Gene Page: RAB7A
Summary ?
| GeneID | 7879 |
| Symbol | RAB7A |
| Synonyms | PRO2706|RAB7 |
| Description | RAB7A, member RAS oncogene family |
| Reference | MIM:602298|HGNC:HGNC:9788|Ensembl:ENSG00000075785|HPRD:03805|Vega:OTTHUMG00000159812 |
| Gene type | protein-coding |
| Map location | 3q21.3 |
| Pascal p-value | 0.802 |
| Sherlock p-value | 0.414 |
| Fetal beta | -0.421 |
| DMG | 1 (# studies) |
| eGene | Cerebellum Meta |
| Support | INTRACELLULAR TRAFFICKING G2Cdb.human_BAYES-COLLINS-HUMAN-PSD-CONSENSUS CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg13284070 | 3 | 128483832 | RAB7A | 1.525E-4 | 0.475 | 0.032 | DMG:Wockner_2014 |
Section II. Transcriptome annotation
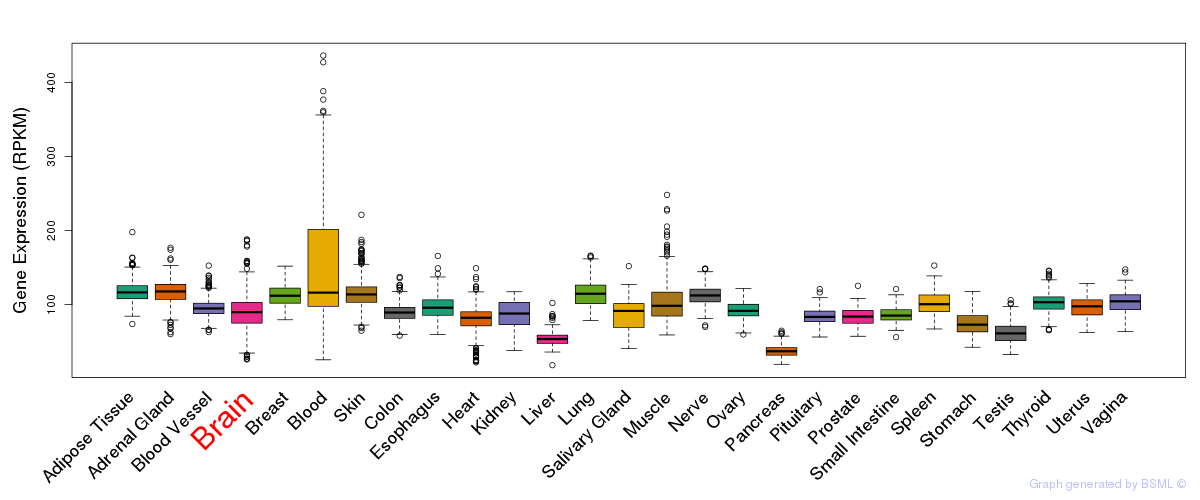
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| CHM | DXS540 | FLJ38564 | GGTA | HSD-32 | MGC102710 | REP-1 | TCD | choroideremia (Rab escort protein 1) | - | HPRD,BioGRID | 9563513 |12576024 |
| EIF1B | GC20 | eukaryotic translation initiation factor 1B | Affinity Capture-MS | BioGRID | 17353931 |
| RAB17 | FLJ12538 | RAB17, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB22A | MGC16770 | RAB22A, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB4A | HRES-1/RAB4 | RAB4 | RAB4A, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB4B | FLJ78649 | MGC52123 | RAB4B, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB5A | RAB5 | RAB5A, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB5B | - | RAB5B, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RAB6A | RAB6 | RAB6A' | RAB6B | RAB6A, member RAS oncogene family | Two-hybrid | BioGRID | 10329441 |
| RABAC1 | PRA1 | PRAF1 | YIP3 | Rab acceptor 1 (prenylated) | - | HPRD | 10329441 |
| RHEB | MGC111559 | RHEB2 | Ras homolog enriched in brain | Co-localization | BioGRID | 15809346 |
| RILP | FLJ31193 | PP10141 | Rab interacting lysosomal protein | - | HPRD,BioGRID | 11179213 |
| RNF115 | BCA2 | ZNF364 | ring finger protein 115 | - | HPRD | 12972561 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| BIOCARTA RAB PATHWAY | 12 | 8 | All SZGR 2.0 genes in this pathway |
| PID IL12 2PATHWAY | 63 | 54 | All SZGR 2.0 genes in this pathway |
| PID AJDISS 2PATHWAY | 48 | 38 | All SZGR 2.0 genes in this pathway |
| PID IL8 CXCR2 PATHWAY | 34 | 26 | All SZGR 2.0 genes in this pathway |
| REACTOME MHC CLASS II ANTIGEN PRESENTATION | 91 | 61 | All SZGR 2.0 genes in this pathway |
| REACTOME IMMUNE SYSTEM | 933 | 616 | All SZGR 2.0 genes in this pathway |
| REACTOME ADAPTIVE IMMUNE SYSTEM | 539 | 350 | All SZGR 2.0 genes in this pathway |
| ONKEN UVEAL MELANOMA DN | 526 | 357 | All SZGR 2.0 genes in this pathway |
| BIDUS METASTASIS UP | 214 | 134 | All SZGR 2.0 genes in this pathway |
| CHIARADONNA NEOPLASTIC TRANSFORMATION KRAS CDC25 UP | 58 | 39 | All SZGR 2.0 genes in this pathway |
| RIZ ERYTHROID DIFFERENTIATION | 77 | 51 | All SZGR 2.0 genes in this pathway |
| RIZ ERYTHROID DIFFERENTIATION CCNE1 | 40 | 26 | All SZGR 2.0 genes in this pathway |
| RASHI RESPONSE TO IONIZING RADIATION 4 | 61 | 34 | All SZGR 2.0 genes in this pathway |
| MARTORIATI MDM4 TARGETS FETAL LIVER DN | 514 | 319 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS UP | 811 | 508 | All SZGR 2.0 genes in this pathway |
| SHEN SMARCA2 TARGETS UP | 424 | 268 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS LATE PROGENITOR | 544 | 307 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE DN | 1237 | 837 | All SZGR 2.0 genes in this pathway |
| BURTON ADIPOGENESIS 8 | 86 | 57 | All SZGR 2.0 genes in this pathway |
| WELCSH BRCA1 TARGETS UP | 198 | 132 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE BY UV | 62 | 43 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE BY 4NQO OR UV | 63 | 44 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE DN | 195 | 135 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE UP | 226 | 164 | All SZGR 2.0 genes in this pathway |
| IWANAGA CARCINOGENESIS BY KRAS PTEN UP | 181 | 112 | All SZGR 2.0 genes in this pathway |
| WANG TUMOR INVASIVENESS UP | 374 | 247 | All SZGR 2.0 genes in this pathway |
| YAO TEMPORAL RESPONSE TO PROGESTERONE CLUSTER 7 | 76 | 46 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 3 UP | 430 | 288 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS UP | 682 | 433 | All SZGR 2.0 genes in this pathway |