Gene Page: DNAH11
Summary ?
| GeneID | 8701 |
| Symbol | DNAH11 |
| Synonyms | CILD7|DNAHBL|DNAHC11|DNHBL|DPL11 |
| Description | dynein axonemal heavy chain 11 |
| Reference | MIM:603339|HGNC:HGNC:2942|Ensembl:ENSG00000105877|HPRD:09134|Vega:OTTHUMG00000152524 |
| Gene type | protein-coding |
| Map location | 7p21 |
| Pascal p-value | 0.138 |
| Fetal beta | -0.331 |
| eGene | Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| CV:PheWAS | Phenome-wide association studies (PheWAS) | 157 SNPs associated with schizophrenia | 1 |
Section I. Genetics and epigenetics annotation
 CV:PheWAS
CV:PheWAS
| SNP ID | Chromosome | Position | Allele | P | Function | Gene | Up/Down Distance |
|---|---|---|---|---|---|---|---|
| rs4487645 | 7 | 0 | null | 1.441 | DNAH11 | null |
Section II. Transcriptome annotation
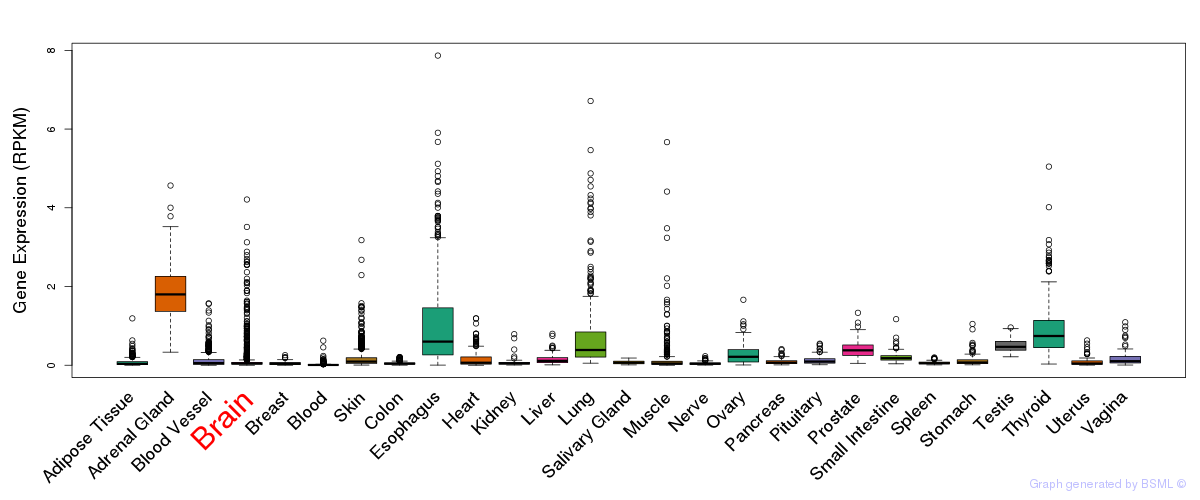
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PHF20 | 0.95 | 0.96 |
| ZMYM4 | 0.95 | 0.95 |
| HMG20A | 0.95 | 0.97 |
| CAPRIN1 | 0.95 | 0.96 |
| WDR48 | 0.95 | 0.96 |
| EPM2AIP1 | 0.95 | 0.96 |
| SPAST | 0.95 | 0.96 |
| DDX21 | 0.95 | 0.95 |
| VPRBP | 0.95 | 0.96 |
| CTCF | 0.94 | 0.96 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.75 | -0.88 |
| AF347015.31 | -0.75 | -0.87 |
| FXYD1 | -0.74 | -0.87 |
| IFI27 | -0.74 | -0.86 |
| AF347015.27 | -0.73 | -0.85 |
| AF347015.33 | -0.72 | -0.83 |
| HIGD1B | -0.72 | -0.86 |
| MT-CYB | -0.72 | -0.84 |
| AF347015.8 | -0.71 | -0.86 |
| C5orf53 | -0.70 | -0.74 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 | 473 | 224 | All SZGR 2.0 genes in this pathway |
| MEISSNER BRAIN HCP WITH H3K4ME3 AND H3K27ME3 | 1069 | 729 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3K4ME2 AND H3K27ME3 | 349 | 234 | All SZGR 2.0 genes in this pathway |