Gene Page: TACR1
Summary ?
| GeneID | 6869 |
| Symbol | TACR1 |
| Synonyms | NK1R|NKIR|SPR|TAC1R |
| Description | tachykinin receptor 1 |
| Reference | MIM:162323|HGNC:HGNC:11526|Ensembl:ENSG00000115353|HPRD:01208|Vega:OTTHUMG00000129973 |
| Gene type | protein-coding |
| Map location | 2p12 |
| Pascal p-value | 0.054 |
| Sherlock p-value | 0.334 |
| Fetal beta | 0.152 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
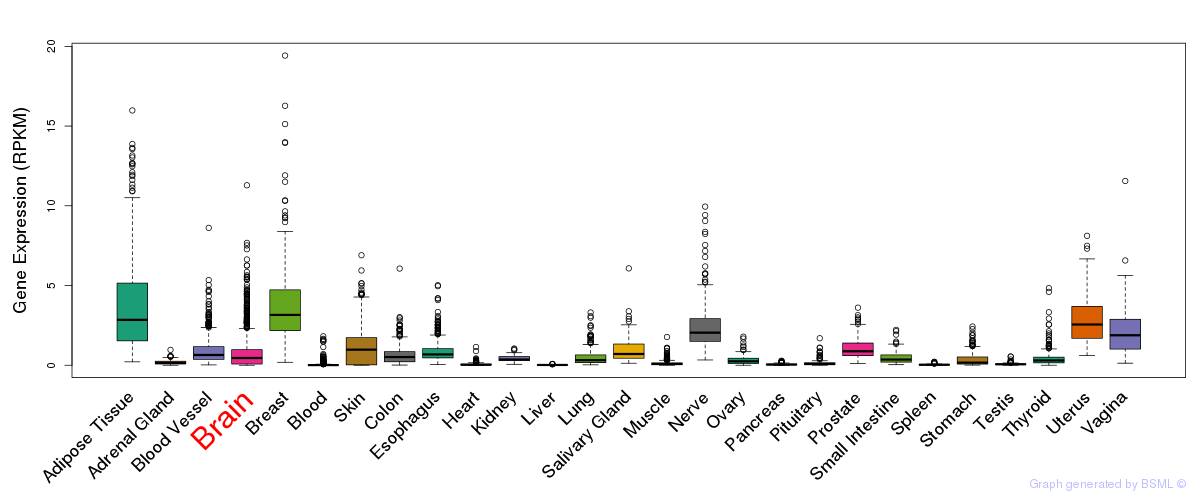
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| GPER | 0.87 | 0.83 |
| FAM38A | 0.83 | 0.84 |
| SLC6A12 | 0.81 | 0.79 |
| SOX18 | 0.80 | 0.79 |
| RIN3 | 0.79 | 0.80 |
| BCAM | 0.78 | 0.81 |
| HLA-E | 0.78 | 0.79 |
| GPR37L1 | 0.78 | 0.71 |
| COL5A3 | 0.77 | 0.73 |
| TINAGL1 | 0.77 | 0.76 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| FRG1 | -0.64 | -0.71 |
| PSMA4 | -0.60 | -0.63 |
| FARSB | -0.60 | -0.60 |
| ASNSD1 | -0.59 | -0.59 |
| SPATA7 | -0.59 | -0.59 |
| C18orf21 | -0.59 | -0.60 |
| COQ3 | -0.59 | -0.59 |
| TXNL1 | -0.58 | -0.60 |
| GGPS1 | -0.58 | -0.56 |
| POLB | -0.57 | -0.64 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0004872 | receptor activity | IEA | - | |
| GO:0004995 | tachykinin receptor activity | IEA | - | |
| GO:0004995 | tachykinin receptor activity | TAS | 8985172 | |
| GO:0005515 | protein binding | IPI | 17986524 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0007200 | G-protein signaling, coupled to IP3 second messenger (phospholipase C activating) | TAS | 1657150 | |
| GO:0007186 | G-protein coupled receptor protein signaling pathway | IEA | - | |
| GO:0007217 | tachykinin signaling pathway | TAS | 8985172 | |
| GO:0007165 | signal transduction | IEA | - | |
| GO:0009582 | detection of abiotic stimulus | TAS | 9537323 | |
| GO:0006954 | inflammatory response | TAS | 10854844 | |
| GO:0007638 | mechanosensory behavior | TAS | 9537323 | |
| GO:0048265 | response to pain | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0016021 | integral to membrane | IEA | - | |
| GO:0005886 | plasma membrane | IEA | - | |
| GO:0005886 | plasma membrane | TAS | 10611312 | |
| GO:0005887 | integral to plasma membrane | TAS | 1657150 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG CALCIUM SIGNALING PATHWAY | 178 | 134 | All SZGR 2.0 genes in this pathway |
| KEGG NEUROACTIVE LIGAND RECEPTOR INTERACTION | 272 | 195 | All SZGR 2.0 genes in this pathway |
| REACTOME GASTRIN CREB SIGNALLING PATHWAY VIA PKC AND MAPK | 205 | 136 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALING BY GPCR | 920 | 449 | All SZGR 2.0 genes in this pathway |
| REACTOME PEPTIDE LIGAND BINDING RECEPTORS | 188 | 108 | All SZGR 2.0 genes in this pathway |
| REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | 305 | 177 | All SZGR 2.0 genes in this pathway |
| REACTOME G ALPHA Q SIGNALLING EVENTS | 184 | 116 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR DOWNSTREAM SIGNALING | 805 | 368 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR LIGAND BINDING | 408 | 246 | All SZGR 2.0 genes in this pathway |
| HERNANDEZ MITOTIC ARREST BY DOCETAXEL 1 UP | 36 | 18 | All SZGR 2.0 genes in this pathway |
| HATADA METHYLATED IN LUNG CANCER UP | 390 | 236 | All SZGR 2.0 genes in this pathway |
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| BENPORATH ES WITH H3K27ME3 | 1118 | 744 | All SZGR 2.0 genes in this pathway |
| MOREAUX MULTIPLE MYELOMA BY TACI UP | 412 | 249 | All SZGR 2.0 genes in this pathway |
| HILLION HMGA1 TARGETS | 90 | 71 | All SZGR 2.0 genes in this pathway |
| KONDO EZH2 TARGETS | 245 | 148 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN NPC HCP WITH H3K4ME3 AND H3K27ME3 | 210 | 148 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3K27ME3 | 590 | 403 | All SZGR 2.0 genes in this pathway |