Gene Page: F8A1
Summary ?
| GeneID | 8263 |
| Symbol | F8A1 |
| Synonyms | DXS522E|F8A|HAP40 |
| Description | coagulation factor VIII-associated 1 |
| Reference | MIM:305423|HGNC:HGNC:3547|Ensembl:ENSG00000277203|HPRD:02374|Vega:OTTHUMG00000013500 |
| Gene type | protein-coding |
| Map location | Xq28 |
| Sherlock p-value | 0.302 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
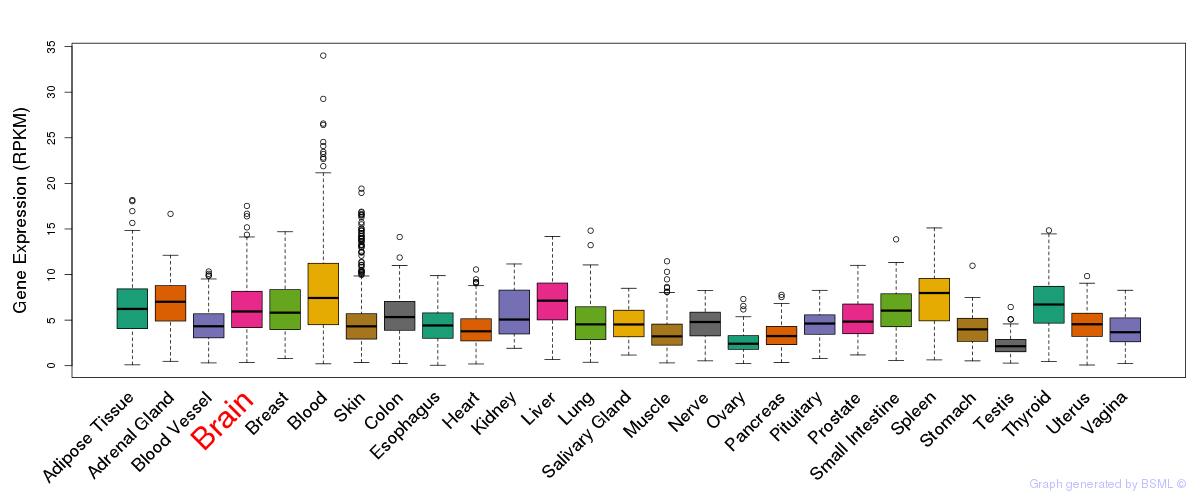
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| DAG1 | 0.79 | 0.84 |
| NCAN | 0.76 | 0.83 |
| POFUT1 | 0.76 | 0.81 |
| CYFIP1 | 0.75 | 0.80 |
| TMEM43 | 0.74 | 0.80 |
| SARM1 | 0.74 | 0.79 |
| FAM57A | 0.74 | 0.78 |
| PTPN21 | 0.73 | 0.79 |
| SALL2 | 0.73 | 0.75 |
| MCART6 | 0.73 | 0.76 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C5orf53 | -0.54 | -0.59 |
| AF347015.27 | -0.54 | -0.62 |
| MT-CO2 | -0.53 | -0.63 |
| AF347015.33 | -0.53 | -0.60 |
| AF347015.31 | -0.53 | -0.62 |
| AF347015.21 | -0.53 | -0.68 |
| S100B | -0.52 | -0.58 |
| AC021016.1 | -0.51 | -0.60 |
| AF347015.8 | -0.51 | -0.62 |
| FXYD1 | -0.51 | -0.56 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| WOOD EBV EBNA1 TARGETS UP | 110 | 71 | All SZGR 2.0 genes in this pathway |
| FERREIRA EWINGS SARCOMA UNSTABLE VS STABLE DN | 98 | 59 | All SZGR 2.0 genes in this pathway |
| LOPEZ MBD TARGETS | 957 | 597 | All SZGR 2.0 genes in this pathway |
| KUMAR TARGETS OF MLL AF9 FUSION | 405 | 264 | All SZGR 2.0 genes in this pathway |
| MCCABE BOUND BY HOXC6 | 469 | 239 | All SZGR 2.0 genes in this pathway |
| ACEVEDO NORMAL TISSUE ADJACENT TO LIVER TUMOR DN | 354 | 216 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| JIANG TIP30 TARGETS DN | 24 | 14 | All SZGR 2.0 genes in this pathway |
| ZHANG TLX TARGETS 36HR UP | 221 | 150 | All SZGR 2.0 genes in this pathway |
| PURBEY TARGETS OF CTBP1 NOT SATB1 DN | 448 | 282 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |