Gene Page: TADA2A
Summary ?
| GeneID | 6871 |
| Symbol | TADA2A |
| Synonyms | ADA2|ADA2A|KL04P|TADA2L|hADA2 |
| Description | transcriptional adaptor 2A |
| Reference | MIM:602276|HGNC:HGNC:11531|Ensembl:ENSG00000276234|HPRD:03784|Vega:OTTHUMG00000188469 |
| Gene type | protein-coding |
| Map location | 17q12-q21 |
| Pascal p-value | 0.275 |
| Fetal beta | 0.883 |
| eGene | Cerebellar Hemisphere Cerebellum Frontal Cortex BA9 Hypothalamus |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CNV:YES | Copy number variation studies | Manual curation | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
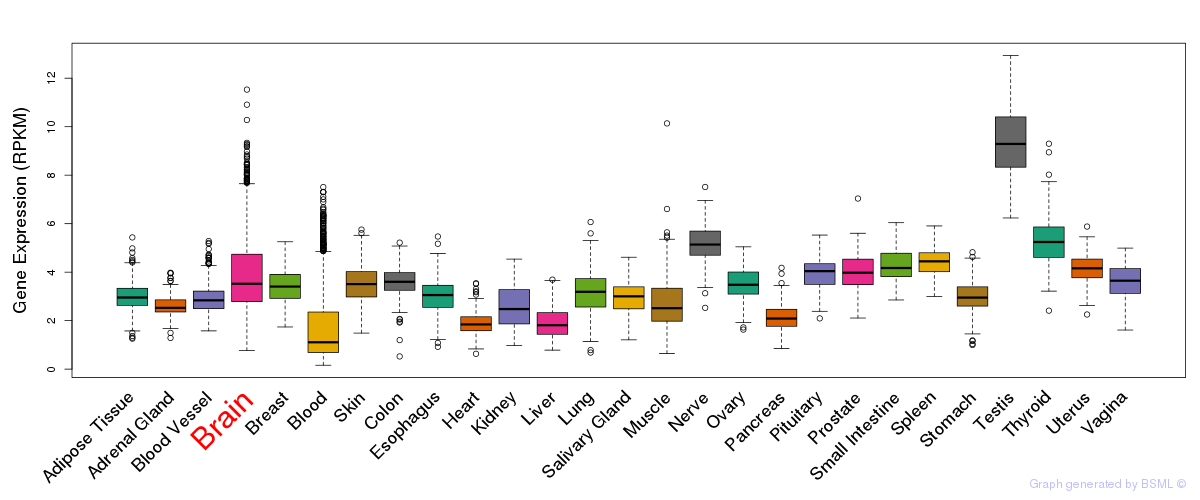
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| EXOC3L | 0.70 | 0.50 |
| MUC1 | 0.69 | 0.50 |
| CCDC88C | 0.68 | 0.49 |
| CASZ1 | 0.68 | 0.49 |
| TNK1 | 0.67 | 0.43 |
| ATG9B | 0.67 | 0.46 |
| PLEKHG4 | 0.66 | 0.50 |
| FBF1 | 0.66 | 0.61 |
| KRTCAP3 | 0.65 | 0.31 |
| NPEPL1 | 0.65 | 0.61 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| CHPT1 | -0.37 | -0.39 |
| KBTBD3 | -0.33 | -0.41 |
| HIGD1A | -0.33 | -0.36 |
| BCAP29 | -0.32 | -0.32 |
| TPMT | -0.32 | -0.32 |
| PLA2G4A | -0.32 | -0.32 |
| UQCRB | -0.32 | -0.38 |
| NDUFB3 | -0.31 | -0.34 |
| NDUFB5 | -0.31 | -0.41 |
| ITM2B | -0.31 | -0.30 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| GAZDA DIAMOND BLACKFAN ANEMIA ERYTHROID DN | 493 | 298 | All SZGR 2.0 genes in this pathway |
| CASORELLI ACUTE PROMYELOCYTIC LEUKEMIA DN | 663 | 425 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 12HR DN | 209 | 122 | All SZGR 2.0 genes in this pathway |
| MIDORIKAWA AMPLIFIED IN LIVER CANCER | 55 | 38 | All SZGR 2.0 genes in this pathway |
| SPIELMAN LYMPHOBLAST EUROPEAN VS ASIAN UP | 479 | 299 | All SZGR 2.0 genes in this pathway |
| ROPERO HDAC2 TARGETS | 114 | 71 | All SZGR 2.0 genes in this pathway |
| NIKOLSKY BREAST CANCER 17Q11 Q21 AMPLICON | 133 | 78 | All SZGR 2.0 genes in this pathway |
| DEBIASI APOPTOSIS BY REOVIRUS INFECTION UP | 314 | 201 | All SZGR 2.0 genes in this pathway |
| KIM GASTRIC CANCER CHEMOSENSITIVITY | 103 | 64 | All SZGR 2.0 genes in this pathway |
| BONOME OVARIAN CANCER SURVIVAL SUBOPTIMAL DEBULKING | 510 | 309 | All SZGR 2.0 genes in this pathway |
| ASGHARZADEH NEUROBLASTOMA POOR SURVIVAL DN | 46 | 30 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE EARLY LATE | 317 | 190 | All SZGR 2.0 genes in this pathway |