Gene Page: KRT36
Summary ?
| GeneID | 8689 |
| Symbol | KRT36 |
| Synonyms | HA6|KRTHA6|hHa6 |
| Description | keratin 36 |
| Reference | MIM:604540|HGNC:HGNC:6454|Ensembl:ENSG00000126337|HPRD:05174|Vega:OTTHUMG00000133431 |
| Gene type | protein-coding |
| Map location | 17q21.2 |
| Pascal p-value | 0.446 |
| Fetal beta | 0.051 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
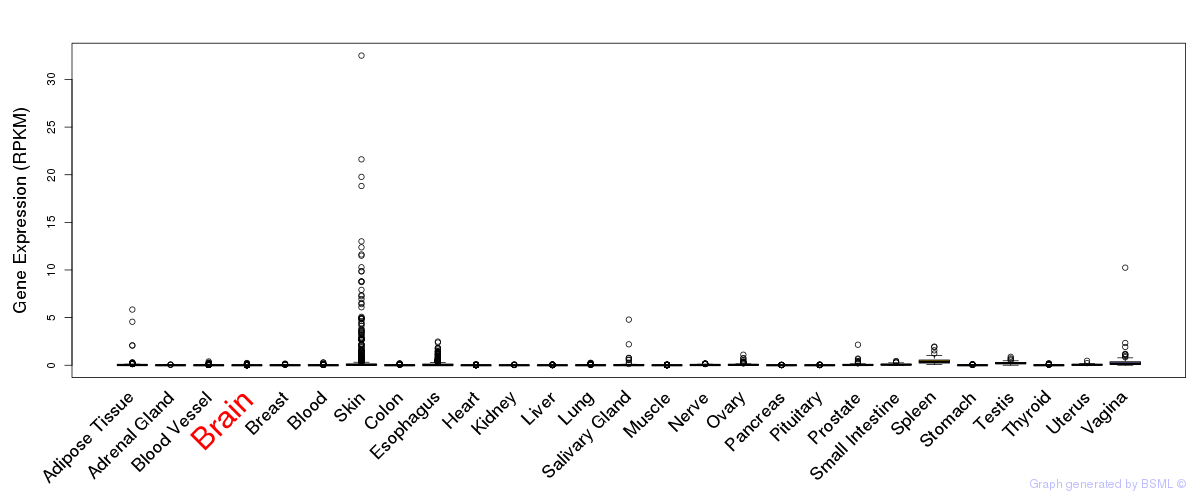
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PARP6 | 0.95 | 0.94 |
| FAM40A | 0.95 | 0.93 |
| BAT3 | 0.95 | 0.95 |
| C7orf26 | 0.94 | 0.92 |
| PRCC | 0.94 | 0.93 |
| FGD1 | 0.94 | 0.95 |
| RAF1 | 0.94 | 0.93 |
| MARK3 | 0.94 | 0.92 |
| PJA1 | 0.94 | 0.93 |
| ISG20L2 | 0.94 | 0.91 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.83 | -0.91 |
| AF347015.27 | -0.83 | -0.90 |
| AF347015.31 | -0.82 | -0.89 |
| AF347015.33 | -0.81 | -0.89 |
| AF347015.8 | -0.80 | -0.90 |
| MT-CYB | -0.80 | -0.88 |
| HLA-F | -0.78 | -0.79 |
| C5orf53 | -0.77 | -0.78 |
| S100B | -0.77 | -0.84 |
| AIFM3 | -0.77 | -0.78 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE LIVE UP | 485 | 293 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE FIMA UP | 544 | 308 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LPS UP | 431 | 237 | All SZGR 2.0 genes in this pathway |
| TANAKA METHYLATED IN ESOPHAGEAL CARCINOMA | 103 | 58 | All SZGR 2.0 genes in this pathway |
| DURCHDEWALD SKIN CARCINOGENESIS UP | 88 | 47 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS UP | 673 | 430 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS DN | 593 | 372 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS DN | 591 | 366 | All SZGR 2.0 genes in this pathway |
| CAMPS COLON CANCER COPY NUMBER UP | 92 | 45 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |
| WANG NFKB TARGETS | 25 | 15 | All SZGR 2.0 genes in this pathway |