Gene Page: LRRC27
Summary ?
| GeneID | 80313 |
| Symbol | LRRC27 |
| Synonyms | - |
| Description | leucine rich repeat containing 27 |
| Reference | HGNC:HGNC:29346|Ensembl:ENSG00000148814|HPRD:17451|Vega:OTTHUMG00000019284 |
| Gene type | protein-coding |
| Map location | 10q26.3 |
| Pascal p-value | 0.802 |
| Sherlock p-value | 0.92 |
| Fetal beta | 0.561 |
| DMG | 1 (# studies) |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cortex Frontal Cortex BA9 Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
| GSMA_IIA | Genome scan meta-analysis (All samples) | Psr: 0.03487 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg27016988 | 10 | 134186967 | LRRC27 | 3.635E-4 | 0.246 | 0.042 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs12269628 | 10 | 134130159 | LRRC27 | ENSG00000148814.13 | 2.863E-8 | 0 | -15455 | gtex_brain_ba24 |
| rs76502537 | 10 | 134139068 | LRRC27 | ENSG00000148814.13 | 7.153E-8 | 0 | -6546 | gtex_brain_ba24 |
| rs6560691 | 10 | 134140245 | LRRC27 | ENSG00000148814.13 | 7.371E-8 | 0 | -5369 | gtex_brain_ba24 |
| rs7085852 | 10 | 134144005 | LRRC27 | ENSG00000148814.13 | 1.116E-8 | 0 | -1609 | gtex_brain_ba24 |
| rs76805118 | 10 | 134144739 | LRRC27 | ENSG00000148814.13 | 3.181E-9 | 0 | -875 | gtex_brain_ba24 |
| rs3923974 | 10 | 134146084 | LRRC27 | ENSG00000148814.13 | 3.181E-9 | 0 | 470 | gtex_brain_ba24 |
| rs2295273 | 10 | 134150920 | LRRC27 | ENSG00000148814.13 | 3.181E-9 | 0 | 5306 | gtex_brain_ba24 |
| rs7912110 | 10 | 134152214 | LRRC27 | ENSG00000148814.13 | 2.239E-9 | 0 | 6600 | gtex_brain_ba24 |
| rs7098749 | 10 | 134157220 | LRRC27 | ENSG00000148814.13 | 1.554E-10 | 0 | 11606 | gtex_brain_ba24 |
| rs11146339 | 10 | 134158448 | LRRC27 | ENSG00000148814.13 | 2.934E-9 | 0 | 12834 | gtex_brain_ba24 |
| rs4612718 | 10 | 134158940 | LRRC27 | ENSG00000148814.13 | 1.78E-9 | 0 | 13326 | gtex_brain_ba24 |
| rs55813610 | 10 | 134163693 | LRRC27 | ENSG00000148814.13 | 2.431E-7 | 0 | 18079 | gtex_brain_ba24 |
| rs7090409 | 10 | 134164622 | LRRC27 | ENSG00000148814.13 | 6.034E-9 | 0 | 19008 | gtex_brain_ba24 |
| rs56678407 | 10 | 134167502 | LRRC27 | ENSG00000148814.13 | 1.38E-9 | 0 | 21888 | gtex_brain_ba24 |
Section II. Transcriptome annotation
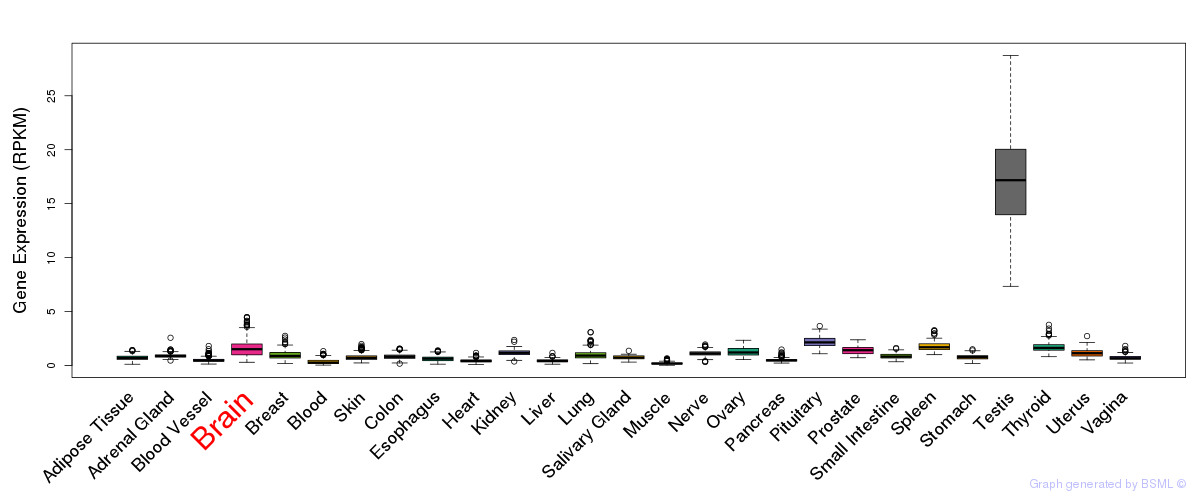
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005515 | protein binding | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 | 473 | 224 | All SZGR 2.0 genes in this pathway |
| LEIN MEDULLA MARKERS | 81 | 48 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| COATES MACROPHAGE M1 VS M2 DN | 78 | 44 | All SZGR 2.0 genes in this pathway |
| CHEN METABOLIC SYNDROM NETWORK | 1210 | 725 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP B | 549 | 316 | All SZGR 2.0 genes in this pathway |
| YANG BCL3 TARGETS UP | 364 | 236 | All SZGR 2.0 genes in this pathway |