Gene Page: HKDC1
Summary ?
| GeneID | 80201 |
| Symbol | HKDC1 |
| Synonyms | - |
| Description | hexokinase domain containing 1 |
| Reference | HGNC:HGNC:23302|Ensembl:ENSG00000156510|HPRD:08006|Vega:OTTHUMG00000018371 |
| Gene type | protein-coding |
| Map location | 10q22.1 |
| Pascal p-value | 0.356 |
| DMG | 1 (# studies) |
| eGene | Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg11639651 | 10 | 70979811 | HKDC1 | 3.106E-4 | -0.491 | 0.04 | DMG:Wockner_2014 |
Section II. Transcriptome annotation
General gene expression (GTEx)
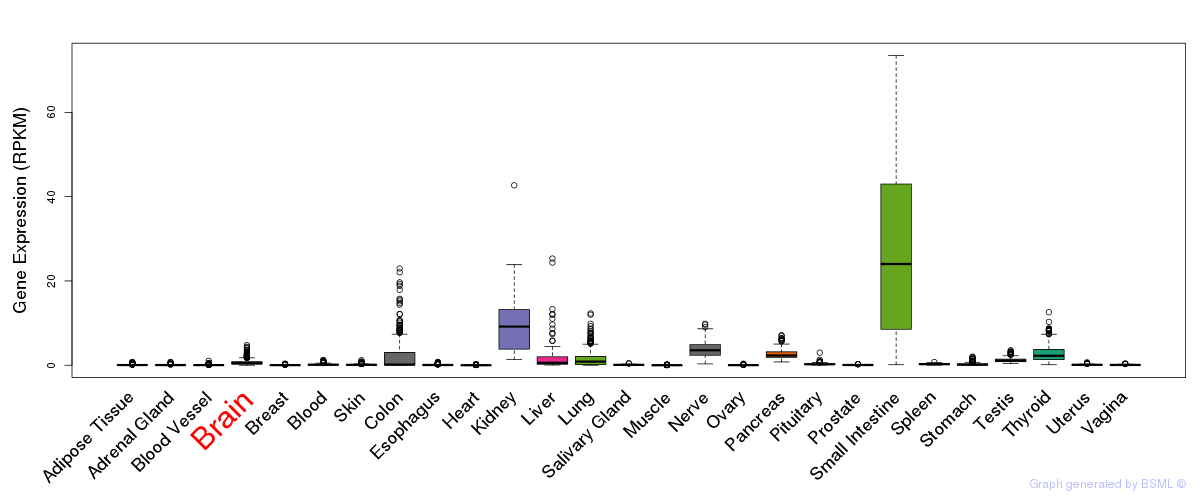
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SH3GLB1 | 0.83 | 0.82 |
| RNF180 | 0.83 | 0.82 |
| B3GALNT1 | 0.82 | 0.81 |
| DERL1 | 0.80 | 0.82 |
| SMARCA1 | 0.80 | 0.83 |
| HMGCS1 | 0.79 | 0.84 |
| FBXO3 | 0.79 | 0.82 |
| PKIA | 0.79 | 0.80 |
| BIRC2 | 0.78 | 0.82 |
| MGAT4C | 0.78 | 0.73 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.60 | -0.70 |
| AF347015.31 | -0.57 | -0.68 |
| AF347015.8 | -0.57 | -0.68 |
| AF347015.27 | -0.57 | -0.65 |
| AF347015.33 | -0.57 | -0.65 |
| MT-CYB | -0.56 | -0.67 |
| FXYD1 | -0.54 | -0.62 |
| AF347015.2 | -0.54 | -0.68 |
| HLA-F | -0.54 | -0.58 |
| AF347015.15 | -0.54 | -0.66 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE LIVE DN | 384 | 220 | All SZGR 2.0 genes in this pathway |
| DEURIG T CELL PROLYMPHOCYTIC LEUKEMIA UP | 368 | 234 | All SZGR 2.0 genes in this pathway |
| TIEN INTESTINE PROBIOTICS 24HR DN | 214 | 133 | All SZGR 2.0 genes in this pathway |
| LIAO METASTASIS | 539 | 324 | All SZGR 2.0 genes in this pathway |
| SCHAEFFER PROSTATE DEVELOPMENT 48HR UP | 487 | 286 | All SZGR 2.0 genes in this pathway |
| CERVERA SDHB TARGETS 2 | 114 | 76 | All SZGR 2.0 genes in this pathway |
| HENDRICKS SMARCA4 TARGETS UP | 56 | 35 | All SZGR 2.0 genes in this pathway |
| ACEVEDO METHYLATED IN LIVER CANCER DN | 940 | 425 | All SZGR 2.0 genes in this pathway |
| WOO LIVER CANCER RECURRENCE UP | 105 | 75 | All SZGR 2.0 genes in this pathway |
| VANDESLUIS COMMD1 TARGETS GROUP 3 UP | 89 | 50 | All SZGR 2.0 genes in this pathway |