Gene Page: NSUN7
Summary ?
| GeneID | 79730 |
| Symbol | NSUN7 |
| Synonyms | - |
| Description | NOP2/Sun RNA methyltransferase family member 7 |
| Reference | HGNC:HGNC:25857|Ensembl:ENSG00000179299|HPRD:07846|Vega:OTTHUMG00000128597 |
| Gene type | protein-coding |
| Map location | 4p14 |
| Pascal p-value | 0.42 |
| Fetal beta | 2.552 |
| DMG | 1 (# studies) |
| eGene | Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:vanEijk_2014 | Genome-wide DNA methylation analysis | This dataset includes 432 differentially methylated CpG sites corresponding to 391 unique transcripts between schizophrenia patients (n=260) and unaffected controls (n=250). | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg19786920 | 4 | 40751770 | NSUN7 | 0.015 | -3.194 | DMG:vanEijk_2014 |
Section II. Transcriptome annotation
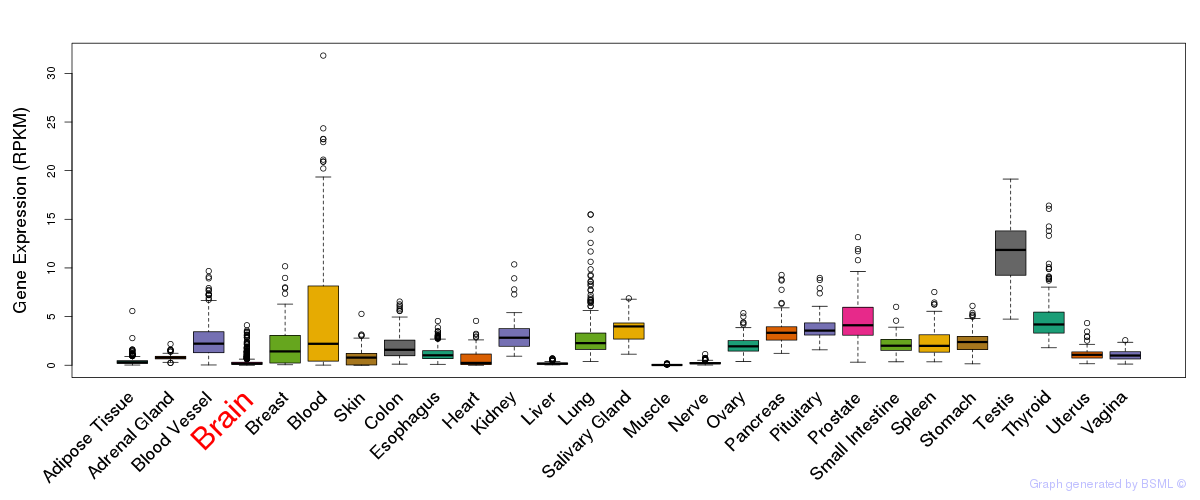
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZCCHC24 | 0.92 | 0.94 |
| SASH1 | 0.91 | 0.88 |
| KCNJ10 | 0.89 | 0.93 |
| TPCN1 | 0.88 | 0.90 |
| ITPKB | 0.88 | 0.92 |
| KANK1 | 0.87 | 0.89 |
| TNS1 | 0.86 | 0.89 |
| ATP1A2 | 0.86 | 0.90 |
| TPP1 | 0.85 | 0.89 |
| EPAS1 | 0.85 | 0.91 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| POLB | -0.62 | -0.74 |
| NR2C2AP | -0.61 | -0.71 |
| TRNAU1AP | -0.60 | -0.68 |
| MED19 | -0.60 | -0.68 |
| ING4 | -0.59 | -0.66 |
| FRG1 | -0.59 | -0.74 |
| DUSP12 | -0.58 | -0.64 |
| STMN1 | -0.58 | -0.63 |
| OCIAD2 | -0.58 | -0.67 |
| COMMD3 | -0.58 | -0.71 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| CHEMNITZ RESPONSE TO PROSTAGLANDIN E2 DN | 391 | 222 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| NUYTTEN EZH2 TARGETS DN | 1024 | 594 | All SZGR 2.0 genes in this pathway |
| ZHAN MULTIPLE MYELOMA CD1 UP | 45 | 29 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 3 | 720 | 440 | All SZGR 2.0 genes in this pathway |
| MATZUK SPERMATOZOA | 114 | 77 | All SZGR 2.0 genes in this pathway |
| HAN SATB1 TARGETS UP | 395 | 249 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3K4ME2 | 491 | 319 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |