Gene Page: PGM2
Summary ?
| GeneID | 55276 |
| Symbol | PGM2 |
| Synonyms | MSTP006 |
| Description | phosphoglucomutase 2 |
| Reference | MIM:172000|HGNC:HGNC:8906|Ensembl:ENSG00000169299|HPRD:11761|Vega:OTTHUMG00000097813 |
| Gene type | protein-coding |
| Map location | 4p14 |
| Pascal p-value | 0.01 |
| Fetal beta | 0.69 |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cortex Frontal Cortex BA9 Hippocampus Nucleus accumbens basal ganglia Putamen basal ganglia |
| Support | CELL METABOLISM |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs2995966 | 4 | 37780576 | PGM2 | ENSG00000169299.9 | 5.712E-7 | 0.02 | -47679 | gtex_brain_ba24 |
| rs7659570 | 4 | 37781158 | PGM2 | ENSG00000169299.9 | 5.712E-7 | 0.02 | -47097 | gtex_brain_ba24 |
| rs12512677 | 4 | 37817932 | PGM2 | ENSG00000169299.9 | 1.556E-6 | 0.02 | -10323 | gtex_brain_ba24 |
| rs1115683 | 4 | 37820396 | PGM2 | ENSG00000169299.9 | 1.556E-6 | 0.02 | -7859 | gtex_brain_ba24 |
| rs35282905 | 4 | 37821847 | PGM2 | ENSG00000169299.9 | 1.524E-6 | 0.02 | -6408 | gtex_brain_ba24 |
| rs11721535 | 4 | 37824325 | PGM2 | ENSG00000169299.9 | 1.557E-6 | 0.02 | -3930 | gtex_brain_ba24 |
| rs3775798 | 4 | 37824855 | PGM2 | ENSG00000169299.9 | 1.557E-6 | 0.02 | -3400 | gtex_brain_ba24 |
| rs3756142 | 4 | 37825550 | PGM2 | ENSG00000169299.9 | 1.557E-6 | 0.02 | -2705 | gtex_brain_ba24 |
| rs4832957 | 4 | 37825940 | PGM2 | ENSG00000169299.9 | 1.557E-6 | 0.02 | -2315 | gtex_brain_ba24 |
| rs13122762 | 4 | 37826536 | PGM2 | ENSG00000169299.9 | 1.557E-6 | 0.02 | -1719 | gtex_brain_ba24 |
| rs59397780 | 4 | 37827578 | PGM2 | ENSG00000169299.9 | 1.538E-6 | 0.02 | -677 | gtex_brain_ba24 |
| rs11096905 | 4 | 37800240 | PGM2 | ENSG00000169299.9 | 5.928E-9 | 0 | -28015 | gtex_brain_putamen_basal |
| rs11938667 | 4 | 37800525 | PGM2 | ENSG00000169299.9 | 5.928E-9 | 0 | -27730 | gtex_brain_putamen_basal |
| rs66475274 | 4 | 37800876 | PGM2 | ENSG00000169299.9 | 5.928E-9 | 0 | -27379 | gtex_brain_putamen_basal |
| rs17578457 | 4 | 37800972 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -27283 | gtex_brain_putamen_basal |
| rs66540038 | 4 | 37801051 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -27204 | gtex_brain_putamen_basal |
| rs17606268 | 4 | 37801163 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -27092 | gtex_brain_putamen_basal |
| rs11737776 | 4 | 37801348 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -26907 | gtex_brain_putamen_basal |
| rs398107204 | 4 | 37801666 | PGM2 | ENSG00000169299.9 | 5.128E-10 | 0 | -26589 | gtex_brain_putamen_basal |
| rs6832381 | 4 | 37801922 | PGM2 | ENSG00000169299.9 | 2.173E-9 | 0 | -26333 | gtex_brain_putamen_basal |
| rs6849324 | 4 | 37801952 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -26303 | gtex_brain_putamen_basal |
| rs11938059 | 4 | 37802139 | PGM2 | ENSG00000169299.9 | 1.188E-8 | 0 | -26116 | gtex_brain_putamen_basal |
| rs6849750 | 4 | 37802141 | PGM2 | ENSG00000169299.9 | 1.187E-8 | 0 | -26114 | gtex_brain_putamen_basal |
| rs112691053 | 4 | 37802449 | PGM2 | ENSG00000169299.9 | 5.889E-9 | 0 | -25806 | gtex_brain_putamen_basal |
| rs76792647 | 4 | 37802625 | PGM2 | ENSG00000169299.9 | 5.949E-9 | 0 | -25630 | gtex_brain_putamen_basal |
| rs13103553 | 4 | 37802850 | PGM2 | ENSG00000169299.9 | 5.908E-9 | 0 | -25405 | gtex_brain_putamen_basal |
| rs34371555 | 4 | 37802906 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -25349 | gtex_brain_putamen_basal |
| rs34274238 | 4 | 37802988 | PGM2 | ENSG00000169299.9 | 2.147E-9 | 0 | -25267 | gtex_brain_putamen_basal |
| rs34856096 | 4 | 37803052 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -25203 | gtex_brain_putamen_basal |
| rs13110103 | 4 | 37803233 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -25022 | gtex_brain_putamen_basal |
| rs1425322 | 4 | 37803423 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -24832 | gtex_brain_putamen_basal |
| rs1834340 | 4 | 37803601 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -24654 | gtex_brain_putamen_basal |
| rs1863283 | 4 | 37803619 | PGM2 | ENSG00000169299.9 | 9.43E-9 | 0 | -24636 | gtex_brain_putamen_basal |
| rs5857588 | 4 | 37803623 | PGM2 | ENSG00000169299.9 | 9.373E-9 | 0 | -24632 | gtex_brain_putamen_basal |
| rs883270 | 4 | 37803934 | PGM2 | ENSG00000169299.9 | 2.172E-9 | 0 | -24321 | gtex_brain_putamen_basal |
| rs35923894 | 4 | 37804347 | PGM2 | ENSG00000169299.9 | 2.29E-6 | 0 | -23908 | gtex_brain_putamen_basal |
| rs1425321 | 4 | 37804819 | PGM2 | ENSG00000169299.9 | 9.43E-9 | 0 | -23436 | gtex_brain_putamen_basal |
| rs56838577 | 4 | 37807521 | PGM2 | ENSG00000169299.9 | 4.54E-9 | 0 | -20734 | gtex_brain_putamen_basal |
| rs12499691 | 4 | 37807681 | PGM2 | ENSG00000169299.9 | 5.339E-9 | 0 | -20574 | gtex_brain_putamen_basal |
| rs4832955 | 4 | 37809773 | PGM2 | ENSG00000169299.9 | 6.589E-9 | 0 | -18482 | gtex_brain_putamen_basal |
| rs1834345 | 4 | 37810182 | PGM2 | ENSG00000169299.9 | 1.644E-6 | 0 | -18073 | gtex_brain_putamen_basal |
| rs4832737 | 4 | 37812202 | PGM2 | ENSG00000169299.9 | 5.942E-9 | 0 | -16053 | gtex_brain_putamen_basal |
| rs4832738 | 4 | 37814993 | PGM2 | ENSG00000169299.9 | 5.477E-9 | 0 | -13262 | gtex_brain_putamen_basal |
| rs7691795 | 4 | 37816597 | PGM2 | ENSG00000169299.9 | 5.314E-9 | 0 | -11658 | gtex_brain_putamen_basal |
| rs4832956 | 4 | 37816879 | PGM2 | ENSG00000169299.9 | 5.294E-9 | 0 | -11376 | gtex_brain_putamen_basal |
| rs12512677 | 4 | 37817932 | PGM2 | ENSG00000169299.9 | 3.319E-9 | 0 | -10323 | gtex_brain_putamen_basal |
| rs1115683 | 4 | 37820396 | PGM2 | ENSG00000169299.9 | 3.417E-9 | 0 | -7859 | gtex_brain_putamen_basal |
| rs2911917 | 4 | 37820533 | PGM2 | ENSG00000169299.9 | 1.495E-6 | 0 | -7722 | gtex_brain_putamen_basal |
| rs34738612 | 4 | 37821794 | PGM2 | ENSG00000169299.9 | 1.244E-8 | 0 | -6461 | gtex_brain_putamen_basal |
| rs35282905 | 4 | 37821847 | PGM2 | ENSG00000169299.9 | 4.401E-10 | 0 | -6408 | gtex_brain_putamen_basal |
| rs2911910 | 4 | 37822075 | PGM2 | ENSG00000169299.9 | 1.391E-6 | 0 | -6180 | gtex_brain_putamen_basal |
| rs11721535 | 4 | 37824325 | PGM2 | ENSG00000169299.9 | 3.401E-9 | 0 | -3930 | gtex_brain_putamen_basal |
| rs3775798 | 4 | 37824855 | PGM2 | ENSG00000169299.9 | 3.598E-9 | 0 | -3400 | gtex_brain_putamen_basal |
| rs3756142 | 4 | 37825550 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | -2705 | gtex_brain_putamen_basal |
| rs4832957 | 4 | 37825940 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | -2315 | gtex_brain_putamen_basal |
| rs13122762 | 4 | 37826536 | PGM2 | ENSG00000169299.9 | 3.409E-9 | 0 | -1719 | gtex_brain_putamen_basal |
| rs59397780 | 4 | 37827578 | PGM2 | ENSG00000169299.9 | 3.413E-9 | 0 | -677 | gtex_brain_putamen_basal |
| rs2059483 | 4 | 37828186 | PGM2 | ENSG00000169299.9 | 4.615E-8 | 0 | -69 | gtex_brain_putamen_basal |
| rs3775796 | 4 | 37830358 | PGM2 | ENSG00000169299.9 | 3.261E-9 | 0 | 2103 | gtex_brain_putamen_basal |
| rs11733779 | 4 | 37833060 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | 4805 | gtex_brain_putamen_basal |
| rs3775793 | 4 | 37840606 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | 12351 | gtex_brain_putamen_basal |
| rs3775792 | 4 | 37840653 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | 12398 | gtex_brain_putamen_basal |
| rs12501402 | 4 | 37844504 | PGM2 | ENSG00000169299.9 | 3.447E-9 | 0 | 16249 | gtex_brain_putamen_basal |
| rs12509391 | 4 | 37844604 | PGM2 | ENSG00000169299.9 | 3.447E-9 | 0 | 16349 | gtex_brain_putamen_basal |
| rs398107206 | 4 | 37845833 | PGM2 | ENSG00000169299.9 | 9.457E-9 | 0 | 17578 | gtex_brain_putamen_basal |
| rs13128974 | 4 | 37848085 | PGM2 | ENSG00000169299.9 | 3.44E-9 | 0 | 19830 | gtex_brain_putamen_basal |
| rs35704131 | 4 | 37850312 | PGM2 | ENSG00000169299.9 | 3.448E-9 | 0 | 22057 | gtex_brain_putamen_basal |
| rs4832959 | 4 | 37852564 | PGM2 | ENSG00000169299.9 | 3.155E-9 | 0 | 24309 | gtex_brain_putamen_basal |
| rs6820161 | 4 | 37853172 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 24917 | gtex_brain_putamen_basal |
| rs6838455 | 4 | 37853243 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 24988 | gtex_brain_putamen_basal |
| rs6820375 | 4 | 37853268 | PGM2 | ENSG00000169299.9 | 1.046E-8 | 0 | 25013 | gtex_brain_putamen_basal |
| rs6820384 | 4 | 37853286 | PGM2 | ENSG00000169299.9 | 1.046E-8 | 0 | 25031 | gtex_brain_putamen_basal |
| rs13146036 | 4 | 37853375 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 25120 | gtex_brain_putamen_basal |
| rs13114137 | 4 | 37853392 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 25137 | gtex_brain_putamen_basal |
| rs13140704 | 4 | 37853407 | PGM2 | ENSG00000169299.9 | 1.033E-8 | 0 | 25152 | gtex_brain_putamen_basal |
| rs35619511 | 4 | 37853557 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 25302 | gtex_brain_putamen_basal |
| rs6842833 | 4 | 37853661 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 25406 | gtex_brain_putamen_basal |
| rs4832960 | 4 | 37854182 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 25927 | gtex_brain_putamen_basal |
| rs4832961 | 4 | 37854327 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 26072 | gtex_brain_putamen_basal |
| rs6531582 | 4 | 37854607 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 26352 | gtex_brain_putamen_basal |
| rs6531583 | 4 | 37854693 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 26438 | gtex_brain_putamen_basal |
| rs6531584 | 4 | 37854756 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 26501 | gtex_brain_putamen_basal |
| rs6531585 | 4 | 37854827 | PGM2 | ENSG00000169299.9 | 1.043E-8 | 0 | 26572 | gtex_brain_putamen_basal |
| rs6531586 | 4 | 37854849 | PGM2 | ENSG00000169299.9 | 1.046E-8 | 0 | 26594 | gtex_brain_putamen_basal |
| rs6531587 | 4 | 37854911 | PGM2 | ENSG00000169299.9 | 1.046E-8 | 0 | 26656 | gtex_brain_putamen_basal |
| rs6827984 | 4 | 37854920 | PGM2 | ENSG00000169299.9 | 1.046E-8 | 0 | 26665 | gtex_brain_putamen_basal |
| rs11730888 | 4 | 37855550 | PGM2 | ENSG00000169299.9 | 1.334E-8 | 0 | 27295 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
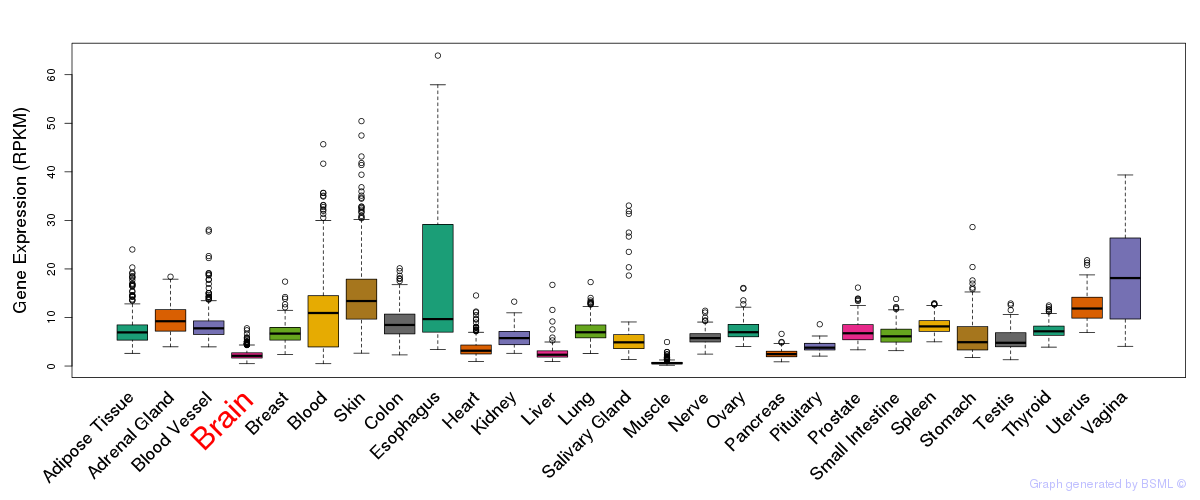
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG GLYCOLYSIS GLUCONEOGENESIS | 62 | 41 | All SZGR 2.0 genes in this pathway |
| KEGG PENTOSE PHOSPHATE PATHWAY | 27 | 14 | All SZGR 2.0 genes in this pathway |
| KEGG GALACTOSE METABOLISM | 26 | 22 | All SZGR 2.0 genes in this pathway |
| KEGG STARCH AND SUCROSE METABOLISM | 52 | 33 | All SZGR 2.0 genes in this pathway |
| KEGG AMINO SUGAR AND NUCLEOTIDE SUGAR METABOLISM | 44 | 30 | All SZGR 2.0 genes in this pathway |
| REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | 18 | 10 | All SZGR 2.0 genes in this pathway |
| REACTOME METABOLISM OF CARBOHYDRATES | 247 | 154 | All SZGR 2.0 genes in this pathway |
| REACTOME GLUCOSE METABOLISM | 69 | 42 | All SZGR 2.0 genes in this pathway |
| NAKAMURA TUMOR ZONE PERIPHERAL VS CENTRAL DN | 634 | 384 | All SZGR 2.0 genes in this pathway |
| FULCHER INFLAMMATORY RESPONSE LECTIN VS LPS UP | 579 | 346 | All SZGR 2.0 genes in this pathway |
| GARY CD5 TARGETS DN | 431 | 263 | All SZGR 2.0 genes in this pathway |
| CHARAFE BREAST CANCER LUMINAL VS BASAL DN | 455 | 304 | All SZGR 2.0 genes in this pathway |
| LASTOWSKA NEUROBLASTOMA COPY NUMBER DN | 800 | 473 | All SZGR 2.0 genes in this pathway |
| MOHANKUMAR TLX1 TARGETS UP | 414 | 287 | All SZGR 2.0 genes in this pathway |
| KIM MYC AMPLIFICATION TARGETS UP | 201 | 127 | All SZGR 2.0 genes in this pathway |
| SHETH LIVER CANCER VS TXNIP LOSS PAM2 | 153 | 102 | All SZGR 2.0 genes in this pathway |
| WEI MYCN TARGETS WITH E BOX | 795 | 478 | All SZGR 2.0 genes in this pathway |
| CUI TCF21 TARGETS 2 DN | 830 | 547 | All SZGR 2.0 genes in this pathway |
| ACEVEDO NORMAL TISSUE ADJACENT TO LIVER TUMOR DN | 354 | 216 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| GOLDRATH ANTIGEN RESPONSE | 346 | 192 | All SZGR 2.0 genes in this pathway |
| CHANG CORE SERUM RESPONSE UP | 212 | 128 | All SZGR 2.0 genes in this pathway |
| SWEET LUNG CANCER KRAS DN | 435 | 289 | All SZGR 2.0 genes in this pathway |
| LEE RECENT THYMIC EMIGRANT | 227 | 128 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP D | 280 | 158 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE EARLY LATE | 317 | 190 | All SZGR 2.0 genes in this pathway |
| YANG BCL3 TARGETS UP | 364 | 236 | All SZGR 2.0 genes in this pathway |
| KRIEG HYPOXIA NOT VIA KDM3A | 770 | 480 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS DN | 784 | 464 | All SZGR 2.0 genes in this pathway |