Gene Page: NSUN5
Summary ?
| GeneID | 55695 |
| Symbol | NSUN5 |
| Synonyms | NOL1|NOL1R|NSUN5A|WBSCR20|WBSCR20A|p120|p120(NOL1) |
| Description | NOP2/Sun RNA methyltransferase family member 5 |
| Reference | MIM:615732|HGNC:HGNC:16385|Ensembl:ENSG00000130305|HPRD:11678|Vega:OTTHUMG00000129869 |
| Gene type | protein-coding |
| Map location | 7q11.23 |
| Pascal p-value | 0.756 |
| Sherlock p-value | 0.441 |
| Fetal beta | 1.182 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CNV:YES | Copy number variation studies | Manual curation |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
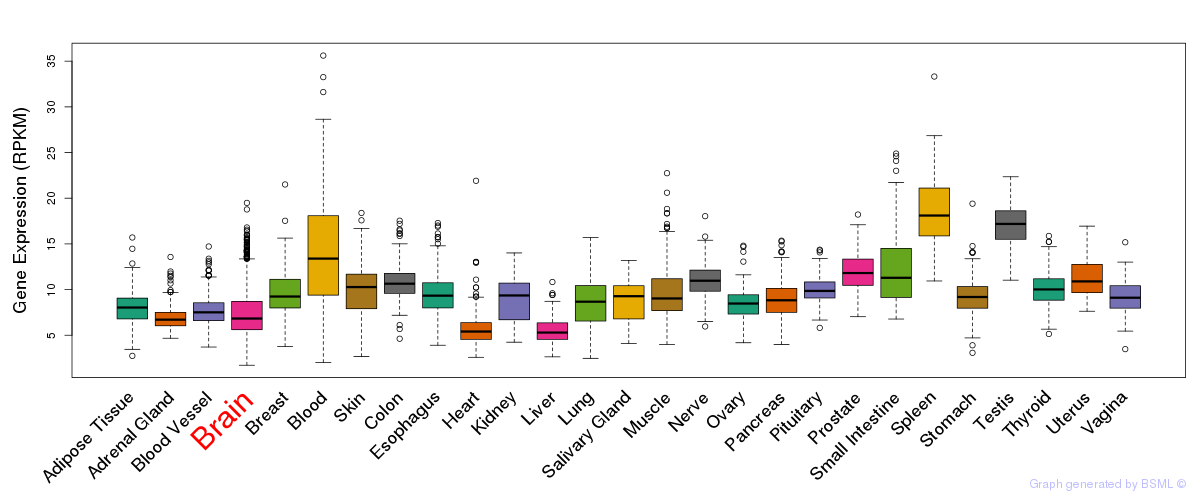
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| GART | 0.97 | 0.92 |
| SLC39A6 | 0.96 | 0.93 |
| DHX9 | 0.96 | 0.93 |
| XRN2 | 0.96 | 0.95 |
| NUP160 | 0.96 | 0.91 |
| SMAD4 | 0.96 | 0.95 |
| SF3B3 | 0.96 | 0.93 |
| GPN1 | 0.96 | 0.92 |
| TRAFD1 | 0.96 | 0.91 |
| KHDRBS1 | 0.96 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| HLA-F | -0.75 | -0.79 |
| AIFM3 | -0.74 | -0.78 |
| C5orf53 | -0.73 | -0.74 |
| MT-CO2 | -0.72 | -0.87 |
| AF347015.27 | -0.72 | -0.84 |
| AF347015.33 | -0.72 | -0.85 |
| FXYD1 | -0.71 | -0.86 |
| AF347015.31 | -0.71 | -0.84 |
| S100B | -0.70 | -0.79 |
| FBXO2 | -0.70 | -0.67 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| LIU SOX4 TARGETS DN | 309 | 191 | All SZGR 2.0 genes in this pathway |
| CHEMNITZ RESPONSE TO PROSTAGLANDIN E2 DN | 391 | 222 | All SZGR 2.0 genes in this pathway |
| VECCHI GASTRIC CANCER EARLY UP | 430 | 232 | All SZGR 2.0 genes in this pathway |
| GINESTIER BREAST CANCER 20Q13 AMPLIFICATION DN | 180 | 101 | All SZGR 2.0 genes in this pathway |
| SHEPARD BMYB MORPHOLINO UP | 205 | 126 | All SZGR 2.0 genes in this pathway |
| SANSOM APC TARGETS REQUIRE MYC | 210 | 123 | All SZGR 2.0 genes in this pathway |
| MASSARWEH TAMOXIFEN RESISTANCE DN | 258 | 160 | All SZGR 2.0 genes in this pathway |
| SENGUPTA EBNA1 ANTICORRELATED | 173 | 85 | All SZGR 2.0 genes in this pathway |
| KOINUMA TARGETS OF SMAD2 OR SMAD3 | 824 | 528 | All SZGR 2.0 genes in this pathway |