Gene Page: KIAA0125
Summary ?
| GeneID | 9834 |
| Symbol | KIAA0125 |
| Synonyms | C14orf110|FAM30A|HSPC053 |
| Description | KIAA0125 |
| Reference | MIM:616623|HGNC:HGNC:19955|Ensembl:ENSG00000226777|HPRD:13779| |
| Gene type | ncRNA |
| Map location | 14q32.33 |
| Fetal beta | 0.084 |
| DMG | 1 (# studies) |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| DMG:Montano_2016 | Genome-wide DNA methylation analysis | This dataset includes 172 replicated associations between CpGs with schizophrenia. | 2 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg04792777 | 14 | 106322429 | KIAA0125 | 1.06E-7 | -0.007 | 0.01 | DMG:Montano_2016 |
| cg21220247 | 14 | 106321936 | KIAA0125 | 1.77E-4 | -0.008 | 0.149 | DMG:Montano_2016 |
Section II. Transcriptome annotation
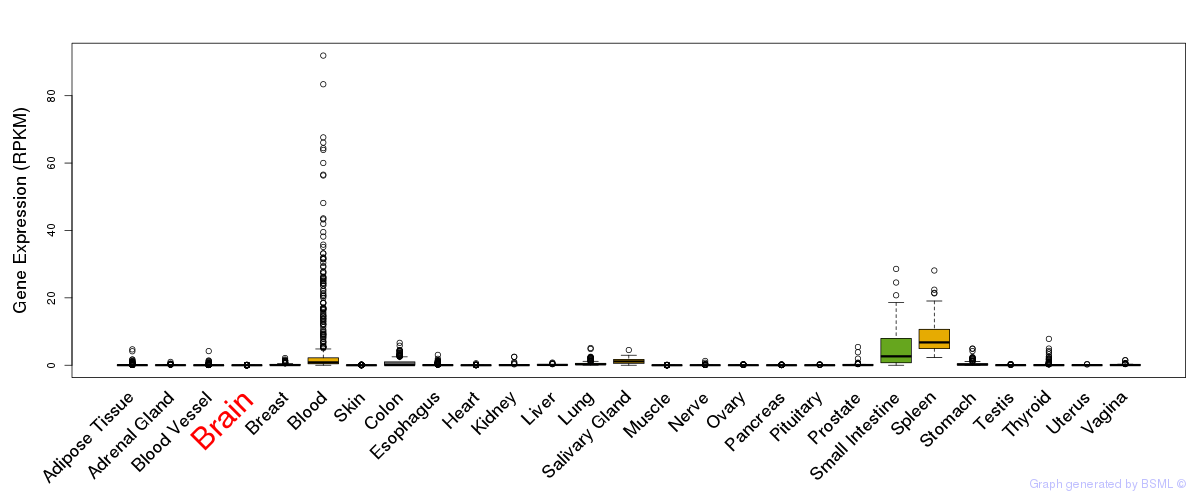
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ATP5L | 0.92 | 0.91 |
| C11orf73 | 0.89 | 0.89 |
| RNF181 | 0.89 | 0.86 |
| COX7A2 | 0.88 | 0.84 |
| EIF3K | 0.88 | 0.85 |
| TIMM8B | 0.88 | 0.85 |
| ATP5O | 0.88 | 0.86 |
| AC009171.1 | 0.88 | 0.86 |
| HINT1 | 0.87 | 0.80 |
| ZNF32 | 0.87 | 0.86 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MYH9 | -0.57 | -0.58 |
| TENC1 | -0.53 | -0.63 |
| RRBP1 | -0.52 | -0.63 |
| FAM38A | -0.52 | -0.65 |
| PEAR1 | -0.52 | -0.62 |
| FYCO1 | -0.50 | -0.62 |
| COBLL1 | -0.50 | -0.56 |
| DOCK6 | -0.50 | -0.59 |
| SHE | -0.49 | -0.58 |
| CGNL1 | -0.49 | -0.62 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| CASORELLI ACUTE PROMYELOCYTIC LEUKEMIA DN | 663 | 425 | All SZGR 2.0 genes in this pathway |
| VECCHI GASTRIC CANCER EARLY DN | 367 | 220 | All SZGR 2.0 genes in this pathway |
| OSMAN BLADDER CANCER DN | 406 | 230 | All SZGR 2.0 genes in this pathway |
| HOEBEKE LYMPHOID STEM CELL UP | 95 | 64 | All SZGR 2.0 genes in this pathway |
| JAATINEN HEMATOPOIETIC STEM CELL UP | 316 | 190 | All SZGR 2.0 genes in this pathway |
| GRAHAM CML QUIESCENT VS NORMAL QUIESCENT DN | 47 | 32 | All SZGR 2.0 genes in this pathway |
| GRAHAM CML DIVIDING VS NORMAL QUIESCENT DN | 95 | 57 | All SZGR 2.0 genes in this pathway |
| BROCKE APOPTOSIS REVERSED BY IL6 | 144 | 98 | All SZGR 2.0 genes in this pathway |
| FERRANDO T ALL WITH MLL ENL FUSION UP | 87 | 67 | All SZGR 2.0 genes in this pathway |
| KAYO AGING MUSCLE UP | 244 | 165 | All SZGR 2.0 genes in this pathway |
| GEORGANTAS HSC MARKERS | 71 | 47 | All SZGR 2.0 genes in this pathway |
| SHEDDEN LUNG CANCER GOOD SURVIVAL A12 | 317 | 177 | All SZGR 2.0 genes in this pathway |
| HOELZEL NF1 TARGETS UP | 139 | 93 | All SZGR 2.0 genes in this pathway |
| WIERENGA STAT5A TARGETS DN | 213 | 127 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |