BIND | 15775976 |
| TP53 | FLJ92943 | LFS1 | TRP53 | p53 | tumor protein p53 | Protein-peptide Reconstituted Complex | BioGRID | 10608806 |15159397 |
| TREX1 | AGS1 | AGS5 | CRV | DKFZp434J0310 | DRN3 | HERNS | three prime repair exonuclease 1 | - | HPRD | 11721054 |
| WRN | DKFZp686C2056 | RECQ3 | RECQL2 | RECQL3 | Werner syndrome | Protein-peptide | BioGRID | 10608806 |
| XRCC5 | FLJ39089 | KARP-1 | KARP1 | KU80 | KUB2 | Ku86 | NFIV | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining) | XRCC5 (Ku80) interacts with ATR. | BIND | 15758953 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG CELL CYCLE | 128 | 84 | All SZGR 2.0 genes in this pathway |
| KEGG P53 SIGNALING PATHWAY | 69 | 45 | All SZGR 2.0 genes in this pathway |
| BIOCARTA G1 PATHWAY | 28 | 21 | All SZGR 2.0 genes in this pathway |
| BIOCARTA G2 PATHWAY | 24 | 20 | All SZGR 2.0 genes in this pathway |
| BIOCARTA ATRBRCA PATHWAY | 21 | 12 | All SZGR 2.0 genes in this pathway |
| PID FANCONI PATHWAY | 47 | 28 | All SZGR 2.0 genes in this pathway |
| PID ATR PATHWAY | 39 | 25 | All SZGR 2.0 genes in this pathway |
| PID TCPTP PATHWAY | 43 | 33 | All SZGR 2.0 genes in this pathway |
| PID CIRCADIAN PATHWAY | 16 | 15 | All SZGR 2.0 genes in this pathway |
| PID BARD1 PATHWAY | 29 | 19 | All SZGR 2.0 genes in this pathway |
| PID P53 REGULATION PATHWAY | 59 | 50 | All SZGR 2.0 genes in this pathway |
| REACTOME MEIOSIS | 116 | 81 | All SZGR 2.0 genes in this pathway |
| REACTOME CELL CYCLE | 421 | 253 | All SZGR 2.0 genes in this pathway |
| REACTOME CELL CYCLE CHECKPOINTS | 124 | 70 | All SZGR 2.0 genes in this pathway |
| REACTOME FANCONI ANEMIA PATHWAY | 25 | 11 | All SZGR 2.0 genes in this pathway |
| REACTOME DNA REPAIR | 112 | 59 | All SZGR 2.0 genes in this pathway |
| REACTOME CHROMOSOME MAINTENANCE | 122 | 80 | All SZGR 2.0 genes in this pathway |
| REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | 38 | 18 | All SZGR 2.0 genes in this pathway |
| REACTOME MEIOTIC SYNAPSIS | 73 | 57 | All SZGR 2.0 genes in this pathway |
| REACTOME G2 M CHECKPOINTS | 45 | 23 | All SZGR 2.0 genes in this pathway |
| REACTOME G2 M DNA DAMAGE CHECKPOINT | 12 | 8 | All SZGR 2.0 genes in this pathway |
| LIU SOX4 TARGETS UP | 137 | 94 | All SZGR 2.0 genes in this pathway |
| GARY CD5 TARGETS DN | 431 | 263 | All SZGR 2.0 genes in this pathway |
| HOEBEKE LYMPHOID STEM CELL UP | 95 | 64 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA DN | 1375 | 806 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED UP | 633 | 376 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA ANAPLASTIC UP | 722 | 443 | All SZGR 2.0 genes in this pathway |
| ENK UV RESPONSE KERATINOCYTE DN | 485 | 334 | All SZGR 2.0 genes in this pathway |
| HAMAI APOPTOSIS VIA TRAIL UP | 584 | 356 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 DN | 855 | 609 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 COMMON DN | 483 | 336 | All SZGR 2.0 genes in this pathway |
| PUJANA BREAST CANCER LIT INT NETWORK | 101 | 73 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA1 PCC NETWORK | 1652 | 1023 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA2 PCC NETWORK | 423 | 265 | All SZGR 2.0 genes in this pathway |
| PUJANA CHEK2 PCC NETWORK | 779 | 480 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| KAUFFMANN DNA REPAIR GENES | 230 | 137 | All SZGR 2.0 genes in this pathway |
| PENG GLUTAMINE DEPRIVATION DN | 337 | 230 | All SZGR 2.0 genes in this pathway |
| FLECHNER PBL KIDNEY TRANSPLANT REJECTED VS OK UP | 63 | 48 | All SZGR 2.0 genes in this pathway |
| HADDAD B LYMPHOCYTE PROGENITOR | 293 | 193 | All SZGR 2.0 genes in this pathway |
| GALE APL WITH FLT3 MUTATED UP | 56 | 35 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE DN | 1237 | 837 | All SZGR 2.0 genes in this pathway |
| WEIGEL OXIDATIVE STRESS BY HNE AND TBH | 60 | 42 | All SZGR 2.0 genes in this pathway |
| DEBIASI APOPTOSIS BY REOVIRUS INFECTION DN | 287 | 208 | All SZGR 2.0 genes in this pathway |
| LY AGING OLD DN | 56 | 35 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR DN | 1011 | 592 | All SZGR 2.0 genes in this pathway |
| ZHANG BREAST CANCER PROGENITORS UP | 425 | 253 | All SZGR 2.0 genes in this pathway |
| EHLERS ANEUPLOIDY UP | 41 | 29 | All SZGR 2.0 genes in this pathway |
| FIRESTEIN PROLIFERATION | 175 | 125 | All SZGR 2.0 genes in this pathway |
| MILI PSEUDOPODIA HAPTOTAXIS UP | 518 | 299 | All SZGR 2.0 genes in this pathway |
| FONTAINE FOLLICULAR THYROID ADENOMA DN | 68 | 45 | All SZGR 2.0 genes in this pathway |
| FONTAINE THYROID TUMOR UNCERTAIN MALIGNANCY DN | 26 | 14 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 3 DN | 918 | 550 | All SZGR 2.0 genes in this pathway |