Gene Page: FBXO31
Summary ?
| GeneID | 79791 |
| Symbol | FBXO31 |
| Synonyms | FBX14|FBXO14|Fbx31|MRT45|pp2386 |
| Description | F-box protein 31 |
| Reference | MIM:609102|HGNC:HGNC:16510|Ensembl:ENSG00000103264|HPRD:10954|Vega:OTTHUMG00000175685 |
| Gene type | protein-coding |
| Map location | 16q24.2 |
| Pascal p-value | 0.031 |
| Sherlock p-value | 0.672 |
| eGene | Cerebellar Hemisphere Cerebellum Myers' cis & trans |
| Support | G2Cdb.humanPSD G2Cdb.humanPSP CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| FBXO31 | chr16 | 87365049 | T | G | NM_024735 NR_024568 | p.489I>L . | missense npcRNA | Schizophrenia | DNM:Fromer_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs16829545 | chr2 | 151977407 | FBXO31 | 79791 | 0 | trans | ||
| rs3845734 | chr2 | 171125572 | FBXO31 | 79791 | 0.01 | trans | ||
| rs7584986 | chr2 | 184111432 | FBXO31 | 79791 | 0.12 | trans | ||
| rs17762315 | chr5 | 76807576 | FBXO31 | 79791 | 0.04 | trans | ||
| rs16955618 | chr15 | 29937543 | FBXO31 | 79791 | 3.322E-6 | trans | ||
| rs1041786 | chr21 | 22617710 | FBXO31 | 79791 | 0.17 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
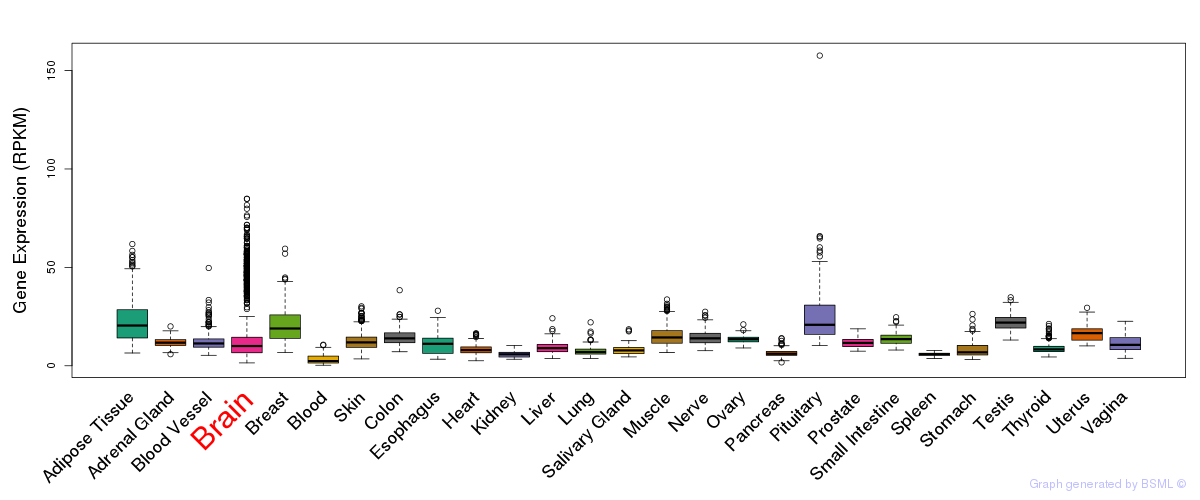
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DOANE BREAST CANCER CLASSES DN | 34 | 26 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| BENPORATH NANOG TARGETS | 988 | 594 | All SZGR 2.0 genes in this pathway |
| GEORGES TARGETS OF MIR192 AND MIR215 | 893 | 528 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER BASAL UP | 648 | 398 | All SZGR 2.0 genes in this pathway |
| LEE LIVER CANCER SURVIVAL UP | 185 | 112 | All SZGR 2.0 genes in this pathway |
| PURBEY TARGETS OF CTBP1 NOT SATB1 DN | 448 | 282 | All SZGR 2.0 genes in this pathway |
| WAKABAYASHI ADIPOGENESIS PPARG RXRA BOUND 8D | 882 | 506 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |