Gene Page: PCID2
Summary ?
| GeneID | 55795 |
| Symbol | PCID2 |
| Synonyms | F10 |
| Description | PCI domain containing 2 |
| Reference | MIM:613713|HGNC:HGNC:25653|Ensembl:ENSG00000126226|HPRD:07749|Vega:OTTHUMG00000017385 |
| Gene type | protein-coding |
| Map location | 13q34 |
| Pascal p-value | 0.177 |
| Sherlock p-value | 0.043 |
| Fetal beta | 0.777 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
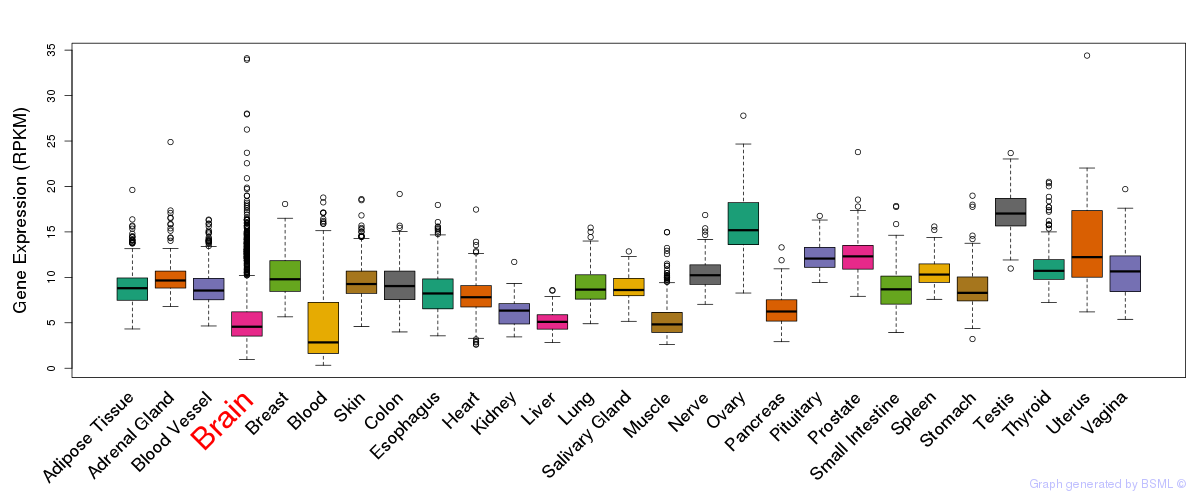
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ERMP1 | 0.84 | 0.81 |
| MFAP3L | 0.84 | 0.85 |
| ENPP4 | 0.83 | 0.76 |
| ENDOD1 | 0.83 | 0.81 |
| MBNL2 | 0.83 | 0.85 |
| RNF13 | 0.83 | 0.81 |
| CLIP4 | 0.82 | 0.73 |
| KIAA0494 | 0.82 | 0.80 |
| LAMP2 | 0.82 | 0.86 |
| UGT8 | 0.81 | 0.81 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| BCL7C | -0.61 | -0.74 |
| KIAA1949 | -0.58 | -0.55 |
| TUBB2B | -0.58 | -0.62 |
| CCDC28B | -0.58 | -0.71 |
| PPP1R14B | -0.57 | -0.71 |
| DYNLT1 | -0.57 | -0.74 |
| PLEKHO1 | -0.57 | -0.62 |
| NR2C2AP | -0.57 | -0.62 |
| SH2B2 | -0.57 | -0.66 |
| NKIRAS2 | -0.56 | -0.48 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005515 | protein binding | IPI | 17353931 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| MAYBURD RESPONSE TO L663536 DN | 56 | 32 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN DN | 770 | 415 | All SZGR 2.0 genes in this pathway |
| GAUSSMANN MLL AF4 FUSION TARGETS F UP | 185 | 119 | All SZGR 2.0 genes in this pathway |
| BERENJENO TRANSFORMED BY RHOA UP | 536 | 340 | All SZGR 2.0 genes in this pathway |
| LOPEZ MBD TARGETS | 957 | 597 | All SZGR 2.0 genes in this pathway |
| LOCKWOOD AMPLIFIED IN LUNG CANCER | 214 | 139 | All SZGR 2.0 genes in this pathway |
| BENPORATH EED TARGETS | 1062 | 725 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR DN | 1011 | 592 | All SZGR 2.0 genes in this pathway |
| ALONSO METASTASIS UP | 198 | 128 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |