Gene Page: SNIP1
Summary ?
| GeneID | 79753 |
| Symbol | SNIP1 |
| Synonyms | PMRED |
| Description | Smad nuclear interacting protein 1 |
| Reference | MIM:608241|HGNC:HGNC:30587|Ensembl:ENSG00000163877|HPRD:06999|Vega:OTTHUMG00000004225 |
| Gene type | protein-coding |
| Map location | 1p34.3 |
| Pascal p-value | 0.316 |
| Sherlock p-value | 0.563 |
| Fetal beta | 0.978 |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs16959318 | chr16 | 12457250 | SNIP1 | 79753 | 0.11 | trans |
Section II. Transcriptome annotation
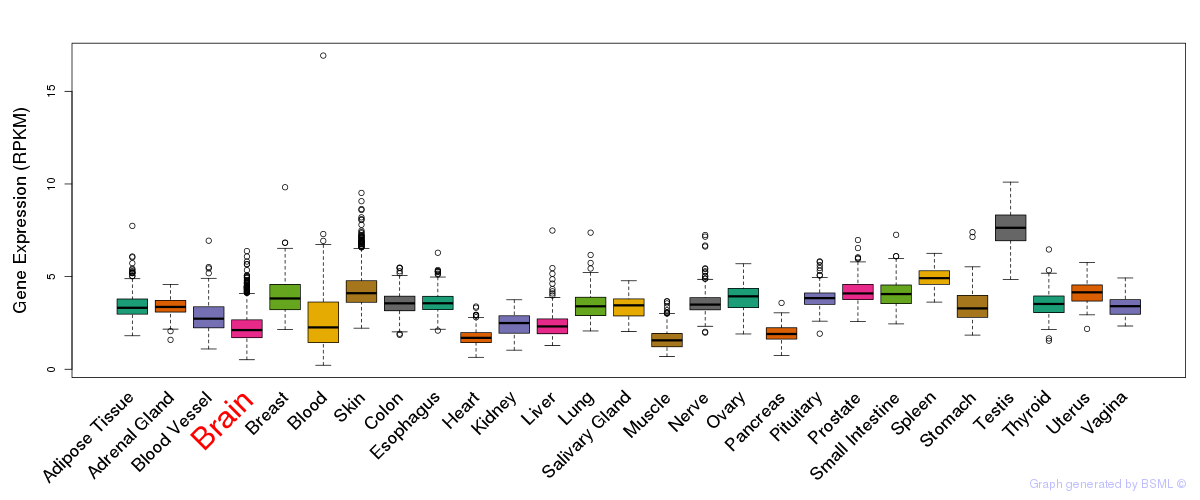
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MARCH2 | 0.85 | 0.76 |
| ISOC2 | 0.85 | 0.81 |
| ASPDH | 0.85 | 0.75 |
| C2orf7 | 0.85 | 0.77 |
| C11orf59 | 0.84 | 0.75 |
| APOA1BP | 0.84 | 0.74 |
| TMEM219 | 0.84 | 0.77 |
| CISD3 | 0.83 | 0.73 |
| COX5B | 0.82 | 0.76 |
| COX4I1 | 0.82 | 0.72 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| RBM25 | -0.51 | -0.57 |
| ZNF326 | -0.51 | -0.55 |
| THOC2 | -0.48 | -0.55 |
| ANKRD12 | -0.47 | -0.50 |
| RTF1 | -0.46 | -0.48 |
| C5orf42 | -0.46 | -0.45 |
| ARID4A | -0.46 | -0.45 |
| AGTPBP1 | -0.46 | -0.47 |
| SFRS12 | -0.46 | -0.49 |
| EIF5B | -0.46 | -0.55 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID SMAD2 3NUCLEAR PATHWAY | 82 | 63 | All SZGR 2.0 genes in this pathway |
| LASTOWSKA NEUROBLASTOMA COPY NUMBER DN | 800 | 473 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS UP | 811 | 508 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RESPONSE TO TRABECTEDIN DN | 271 | 175 | All SZGR 2.0 genes in this pathway |
| GRADE COLON CANCER UP | 871 | 505 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |