Gene Page: TP53BP1
Summary ?
| GeneID | 7158 |
| Symbol | TP53BP1 |
| Synonyms | 53BP1|TDRD30|TP53|p202|p53BP1 |
| Description | tumor protein p53 binding protein 1 |
| Reference | MIM:605230|HGNC:HGNC:11999|Ensembl:ENSG00000067369|HPRD:05569|Vega:OTTHUMG00000059757 |
| Gene type | protein-coding |
| Map location | 15q15-q21 |
| Pascal p-value | 4.038E-4 |
| Sherlock p-value | 0.038 |
| Fetal beta | 0.318 |
| Support | G2Cdb.humanNRC G2Cdb.humanPSP Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
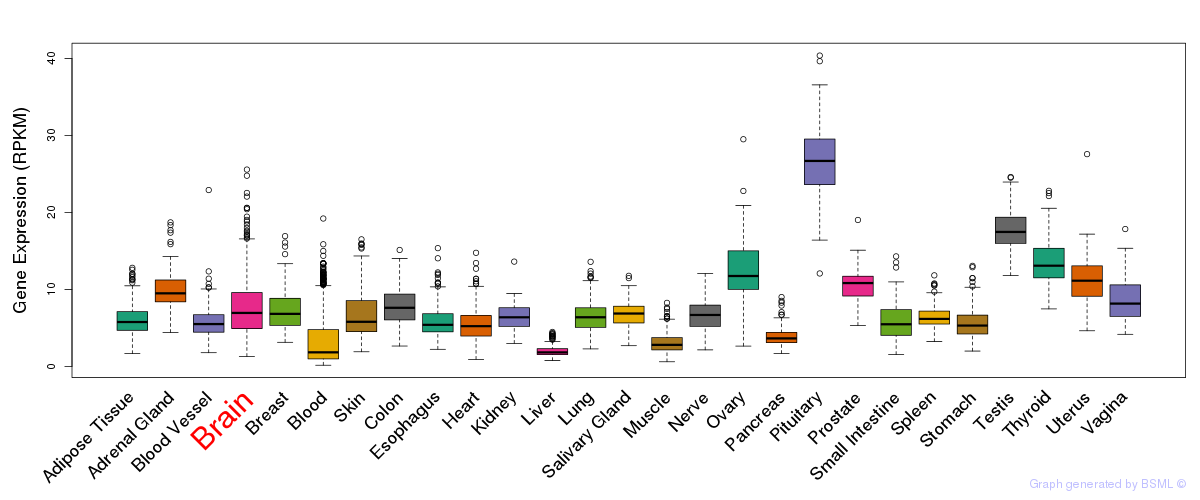
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| OCM | 0.60 | 0.33 |
| IGSF10 | 0.54 | 0.61 |
| TACR3 | 0.54 | 0.53 |
| ARHGAP6 | 0.53 | 0.30 |
| ODZ1 | 0.48 | 0.68 |
| LUZP2 | 0.48 | 0.52 |
| SLC41A2 | 0.47 | 0.65 |
| RP3-402H5.1 | 0.47 | 0.56 |
| SCN9A | 0.47 | 0.65 |
| CSMD3 | 0.47 | 0.68 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.26 | -0.50 |
| HIGD1B | -0.26 | -0.51 |
| AC098691.2 | -0.26 | -0.49 |
| AF347015.2 | -0.26 | -0.47 |
| C11orf67 | -0.25 | -0.45 |
| CSAG1 | -0.25 | -0.42 |
| MT-CYB | -0.25 | -0.47 |
| AF347015.8 | -0.24 | -0.48 |
| AF347015.33 | -0.24 | -0.46 |
| AF347015.31 | -0.24 | -0.47 |
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| ATM | AT1 | ATA | ATC | ATD | ATDC | ATE | DKFZp781A0353 | MGC74674 | TEL1 | TELO1 | ataxia telangiectasia mutated | Biochemical Activity Reconstituted Complex | BioGRID | 12447390 |12697768 |
| BLM | BS | MGC126616 | MGC131618 | MGC131620 | RECQ2 | RECQL2 | RECQL3 | Bloom syndrome | Affinity Capture-Western Co-localization | BioGRID | 15364958 |
| BRCA1 | BRCAI | BRCC1 | IRIS | PSCP | RNF53 | breast cancer 1, early onset | Affinity Capture-Western | BioGRID | 17525332 |
| CHEK1 | CHK1 | CHK1 checkpoint homolog (S. pombe) | Co-localization | BioGRID | 15364958 |
| CHEK2 | CDS1 | CHK2 | HuCds1 | LFS2 | PP1425 | RAD53 | CHK2 checkpoint homolog (S. pombe) | Affinity Capture-Western | BioGRID | 17525332 |
| DCLRE1A | KIAA0086 | PSO2 | SNM1 | SNM1A | hSNM1 | DNA cross-link repair 1A (PSO2 homolog, S. cerevisiae) | Affinity Capture-Western | BioGRID | 12446782 |
| DCLRE1C | A-SCID | DCLREC1C | FLJ11360 | FLJ36438 | RS-SCID | SCIDA | SNM1C | DNA cross-link repair 1C (PSO2 homolog, S. cerevisiae) | Artemis interacts with 53BP1. | BIND | 15574327 |
| DCTN1 | DAP-150 | DP-150 | HMN7B | P135 | dynactin 1 (p150, glued homolog, Drosophila) | Affinity Capture-Western | BioGRID | 15611139 |
| DYNLL1 | DLC1 | DLC8 | DNCL1 | DNCLC1 | LC8 | LC8a | MGC126137 | MGC126138 | PIN | hdlc1 | dynein, light chain, LC8-type 1 | Affinity Capture-Western Protein-peptide Two-hybrid | BioGRID | 15611139 |
| E2F1 | E2F-1 | RBAP1 | RBBP3 | RBP3 | E2F transcription factor 1 | Affinity Capture-Western Reconstituted Complex | BioGRID | 8896460 |
| H2AFX | H2A.X | H2A/X | H2AX | H2A histone family, member X | - | HPRD,BioGRID | 12697768 |
| MDC1 | DKFZp781A0122 | KIAA0170 | MGC166888 | NFBD1 | mediator of DNA damage checkpoint 1 | - | HPRD | 12607005 |
| NEK1 | DKFZp686D06121 | DKFZp686K12169 | KIAA1901 | MGC138800 | NY-REN-55 | NIMA (never in mitosis gene a)-related kinase 1 | NEK1 interacts with 53BP1. | BIND | 14690447 |
| NFKB1 | DKFZp686C01211 | EBP-1 | KBF1 | MGC54151 | NF-kappa-B | NFKB-p105 | NFKB-p50 | p105 | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 | Affinity Capture-Western | BioGRID | 10646849 |
| PAXIP1 | CAGF28 | CAGF29 | FLJ41049 | PACIP1 | PAXIP1L | PTIP | TNRC2 | PAX interacting (with transcription-activation domain) protein 1 | Affinity Capture-Western Reconstituted Complex | BioGRID | 14576432 |15456759 |
| RPA1 | HSSB | REPA1 | RF-A | RP-A | RPA70 | replication protein A1, 70kDa | Affinity Capture-MS Affinity Capture-Western | BioGRID | 15856006 |
| RPA2 | REPA2 | RPA32 | replication protein A2, 32kDa | Affinity Capture-MS Affinity Capture-Western | BioGRID | 15856006 |
| TFDP1 | DP1 | DRTF1 | Dp-1 | transcription factor Dp-1 | Reconstituted Complex | BioGRID | 8896460 |
| TP53 | FLJ92943 | LFS1 | TRP53 | p53 | tumor protein p53 | Affinity Capture-Western Co-crystal Structure Co-localization Reconstituted Complex Two-hybrid | BioGRID | 8016121 |11877378 |12110597 |12351827 |14978302 |15364958 |15611139 |
| TP53 | FLJ92943 | LFS1 | TRP53 | p53 | tumor protein p53 | - | HPRD | 8016121 |12110597 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID ATM PATHWAY | 34 | 25 | All SZGR 2.0 genes in this pathway |
| REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | 17 | 11 | All SZGR 2.0 genes in this pathway |
| REACTOME DOUBLE STRAND BREAK REPAIR | 24 | 14 | All SZGR 2.0 genes in this pathway |
| REACTOME DNA REPAIR | 112 | 59 | All SZGR 2.0 genes in this pathway |
| ENK UV RESPONSE KERATINOCYTE DN | 485 | 334 | All SZGR 2.0 genes in this pathway |
| MARTIN INTERACT WITH HDAC | 44 | 31 | All SZGR 2.0 genes in this pathway |
| PATIL LIVER CANCER | 747 | 453 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 DN | 855 | 609 | All SZGR 2.0 genes in this pathway |
| PUJANA BREAST CANCER LIT INT NETWORK | 101 | 73 | All SZGR 2.0 genes in this pathway |
| RICKMAN METASTASIS UP | 344 | 180 | All SZGR 2.0 genes in this pathway |
| KAUFFMANN DNA REPAIR GENES | 230 | 137 | All SZGR 2.0 genes in this pathway |
| HESS TARGETS OF HOXA9 AND MEIS1 UP | 65 | 44 | All SZGR 2.0 genes in this pathway |
| CHIARETTI ACUTE LYMPHOBLASTIC LEUKEMIA ZAP70 | 67 | 33 | All SZGR 2.0 genes in this pathway |
| ZHU CMV 8 HR DN | 53 | 40 | All SZGR 2.0 genes in this pathway |
| TAKAO RESPONSE TO UVB RADIATION DN | 98 | 67 | All SZGR 2.0 genes in this pathway |
| ZHU CMV ALL DN | 128 | 93 | All SZGR 2.0 genes in this pathway |
| ZHANG BREAST CANCER PROGENITORS UP | 425 | 253 | All SZGR 2.0 genes in this pathway |
| YAUCH HEDGEHOG SIGNALING PARACRINE DN | 264 | 159 | All SZGR 2.0 genes in this pathway |
| HOSHIDA LIVER CANCER SUBCLASS S1 | 237 | 159 | All SZGR 2.0 genes in this pathway |
| WONG ADULT TISSUE STEM MODULE | 721 | 492 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS DN | 1972 | 1213 | All SZGR 2.0 genes in this pathway |
| RAO BOUND BY SALL4 | 227 | 149 | All SZGR 2.0 genes in this pathway |