Gene Page: SRY
Summary ?
| GeneID | 6736 |
| Symbol | SRY |
| Synonyms | SRXX1|SRXY1|TDF|TDY |
| Description | sex determining region Y |
| Reference | MIM:480000|HGNC:HGNC:11311|Ensembl:ENSG00000184895|HPRD:08364|Vega:OTTHUMG00000036084 |
| Gene type | protein-coding |
| Map location | Yp11.3 |
| Fetal beta | 0.533 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenics,schizophrenias,schizotypal | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
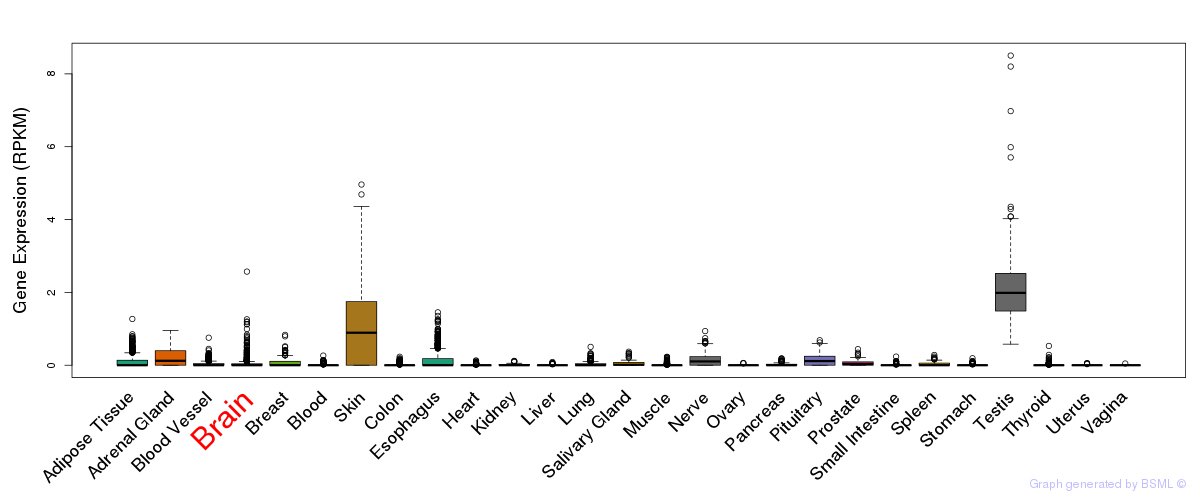
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| GM2A | 0.81 | 0.82 |
| ZCCHC24 | 0.80 | 0.84 |
| TPP1 | 0.80 | 0.83 |
| CSF1 | 0.78 | 0.82 |
| KANK1 | 0.78 | 0.83 |
| ITPKB | 0.78 | 0.83 |
| CDH20 | 0.77 | 0.78 |
| SSFA2 | 0.77 | 0.78 |
| DAAM2 | 0.77 | 0.80 |
| SASH1 | 0.77 | 0.80 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| NR2C2AP | -0.54 | -0.59 |
| MED19 | -0.54 | -0.60 |
| ALKBH2 | -0.51 | -0.59 |
| POLB | -0.51 | -0.59 |
| COMMD3 | -0.51 | -0.58 |
| PLEKHO1 | -0.51 | -0.52 |
| FRG1 | -0.51 | -0.59 |
| STMN1 | -0.50 | -0.51 |
| ING4 | -0.50 | -0.53 |
| ZNF821 | -0.49 | -0.47 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID AR TF PATHWAY | 53 | 38 | All SZGR 2.0 genes in this pathway |
| SCHLOSSER SERUM RESPONSE DN | 712 | 443 | All SZGR 2.0 genes in this pathway |
| KAYO CALORIE RESTRICTION MUSCLE UP | 95 | 64 | All SZGR 2.0 genes in this pathway |
| BANDRES RESPONSE TO CARMUSTIN MGMT 48HR DN | 161 | 105 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 7 UP | 118 | 68 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP A | 898 | 516 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |