Introduction of the 6mA-Finder :
DNA N6-methyladenine (6mA) represents an essential epigenetic modification in the genomes of diverse species, and plays an important role in the regulation of various biological processes, including restriction-modification system, DNA repair and replication, gene expression and the nucleoid segregation. Moreover, recent studies have shown that abnormal status of DNA 6mA modification was closely related to human cancers. The identification of new 6mA locus is fundamental for understanding the molecular mechanisms and regulatory roles of 6mA. In contrast with labor-intensive and expensive experiments, computational prediction of 6mA sites provided an alternatively rapid, accurate and cheap means.
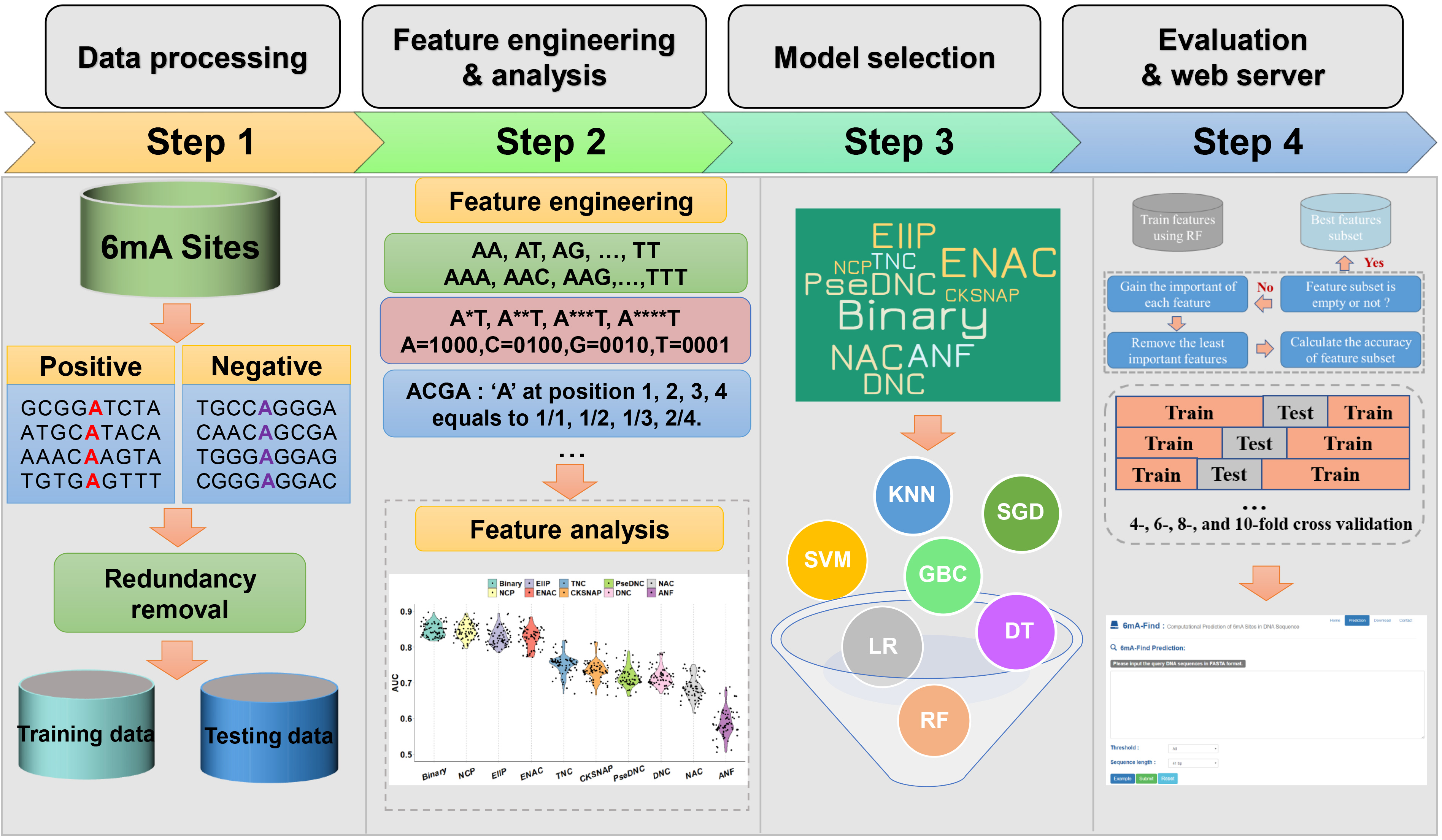
In this work, we carefully evaluated ten types of sequence and physicochemical features by seven conventional machine-learning algorithms such as Support Vector Machine (SVM), Random Forests (RF), Logistic Regression (LR), Stochastic Gradient Descent (SGD), Decision Trees (DT), K-Nearest Neighbors (KNN) and Gradient Boosting (GB), and observed all the features to be informative for predicting 6mA sites, while RF represents most powerful classifier. To avoid overfitting and improve the performance of the model, recursive feature elimination (RFE) strategy was used to select the optimal feature group. By exhaustively testing, here we integrated the ten types of features, and merged optimal features to develop a new tool of 6mA-Finder, with average area under curve (AUC) values of 0.9201, 0.9938 and 0.9367 based on multiple cross-validations for the general, mouse and rice prediction of 6mA sites. 6mA-Finder outperforms its peer tools for general and species-specific 6mA site prediction, suggesting it can provide a useful resource for further experimental investigation of DNA 6mA modification.