Gene Page: MBD2
Summary ?
| GeneID | 8932 |
| Symbol | MBD2 |
| Synonyms | DMTase|NY-CO-41 |
| Description | methyl-CpG binding domain protein 2 |
| Reference | MIM:603547|HGNC:HGNC:6917|Ensembl:ENSG00000134046|HPRD:04647|Vega:OTTHUMG00000132705 |
| Gene type | protein-coding |
| Map location | 18q21 |
| Pascal p-value | 0.351 |
| Fetal beta | -0.554 |
| Support | Chromatin Remodeling Genes |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
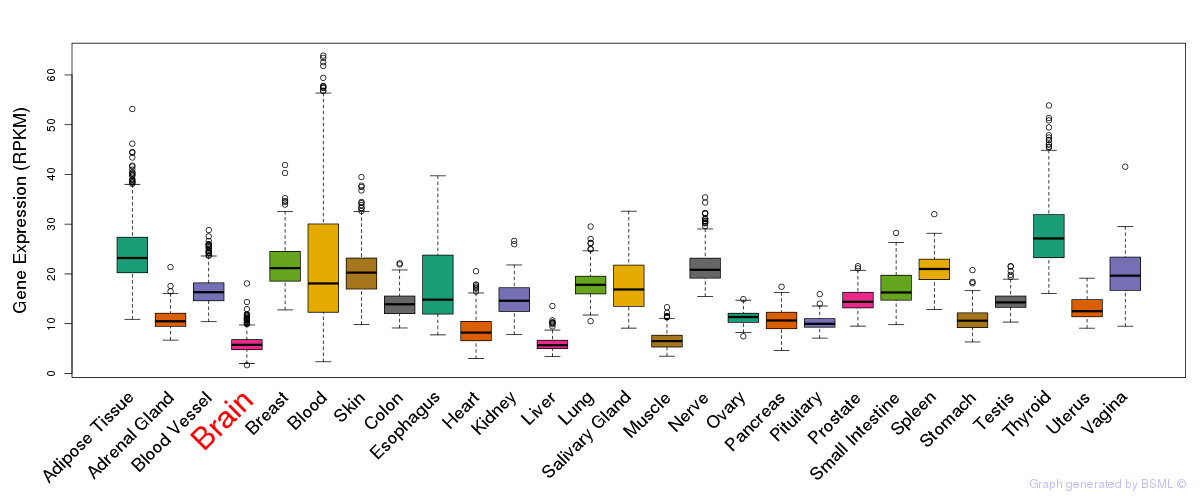
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| FRZB | 0.55 | 0.50 |
| GDF10 | 0.55 | 0.55 |
| GLT8D2 | 0.54 | 0.53 |
| HCRTR1 | 0.51 | 0.49 |
| TM6SF1 | 0.51 | 0.41 |
| CYP1A1 | 0.50 | 0.51 |
| TFR2 | 0.49 | 0.50 |
| OR2W3 | 0.48 | 0.43 |
| AC132186.2 | 0.48 | 0.49 |
| CCDC3 | 0.47 | 0.44 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C11orf67 | -0.23 | -0.22 |
| CHMP4A | -0.23 | -0.23 |
| NKAPL | -0.22 | -0.20 |
| AF347015.21 | -0.22 | -0.17 |
| MT-ATP8 | -0.22 | -0.14 |
| ANP32B | -0.22 | -0.22 |
| AF347015.27 | -0.22 | -0.16 |
| SAP30 | -0.22 | -0.23 |
| AIF1L | -0.22 | -0.20 |
| AF347015.18 | -0.22 | -0.15 |
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| CHD3 | Mi-2a | Mi2-ALPHA | ZFH | chromodomain helicase DNA binding protein 3 | Reconstituted Complex | BioGRID | 10444591 |
| CREBBP | CBP | KAT3A | RSTS | CREB binding protein | - | HPRD | 12665568 |
| DHX9 | DDX9 | LKP | NDHII | RHA | DEAH (Asp-Glu-Ala-His) box polypeptide 9 | - | HPRD,BioGRID | 12665568 |
| DNMT1 | AIM | CXXC9 | DNMT | FLJ16293 | MCMT | MGC104992 | DNA (cytosine-5-)-methyltransferase 1 | - | HPRD | 10947852 |
| GATAD2A | FLJ20085 | FLJ21017 | p66alpha | GATA zinc finger domain containing 2A | - | HPRD,BioGRID | 12183469 |
| GATAD2B | FLJ37346 | KIAA1150 | MGC138257 | MGC138285 | P66beta | RP11-216N14.6 | GATA zinc finger domain containing 2B | - | HPRD,BioGRID | 11756549 |12183469 |
| GPN1 | ATPBD1A | MBDIN | NTPBP | XAB1 | GPN-loop GTPase 1 | - | HPRD,BioGRID | 12588985 |
| HDAC1 | DKFZp686H12203 | GON-10 | HD1 | RPD3 | RPD3L1 | histone deacetylase 1 | - | HPRD,BioGRID | 10471499 |
| HDAC2 | RPD3 | YAF1 | histone deacetylase 2 | - | HPRD,BioGRID | 10471499 |
| MBD2 | DKFZp586O0821 | DMTase | NY-CO-41 | methyl-CpG binding domain protein 2 | - | HPRD | 10947852 |
| MBD3 | - | methyl-CpG binding domain protein 3 | - | HPRD | 10947852 |
| MBD3 | - | methyl-CpG binding domain protein 3 | Reconstituted Complex Two-hybrid | BioGRID | 10444591 |15456747 |
| MBD3L1 | MBD3L | MGC138263 | MGC138269 | methyl-CpG binding domain protein 3-like 1 | - | HPRD,BioGRID | 15456747 |
| MIZF | DKFZp434F162 | HiNF-P | ZNF743 | MBD2-interacting zinc finger | - | HPRD,BioGRID | 11553631 |
| MTA2 | DKFZp686F2281 | MTA1L1 | PID | metastasis associated 1 family, member 2 | Reconstituted Complex | BioGRID | 10444591 |
| RBBP4 | NURF55 | RBAP48 | retinoblastoma binding protein 4 | - | HPRD,BioGRID | 10471499 |
| SIN3A | DKFZp434K2235 | FLJ90319 | KIAA0700 | SIN3 homolog A, transcription regulator (yeast) | Reconstituted Complex Two-hybrid | BioGRID | 10950960 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID HDAC CLASSI PATHWAY | 66 | 50 | All SZGR 2.0 genes in this pathway |
| REACTOME RNA POL I TRANSCRIPTION | 89 | 64 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSCRIPTION | 210 | 127 | All SZGR 2.0 genes in this pathway |
| REACTOME RNA POL I RNA POL III AND MITOCHONDRIAL TRANSCRIPTION | 122 | 80 | All SZGR 2.0 genes in this pathway |
| REACTOME RNA POL I PROMOTER OPENING | 62 | 49 | All SZGR 2.0 genes in this pathway |
| HOLLMANN APOPTOSIS VIA CD40 UP | 201 | 125 | All SZGR 2.0 genes in this pathway |
| ONKEN UVEAL MELANOMA UP | 783 | 507 | All SZGR 2.0 genes in this pathway |
| ENK UV RESPONSE KERATINOCYTE DN | 485 | 334 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN DN | 770 | 415 | All SZGR 2.0 genes in this pathway |
| SCHLOSSER SERUM RESPONSE DN | 712 | 443 | All SZGR 2.0 genes in this pathway |
| KORKOLA TERATOMA | 39 | 25 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 DN | 855 | 609 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 COMMON DN | 483 | 336 | All SZGR 2.0 genes in this pathway |
| OXFORD RALA AND RALB TARGETS UP | 10 | 7 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA1 PCC NETWORK | 1652 | 1023 | All SZGR 2.0 genes in this pathway |
| PUJANA ATM PCC NETWORK | 1442 | 892 | All SZGR 2.0 genes in this pathway |
| KAUFFMANN DNA REPAIR GENES | 230 | 137 | All SZGR 2.0 genes in this pathway |
| SHEPARD CRUSH AND BURN MUTANT DN | 185 | 111 | All SZGR 2.0 genes in this pathway |
| SHEPARD BMYB MORPHOLINO UP | 205 | 126 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS | 882 | 572 | All SZGR 2.0 genes in this pathway |
| ZHAN MULTIPLE MYELOMA LB UP | 48 | 28 | All SZGR 2.0 genes in this pathway |
| CROMER METASTASIS DN | 81 | 58 | All SZGR 2.0 genes in this pathway |
| GENTILE UV HIGH DOSE DN | 312 | 203 | All SZGR 2.0 genes in this pathway |
| DAZARD RESPONSE TO UV NHEK DN | 318 | 220 | All SZGR 2.0 genes in this pathway |
| DAZARD UV RESPONSE CLUSTER G6 | 153 | 112 | All SZGR 2.0 genes in this pathway |
| GENTILE RESPONSE CLUSTER D3 | 61 | 39 | All SZGR 2.0 genes in this pathway |
| FUJII YBX1 TARGETS UP | 43 | 28 | All SZGR 2.0 genes in this pathway |
| MUELLER PLURINET | 299 | 189 | All SZGR 2.0 genes in this pathway |
| SHEDDEN LUNG CANCER GOOD SURVIVAL A12 | 317 | 177 | All SZGR 2.0 genes in this pathway |
| SASSON RESPONSE TO GONADOTROPHINS DN | 87 | 66 | All SZGR 2.0 genes in this pathway |
| SASSON RESPONSE TO FORSKOLIN UP | 90 | 58 | All SZGR 2.0 genes in this pathway |
| PHESSE TARGETS OF APC AND MBD2 DN | 13 | 11 | All SZGR 2.0 genes in this pathway |
| CARD MIR302A TARGETS | 77 | 62 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 3 DN | 918 | 550 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |
| JUBAN TARGETS OF SPI1 AND FLI1 DN | 92 | 60 | All SZGR 2.0 genes in this pathway |
| KRIEG HYPOXIA NOT VIA KDM3A | 770 | 480 | All SZGR 2.0 genes in this pathway |