Gene Page: OR5V1
Summary ?
| GeneID | 81696 |
| Symbol | OR5V1 |
| Synonyms | 6M1-21|hs6M1-21 |
| Description | olfactory receptor family 5 subfamily V member 1 |
| Reference | HGNC:HGNC:13972|Ensembl:ENSG00000243729|HPRD:17771| |
| Gene type | protein-coding |
| Map location | 6p22.1 |
| Pascal p-value | 1E-12 |
| Fetal beta | -0.187 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| GSMA_I | Genome scan meta-analysis | Psr: 0.033 |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
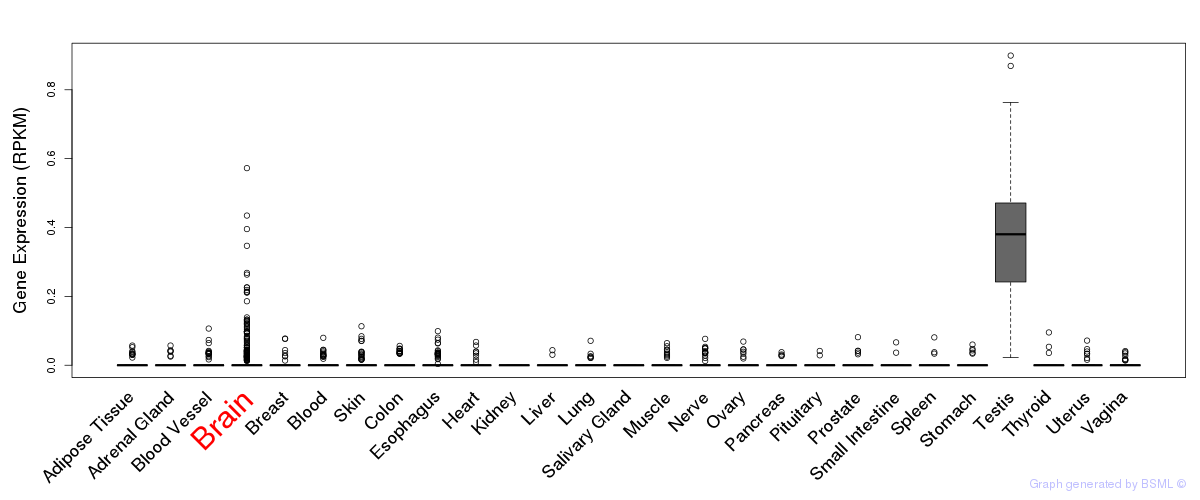
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZBTB44 | 0.93 | 0.95 |
| CCNT1 | 0.92 | 0.94 |
| KIAA1219 | 0.91 | 0.94 |
| TAOK1 | 0.91 | 0.91 |
| GSK3B | 0.90 | 0.93 |
| RC3H2 | 0.90 | 0.94 |
| RLIM | 0.90 | 0.94 |
| AVL9 | 0.90 | 0.94 |
| AFF4 | 0.90 | 0.92 |
| BMPR2 | 0.90 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| FXYD1 | -0.66 | -0.77 |
| MT-CO2 | -0.63 | -0.76 |
| HSD17B14 | -0.63 | -0.71 |
| HIGD1B | -0.63 | -0.75 |
| TLCD1 | -0.62 | -0.72 |
| AF347015.31 | -0.62 | -0.73 |
| AF347015.8 | -0.62 | -0.75 |
| CST3 | -0.62 | -0.73 |
| AF347015.33 | -0.62 | -0.71 |
| MT-CYB | -0.61 | -0.72 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0004872 | receptor activity | IEA | - | |
| GO:0004984 | olfactory receptor activity | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0007186 | G-protein coupled receptor protein signaling pathway | IEA | - | |
| GO:0007165 | signal transduction | IEA | - | |
| GO:0007608 | sensory perception of smell | IEA | - | |
| GO:0050896 | response to stimulus | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0016020 | membrane | IEA | - | |
| GO:0016021 | integral to membrane | IEA | - | |
| GO:0005886 | plasma membrane | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG OLFACTORY TRANSDUCTION | 389 | 85 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALING BY GPCR | 920 | 449 | All SZGR 2.0 genes in this pathway |
| REACTOME OLFACTORY SIGNALING PATHWAY | 328 | 60 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR DOWNSTREAM SIGNALING | 805 | 368 | All SZGR 2.0 genes in this pathway |
| ODONNELL TFRC TARGETS UP | 456 | 228 | All SZGR 2.0 genes in this pathway |
| KONDO PROSTATE CANCER WITH H3K27ME3 | 196 | 93 | All SZGR 2.0 genes in this pathway |