Gene Page: SHBG
Summary ?
| GeneID | 6462 |
| Symbol | SHBG |
| Synonyms | ABP|SBP|TEBG |
| Description | sex hormone-binding globulin |
| Reference | MIM:182205|HGNC:HGNC:10839|Ensembl:ENSG00000129214|HPRD:01646|Vega:OTTHUMG00000108153 |
| Gene type | protein-coding |
| Map location | 17p13.1 |
| Pascal p-value | 0.38 |
| eGene | Caudate basal ganglia |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
| Network | Shortest path distance of core genes in the Human protein-protein interaction network | Contribution to shortest path in PPI network: 0.5587 |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
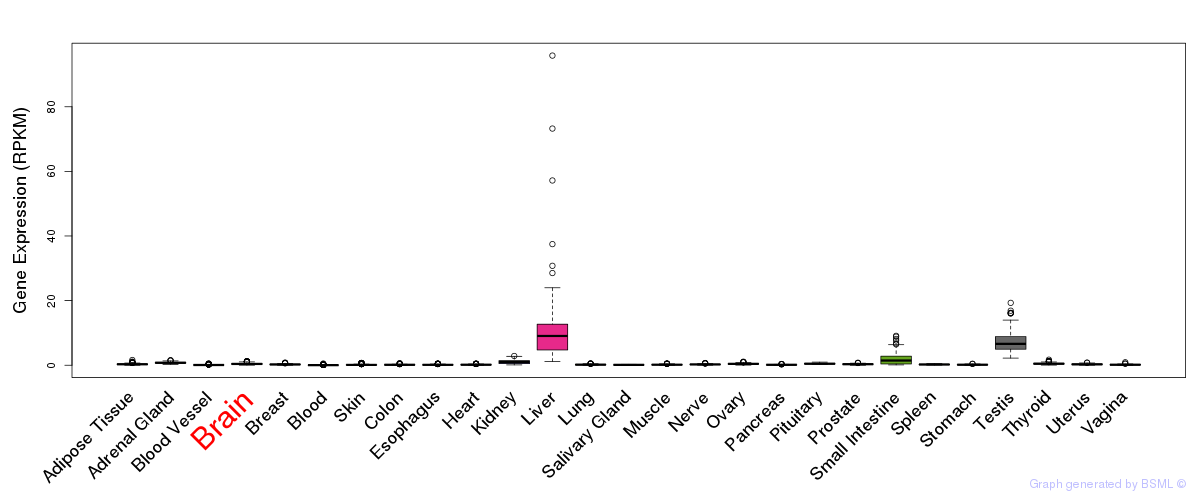
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZYG11B | 0.91 | 0.95 |
| TBL1XR1 | 0.90 | 0.95 |
| RLIM | 0.90 | 0.94 |
| PRKCI | 0.89 | 0.92 |
| ZNF295 | 0.89 | 0.94 |
| KIAA1219 | 0.89 | 0.94 |
| ZBTB44 | 0.89 | 0.94 |
| SCAMP1 | 0.89 | 0.92 |
| YTHDF3 | 0.88 | 0.93 |
| GTF2A1 | 0.88 | 0.94 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| FXYD1 | -0.66 | -0.74 |
| AF347015.31 | -0.64 | -0.73 |
| MT-CO2 | -0.64 | -0.74 |
| HIGD1B | -0.63 | -0.72 |
| CST3 | -0.63 | -0.71 |
| HSD17B14 | -0.62 | -0.67 |
| AF347015.21 | -0.62 | -0.74 |
| TLCD1 | -0.62 | -0.68 |
| MT-CYB | -0.62 | -0.70 |
| AF347015.8 | -0.61 | -0.72 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005497 | androgen binding | TAS | 2587256 | |
| GO:0042803 | protein homodimerization activity | NAS | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0009914 | hormone transport | NAS | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0005576 | extracellular region | NAS | 14718574 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID AR NONGENOMIC PATHWAY | 31 | 27 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER TUMOR VS NORMAL ADJACENT TISSUE DN | 274 | 165 | All SZGR 2.0 genes in this pathway |
| CAIRO HEPATOBLASTOMA DN | 267 | 160 | All SZGR 2.0 genes in this pathway |