Gene Page: ZNF407
Summary ?
| GeneID | 55628 |
| Symbol | ZNF407 |
| Synonyms | - |
| Description | zinc finger protein 407 |
| Reference | MIM:615894|HGNC:HGNC:19904|Ensembl:ENSG00000215421|HPRD:11713|Vega:OTTHUMG00000179122 |
| Gene type | protein-coding |
| Map location | 18q23 |
| Pascal p-value | 5.08E-4 |
| TADA p-value | 0.006 |
| eGene | Meta |
| Support | CompositeSet Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ZNF407 | chr18 | 72775784 | A | AG | NM_017757 | . | frameshift | Schizophrenia | DNM:Fromer_2014 |
Section II. Transcriptome annotation
General gene expression (GTEx)
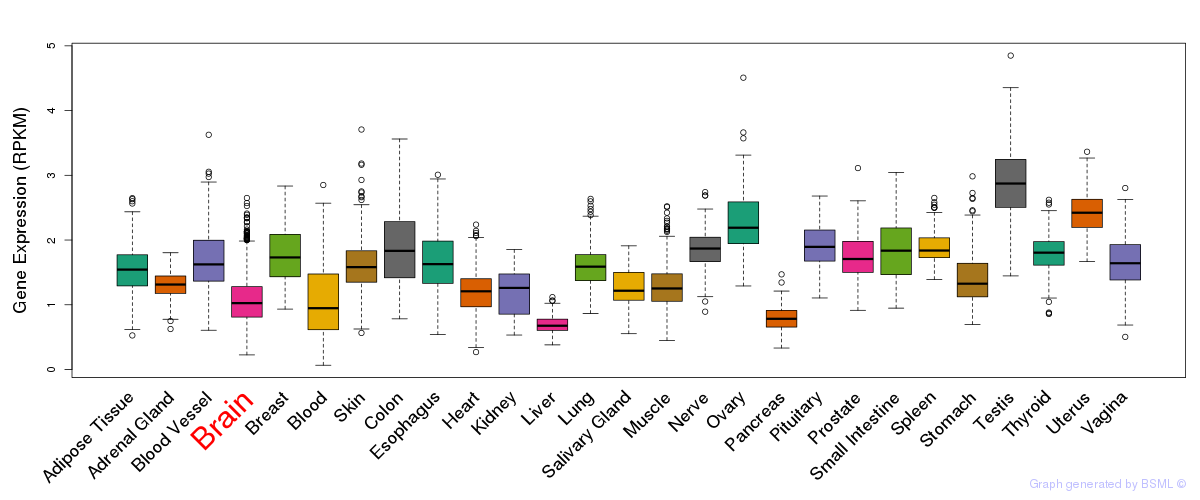
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AC112484.1 | 0.68 | 0.44 |
| C20orf26 | 0.63 | 0.44 |
| LMCD1 | 0.61 | 0.73 |
| CLDN3 | 0.61 | 0.27 |
| MYH7 | 0.61 | 0.56 |
| C17orf72 | 0.60 | 0.47 |
| IQCH | 0.60 | 0.34 |
| AC015908.2 | 0.60 | 0.55 |
| PGAM2 | 0.60 | 0.37 |
| KRT18P19 | 0.60 | -0.08 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ISLR2 | -0.33 | -0.36 |
| DPP4 | -0.32 | -0.42 |
| FOXG1B | -0.32 | -0.37 |
| TIAM2 | -0.32 | -0.39 |
| KLHL1 | -0.31 | -0.35 |
| SSBP2 | -0.31 | -0.38 |
| SLA | -0.31 | -0.33 |
| TTC28 | -0.31 | -0.31 |
| NEUROD6 | -0.31 | -0.41 |
| SEMA3A | -0.31 | -0.39 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| GARY CD5 TARGETS DN | 431 | 263 | All SZGR 2.0 genes in this pathway |
| WANG LMO4 TARGETS DN | 352 | 225 | All SZGR 2.0 genes in this pathway |
| HAMAI APOPTOSIS VIA TRAIL UP | 584 | 356 | All SZGR 2.0 genes in this pathway |
| STAMBOLSKY TARGETS OF MUTATED TP53 UP | 49 | 35 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS DN | 1972 | 1213 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 1ST EGF PULSE ONLY | 1839 | 928 | All SZGR 2.0 genes in this pathway |