Gene Page: SAG
Summary ?
| GeneID | 6295 |
| Symbol | SAG |
| Synonyms | RP47|S-AG |
| Description | S-antigen; retina and pineal gland (arrestin) |
| Reference | MIM:181031|HGNC:HGNC:10521|Ensembl:ENSG00000130561|HPRD:01625|Vega:OTTHUMG00000153213 |
| Gene type | protein-coding |
| Map location | 2q37.1 |
| Pascal p-value | 0.875 |
| Fetal beta | 0.086 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
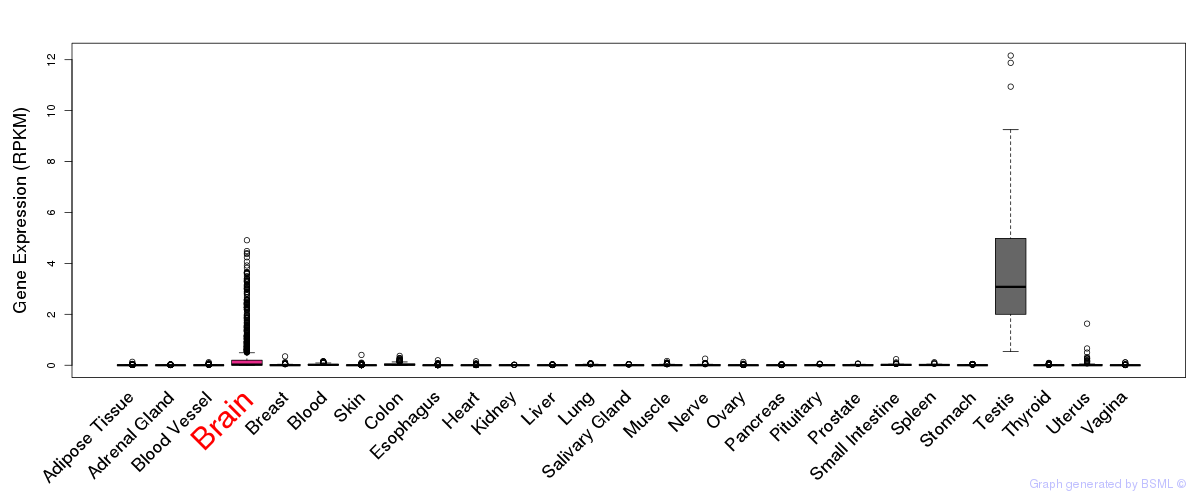
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PDE1B | 0.91 | 0.88 |
| KCNAB1 | 0.89 | 0.84 |
| RASD2 | 0.88 | 0.85 |
| ANO3 | 0.87 | 0.82 |
| ENTPD3 | 0.84 | 0.82 |
| AC005512.1 | 0.82 | 0.55 |
| GPR52 | 0.81 | 0.75 |
| IL17RA | 0.81 | 0.71 |
| DGKB | 0.80 | 0.82 |
| SH3D20 | 0.80 | 0.42 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SH2B2 | -0.29 | -0.42 |
| BCL7C | -0.29 | -0.54 |
| KIAA1949 | -0.28 | -0.30 |
| TUBB2B | -0.28 | -0.40 |
| RPL18 | -0.28 | -0.51 |
| PPP1R14B | -0.28 | -0.46 |
| SH3BP2 | -0.28 | -0.41 |
| TRAF4 | -0.28 | -0.45 |
| RPL38 | -0.27 | -0.55 |
| RBMX2 | -0.27 | -0.45 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| SIG CHEMOTAXIS | 45 | 37 | All SZGR 2.0 genes in this pathway |
| SIG PIP3 SIGNALING IN B LYMPHOCYTES | 36 | 31 | All SZGR 2.0 genes in this pathway |
| PID RHODOPSIN PATHWAY | 24 | 10 | All SZGR 2.0 genes in this pathway |
| ZHENG BOUND BY FOXP3 | 491 | 310 | All SZGR 2.0 genes in this pathway |
| YAGI AML WITH 11Q23 REARRANGED | 351 | 238 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP D | 280 | 158 | All SZGR 2.0 genes in this pathway |