|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |
| Phenotypic Information (metabolism pathway, cancer, disease, phenome) |
| |
| |
| Gene-Gene Network Information: Co-Expression Network, Interacting Genes & KEGG |
| |
|
| Gene Summary for DPYD |
| Basic gene info. | Gene symbol | DPYD |
| Gene name | dihydropyrimidine dehydrogenase | |
| Synonyms | DHP|DHPDHASE|DPD | |
| Cytomap | UCSC genome browser: 1p22 | |
| Genomic location | chr1 :97543299-98386615 | |
| Type of gene | protein-coding | |
| RefGenes | NM_000110.3, NM_001160301.1, | |
| Ensembl id | ENSG00000188641 | |
| Description | dihydropyrimidine dehydrogenase [NADP(+)]dihydrothymine dehydrogenasedihydrouracil dehydrogenase | |
| Modification date | 20141207 | |
| dbXrefs | MIM : 612779 | |
| HGNC : HGNC | ||
| Ensembl : ENSG00000188641 | ||
| HPRD : 02036 | ||
| Vega : OTTHUMG00000039683 | ||
| Protein | UniProt: Q12882 go to UniProt's Cross Reference DB Table | |
| Expression | CleanEX: HS_DPYD | |
| BioGPS: 1806 | ||
| Gene Expression Atlas: ENSG00000188641 | ||
| The Human Protein Atlas: ENSG00000188641 | ||
| Pathway | NCI Pathway Interaction Database: DPYD | |
| KEGG: DPYD | ||
| REACTOME: DPYD | ||
| ConsensusPathDB | ||
| Pathway Commons: DPYD | ||
| Metabolism | MetaCyc: DPYD | |
| HUMANCyc: DPYD | ||
| Regulation | Ensembl's Regulation: ENSG00000188641 | |
| miRBase: chr1 :97,543,299-98,386,615 | ||
| TargetScan: NM_000110 | ||
| cisRED: ENSG00000188641 | ||
| Context | iHOP: DPYD | |
| cancer metabolism search in PubMed: DPYD | ||
| UCL Cancer Institute: DPYD | ||
| Assigned class in ccmGDB | A - This gene has a literature evidence and it belongs to cancer gene. | |
| References showing role of DPYD in cancer cell metabolism | 1. Metabolic rewiring is required for epithelial-mesenchymal transition. Cancer Discov 2014, 4: OF20. doi: 10.1158/2159-8290.CD-RW2014-190. go to article 2. Offer SM, Wegner NJ, Fossum C, Wang K, Diasio RB (2013) Phenotypic profiling of DPYD variations relevant to 5-fluorouracil sensitivity using real-time cellular analysis and in vitro measurement of enzyme activity. Cancer Res 73: 1958-1968. doi: 10.1158/0008-5472.CAN-12-3858. pmid: 3602211. go to article 3. Offer SM, Butterfield GL, Jerde CR, Fossum CC, Wegner NJ, et al. (2014) microRNAs miR-27a and miR-27b directly regulate liver dihydropyrimidine dehydrogenase expression through two conserved binding sites. Mol Cancer Ther 13: 742-751. doi: 10.1158/1535-7163.MCT-13-0878. pmid: 3954441. go to article 4. Shaul YD, Freinkman E, Comb WC, Cantor JR, Tam WL, et al. (2014) Dihydropyrimidine accumulation is required for the epithelial-mesenchymal transition. Cell 158: 1094-1109. doi: 10.1016/j.cell.2014.07.032. pmid: 4250222. go to article | |
| Top |
| Phenotypic Information for DPYD(metabolism pathway, cancer, disease, phenome) |
| Cancer | CGAP: DPYD |
| Familial Cancer Database: DPYD | |
| * This gene is included in those cancer gene databases. |
|
|
|
|
|
| . | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Oncogene 1 | Significant driver gene in | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| cf) number; DB name 1 Oncogene; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 2 Tumor Suppressor gene; https://bioinfo.uth.edu/TSGene/, 3 Cancer Gene Census; http://www.nature.com/nrc/journal/v4/n3/abs/nrc1299.html, 4 CancerGenes; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 5 Network of Cancer Gene; http://ncg.kcl.ac.uk/index.php, 1Therapeutic Vulnerabilities in Cancer; http://cbio.mskcc.org/cancergenomics/statius/ |
| KEGG_PYRIMIDINE_METABOLISM KEGG_BETA_ALANINE_METABOLISM KEGG_DRUG_METABOLISM_OTHER_ENZYMES REACTOME_METABOLISM_OF_NUCLEOTIDES REACTOME_PYRIMIDINE_METABOLISM | |
| OMIM | 274270; phenotype. 274270; phenotype. 612779; gene. 612779; gene. |
| Orphanet | 1675; Dihydropyrimidine dehydrogenase deficiency. 1675; Dihydropyrimidine dehydrogenase deficiency. 240839; 5-fluorouracil toxicity. 240839; 5-fluorouracil toxicity. 240855; Capecitabine toxicity. 240855; Capecitabine toxicity. 240955; Susceptibility to adverse reaction due to 5-fluorouracil treatment. 240955; Susceptibility to adverse reaction due to 5-fluorouracil treatment. 240963; Susceptibility to adverse reaction due to capecitabine treatment. 240963; Susceptibility to adverse reaction due to capecitabine treatment. 293948; 1p21.3 microdeletion syndrome. 293948; 1p21.3 microdeletion syndrome. |
| Disease | KEGG Disease: DPYD |
| MedGen: DPYD (Human Medical Genetics with Condition) | |
| ClinVar: DPYD | |
| Phenotype | MGI: DPYD (International Mouse Phenotyping Consortium) |
| PhenomicDB: DPYD | |
| Mutations for DPYD |
| * Under tables are showing count per each tissue to give us broad intuition about tissue specific mutation patterns.You can go to the detailed page for each mutation database's web site. |
| - Statistics for Tissue and Mutation type | Top |
 |
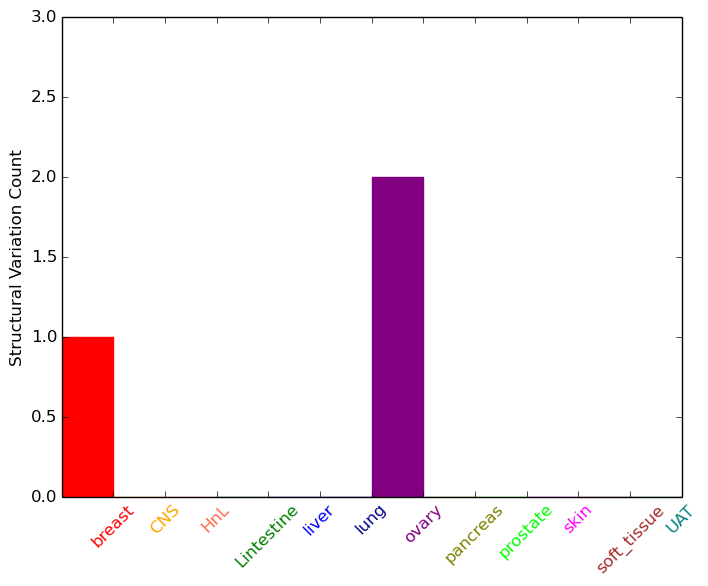
| - For Inter-chromosomal Variations |
| * Inter-chromosomal variantions includes 'interchromosomal amplicon to amplicon', 'interchromosomal amplicon to non-amplified dna', 'interchromosomal insertion', 'Interchromosomal unknown type'. |
 |
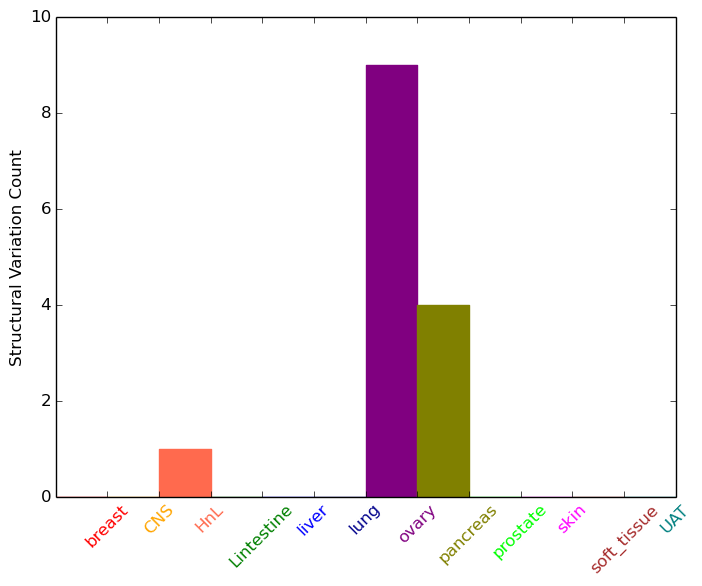
| - For Intra-chromosomal Variations |
| * Intra-chromosomal variantions includes 'intrachromosomal amplicon to amplicon', 'intrachromosomal amplicon to non-amplified dna', 'intrachromosomal deletion', 'intrachromosomal fold-back inversion', 'intrachromosomal inversion', 'intrachromosomal tandem duplication', 'Intrachromosomal unknown type', 'intrachromosomal with inverted orientation', 'intrachromosomal with non-inverted orientation'. |
 |
| Sample | Symbol_a | Chr_a | Start_a | End_a | Symbol_b | Chr_b | Start_b | End_b |
| breast | DPYD | chr1 | 98012751 | 98012751 | IQSEC2 | chr23 | 53278229 | 53278229 |
| haematopoietic_and_lymphoid_tissue | DPYD | chr1 | 97613255 | 97613255 | DPYD | chr1 | 97596846 | 97596846 |
| ovary | DPYD | chr1 | 97634042 | 97634062 | AFF3 | chr2 | 100567111 | 100567131 |
| ovary | DPYD | chr1 | 97634475 | 97634495 | DPYD | chr1 | 97634579 | 97634599 |
| ovary | DPYD | chr1 | 97686510 | 97686530 | PTBP2 | chr1 | 97242249 | 97242269 |
| ovary | DPYD | chr1 | 97717572 | 97717592 | DPYD | chr1 | 98258856 | 98258876 |
| ovary | DPYD | chr1 | 97758588 | 97758608 | DPYD | chr1 | 97754020 | 97754040 |
| ovary | DPYD | chr1 | 97758591 | 97758611 | DPYD | chr1 | 97754023 | 97754043 |
| ovary | DPYD | chr1 | 97758591 | 97758611 | INTS9 | chr8 | 28677032 | 28677052 |
| ovary | DPYD | chr1 | 97887428 | 97887448 | DPYD | chr1 | 97885061 | 97885081 |
| ovary | DPYD | chr1 | 97956955 | 97956975 | DPYD | chr1 | 97957015 | 97957035 |
| ovary | DPYD | chr1 | 97988432 | 97988452 | DPYD | chr1 | 98248676 | 98248696 |
| pancreas | DPYD | chr1 | 97662539 | 97662559 | DPYD | chr1 | 97687212 | 97687232 |
| pancreas | DPYD | chr1 | 97662539 | 97662559 | DPYD | chr1 | 97687217 | 97687237 |
| pancreas | DPYD | chr1 | 97726352 | 97726372 | DPYD | chr1 | 97740442 | 97740462 |
| pancreas | DPYD | chr1 | 97838203 | 97838223 | chr1 | 66178314 | 66178334 |
| cf) Tissue number; Tissue name (1;Breast, 2;Central_nervous_system, 3;Haematopoietic_and_lymphoid_tissue, 4;Large_intestine, 5;Liver, 6;Lung, 7;Ovary, 8;Pancreas, 9;Prostate, 10;Skin, 11;Soft_tissue, 12;Upper_aerodigestive_tract) |
| * From mRNA Sanger sequences, Chitars2.0 arranged chimeric transcripts. This table shows DPYD related fusion information. |
| ID | Head Gene | Tail Gene | Accession | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a |
| BG988212 | DPYD | 20 | 303 | 1 | 98128967 | 98129251 | LOC100507373 | 301 | 622 | 19 | 14280890 | 14281216 | |
| BP319012 | DPYD | 1 | 356 | 1 | 98320818 | 98386590 | COX5B | 345 | 437 | 2 | 98264565 | 98264657 | |
| CN412218 | DPYD | 481 | 502 | 1 | 97929526 | 97929547 | KXD1 | 496 | 601 | 19 | 18670041 | 18670146 | |
| CV375938 | SRRM2 | 1 | 92 | 16 | 2804703 | 2804796 | DPYD | 83 | 516 | 1 | 98322148 | 98322581 | |
| BI603836 | DPYD | 4 | 550 | 1 | 98048815 | 98049359 | NRIP3 | 549 | 750 | 11 | 9002563 | 9002769 | |
| AB042557 | NBPF9 | 1 | 1132 | 1 | 144951765 | 144994909 | DPYD | 1130 | 1155 | 1 | 97956375 | 97956402 | |
| Top |
| Mutation type/ Tissue ID | brca | cns | cerv | endome | haematopo | kidn | Lintest | liver | lung | ns | ovary | pancre | prost | skin | stoma | thyro | urina | |||
| Total # sample | 1 | 1 | 3 | 1 | 1 | 2 | ||||||||||||||
| GAIN (# sample) | 1 | 2 | 1 | 2 | ||||||||||||||||
| LOSS (# sample) | 1 | 2 | 1 |
| cf) Tissue ID; Tissue type (1; Breast, 2; Central_nervous_system, 3; Cervix, 4; Endometrium, 5; Haematopoietic_and_lymphoid_tissue, 6; Kidney, 7; Large_intestine, 8; Liver, 9; Lung, 10; NS, 11; Ovary, 12; Pancreas, 13; Prostate, 14; Skin, 15; Stomach, 16; Thyroid, 17; Urinary_tract) |
| Top |
|
 |
| Top |
| Stat. for Non-Synonymous SNVs (# total SNVs=154) | (# total SNVs=34) |
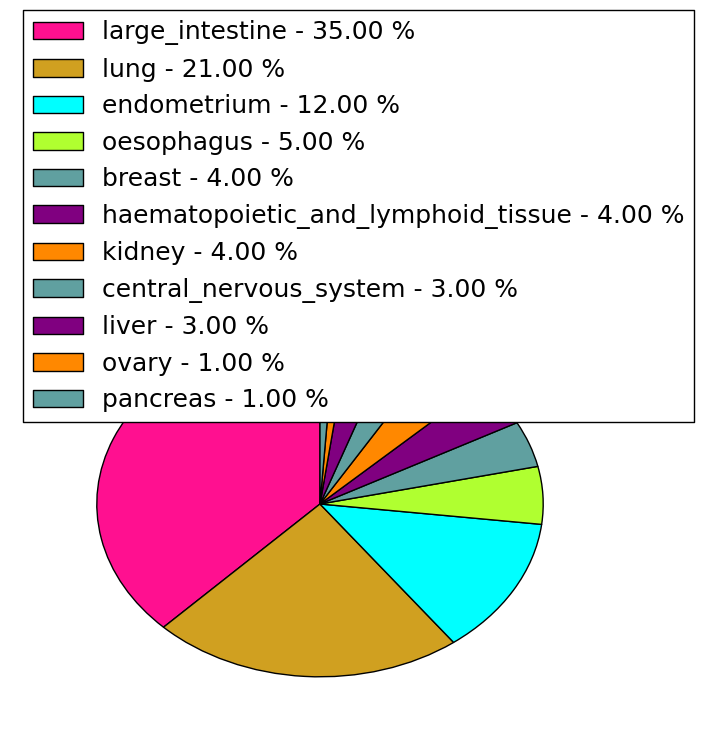 | 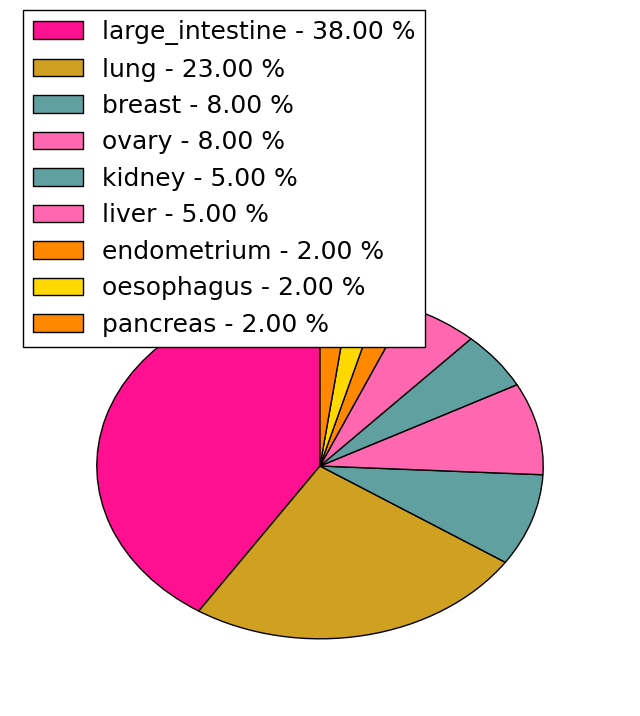 |
(# total SNVs=0) | (# total SNVs=0) |
| Top |
| * When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site,primary_histology,mutation(aa),pubmedID. |
| GRCh37 position | Mutation(aa) | Unique sampleID count |
| chr1:98164976-98164976 | p.S204F | 4 |
| chr1:97700485-97700485 | p.P789S | 4 |
| chr1:98165013-98165013 | p.L192F | 3 |
| chr1:98039426-98039426 | p.R410Q | 3 |
| chr1:97544582-97544582 | p.P1010T | 3 |
| chr1:97981407-97981407 | p.G539R | 3 |
| chr1:97981340-97981340 | p.R561Q | 3 |
| chr1:97847978-97847978 | p.D649Y | 3 |
| chr1:98205983-98205983 | p.D96N | 3 |
| chr1:97839164-97839164 | p.C671G | 3 |
| Top |
|
 |
| Point Mutation/ Tissue ID | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 |
| # sample | 5 | 1 | 39 | 2 | 10 | 3 | 15 | 7 | 3 | 2 | 53 | 17 | 15 | |||||||
| # mutation | 6 | 1 | 44 | 2 | 12 | 3 | 21 | 11 | 3 | 2 | 61 | 20 | 20 | |||||||
| nonsynonymous SNV | 5 | 35 | 2 | 8 | 2 | 17 | 8 | 1 | 2 | 47 | 12 | 19 | ||||||||
| synonymous SNV | 1 | 1 | 9 | 4 | 1 | 4 | 3 | 2 | 14 | 8 | 1 |
| cf) Tissue ID; Tissue type (1; BLCA[Bladder Urothelial Carcinoma], 2; BRCA[Breast invasive carcinoma], 3; CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], 4; COAD[Colon adenocarcinoma], 5; GBM[Glioblastoma multiforme], 6; Glioma Low Grade, 7; HNSC[Head and Neck squamous cell carcinoma], 8; KICH[Kidney Chromophobe], 9; KIRC[Kidney renal clear cell carcinoma], 10; KIRP[Kidney renal papillary cell carcinoma], 11; LAML[Acute Myeloid Leukemia], 12; LUAD[Lung adenocarcinoma], 13; LUSC[Lung squamous cell carcinoma], 14; OV[Ovarian serous cystadenocarcinoma ], 15; PAAD[Pancreatic adenocarcinoma], 16; PRAD[Prostate adenocarcinoma], 17; SKCM[Skin Cutaneous Melanoma], 18:STAD[Stomach adenocarcinoma], 19:THCA[Thyroid carcinoma], 20:UCEC[Uterine Corpus Endometrial Carcinoma]) |
| Top |
| * We represented just top 10 SNVs. When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site, primary_histology, mutation(aa), pubmedID. |
| Genomic Position | Mutation(aa) | Unique sampleID count |
| chr1:98157281 | p.I889I | 2 |
| chr1:97658647 | p.V362I | 2 |
| chr1:98039506 | p.P789S | 2 |
| chr1:97839162 | p.S204F | 2 |
| chr1:97700485 | p.I361I | 2 |
| chr1:98348842 | p.N955N | 2 |
| chr1:97981388 | p.A437V | 2 |
| chr1:98205983 | p.R867L | 2 |
| chr1:98058899 | p.D949N | 2 |
| chr1:98039345 | p.E415K | 2 |
| * Copy number data were extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered on Jan-05-2015. Function ProcessCNAData in TCGA-Assembler package was used to obtain gene-level copy number value which is calculated as the average copy number of the genomic region of a gene. |
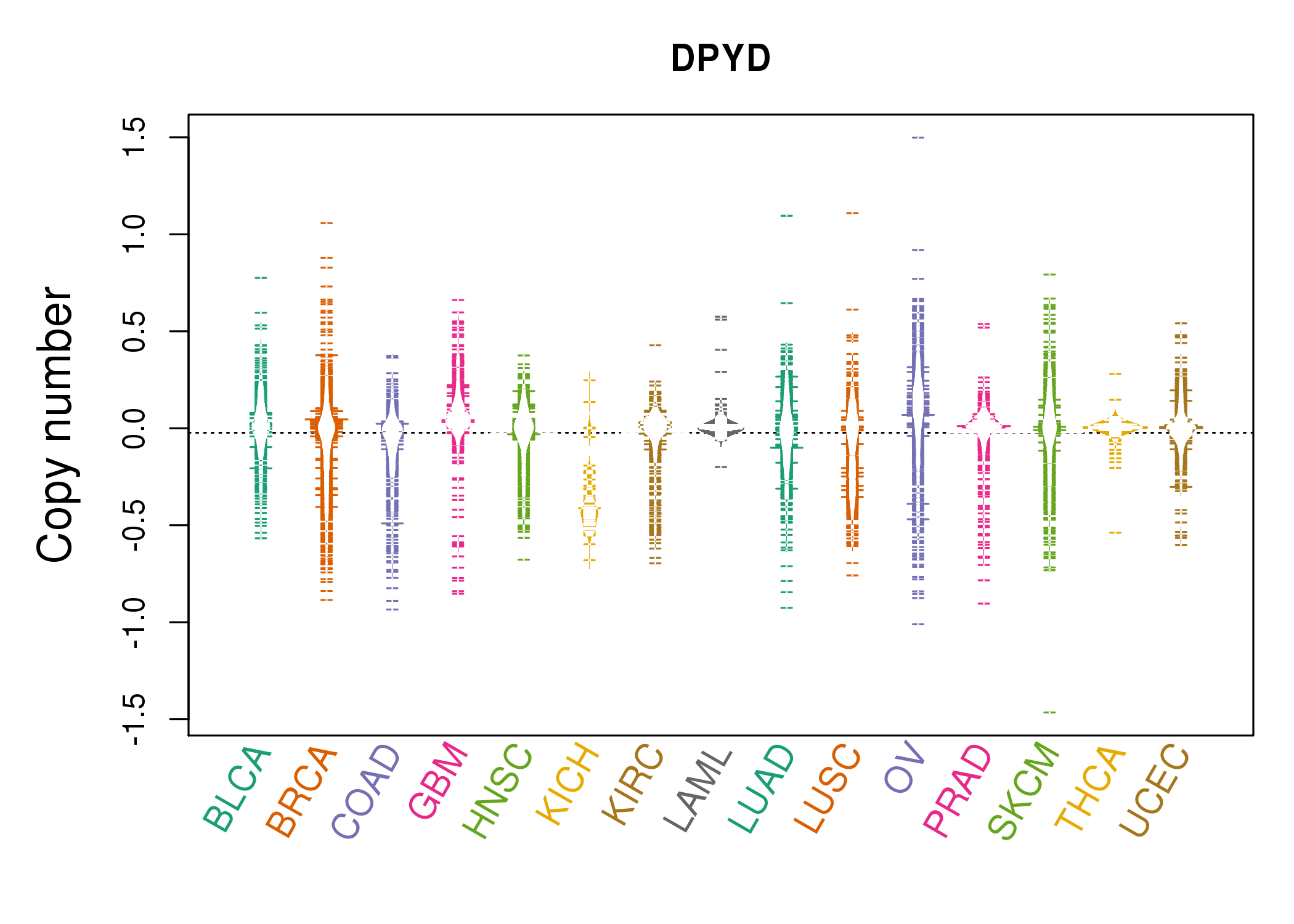 |
| cf) Tissue ID[Tissue type]: BLCA[Bladder Urothelial Carcinoma], BRCA[Breast invasive carcinoma], CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], COAD[Colon adenocarcinoma], GBM[Glioblastoma multiforme], Glioma Low Grade, HNSC[Head and Neck squamous cell carcinoma], KICH[Kidney Chromophobe], KIRC[Kidney renal clear cell carcinoma], KIRP[Kidney renal papillary cell carcinoma], LAML[Acute Myeloid Leukemia], LUAD[Lung adenocarcinoma], LUSC[Lung squamous cell carcinoma], OV[Ovarian serous cystadenocarcinoma ], PAAD[Pancreatic adenocarcinoma], PRAD[Prostate adenocarcinoma], SKCM[Skin Cutaneous Melanoma], STAD[Stomach adenocarcinoma], THCA[Thyroid carcinoma], UCEC[Uterine Corpus Endometrial Carcinoma] |
| Top |
| Gene Expression for DPYD |
| * CCLE gene expression data were extracted from CCLE_Expression_Entrez_2012-10-18.res: Gene-centric RMA-normalized mRNA expression data. |
 |
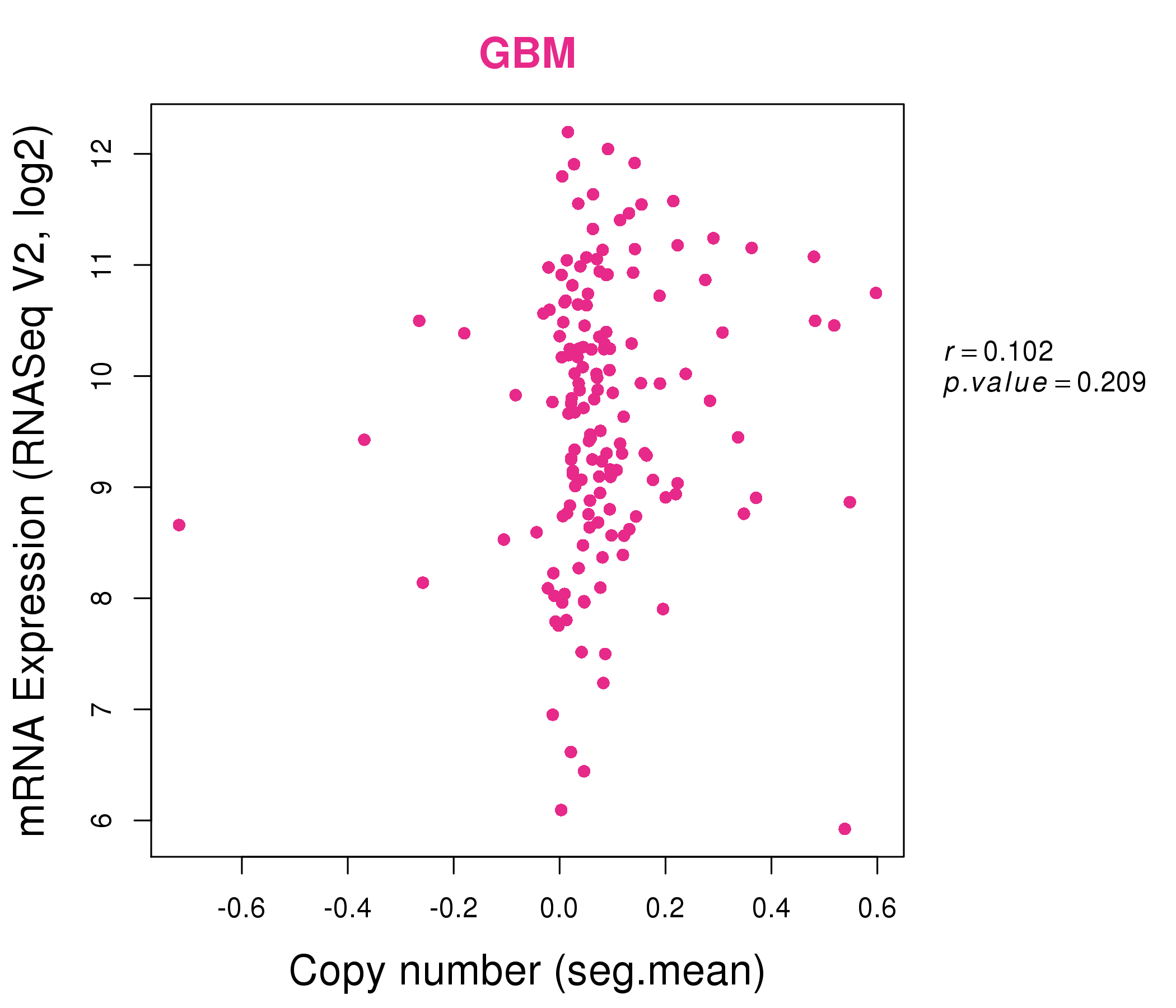
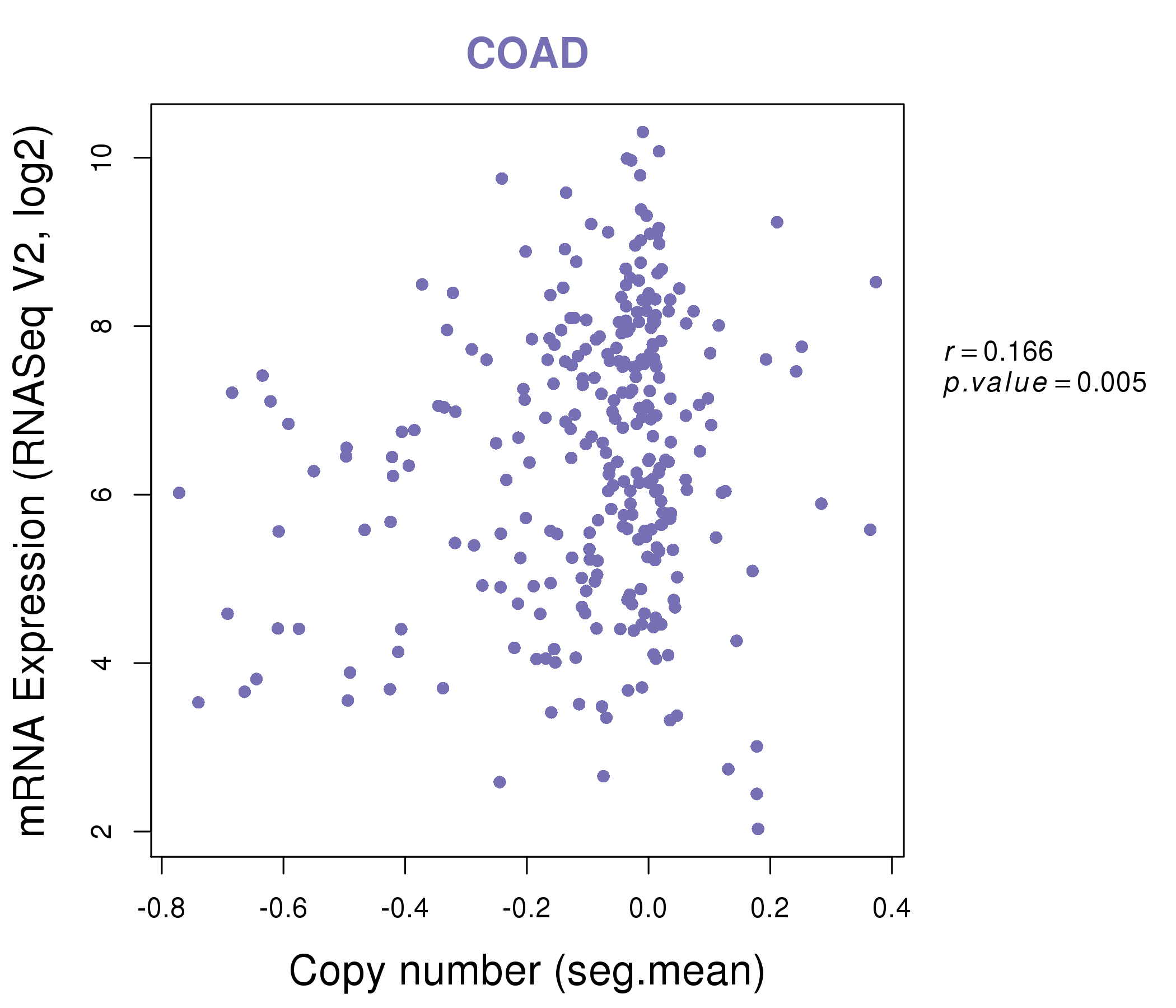
| * Normalized gene expression data of RNASeqV2 was extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered at Jan-05-2015. Only eight cancer types have enough normal control samples for differential expression analysis. (t test, adjusted p<0.05 (using Benjamini-Hochberg FDR)) |
 |
| Top |
| * This plots show the correlation between CNV and gene expression. |
: Open all plots for all cancer types
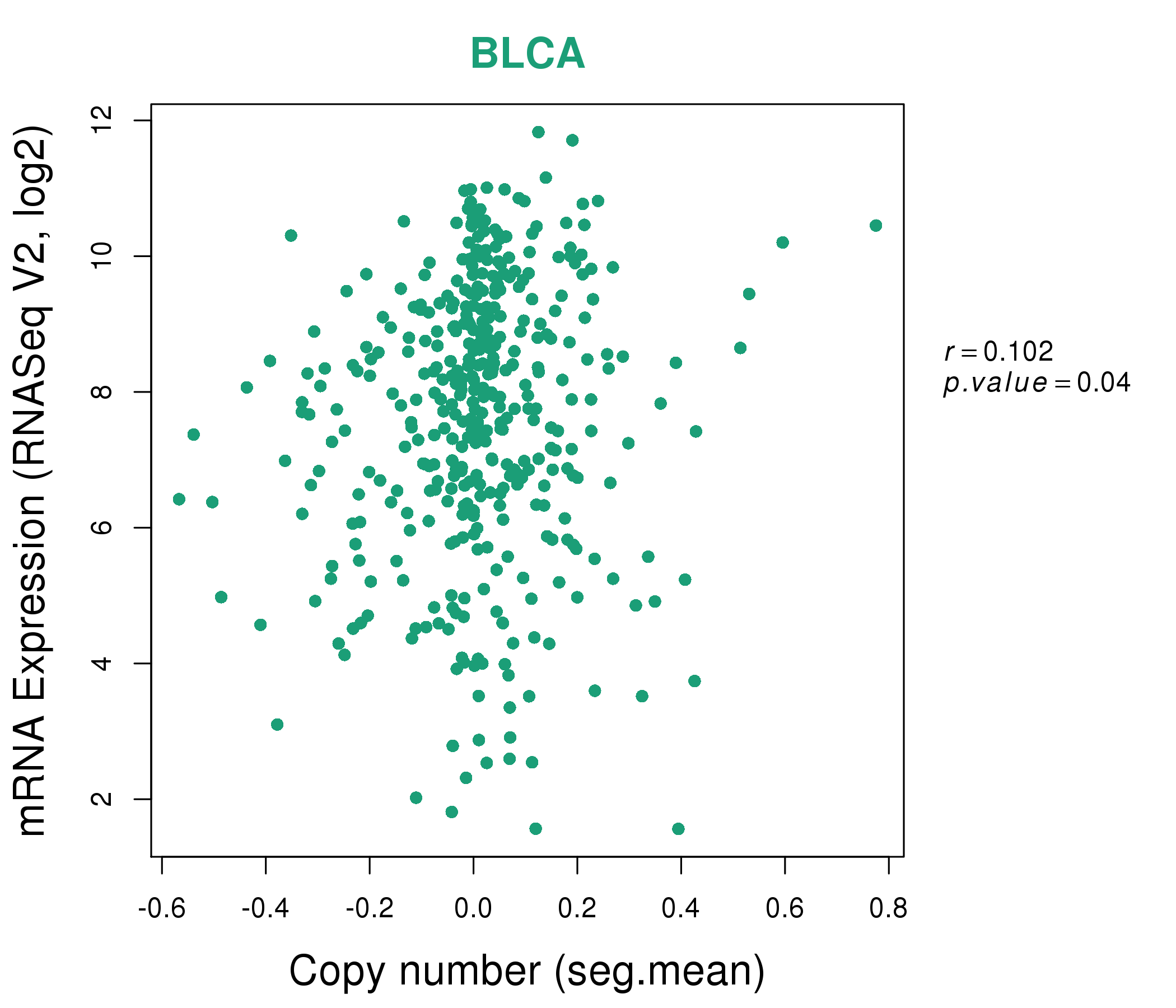 |
|
 |
|
| Top |
| Gene-Gene Network Information |
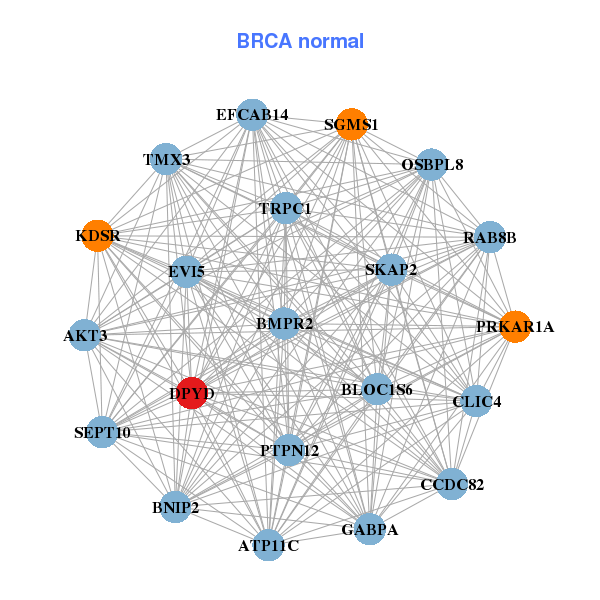
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
 |
| ||||
| ARAP2,C10orf128,CYBB,DPYD,FGL2,HLA-DRA,KCTD12, LHFPL2,MCTP1,MPEG1,NCKAP1L,PIK3CG,PTAFR,RAB8B, RGS18,SLC8A1,SLCO2B1,TFEC,TLR4,WIPF1,ZEB2 | AKT3,ATP11C,BMPR2,BNIP2,CCDC82,CLIC4,DPYD, EVI5,GABPA,KDSR,EFCAB14,OSBPL8,BLOC1S6,PRKAR1A, PTPN12,RAB8B,SEPT10,SGMS1,SKAP2,TMX3,TRPC1 | ||||
 |
| ||||
| C3AR1,CMKLR1,CSF1,DPYD,HAVCR2,IL2RA,LAIR1, LAPTM5,LCP2,LILRB1,MAFB,MS4A4A,MS4A6A,MYO5A, NRP1,PDCD1LG2,PLEKHO2,SLAMF8,SLC15A3,STX11,TNFSF13B | PRR34,CA11,CDKN2C,CTSO,DISP1,DOCK11,DPYD, FAM189B,LMO4,MCFD2,NAB2,NINJ1,P2RX7,PDE1B, CPQ,SAP30,SCPEP1,SEC22C,SHF,SNX18,THOC5 |
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
| Top |
: Open all interacting genes' information including KEGG pathway for all interacting genes from DAVID
| Top |
| Pharmacological Information for DPYD |
| DB Category | DB Name | DB's ID and Url link |
| Chemistry | BindingDB | Q12882; -. |
| Chemistry | ChEMBL | CHEMBL3172; -. |
| Chemistry | BindingDB | Q12882; -. |
| Chemistry | ChEMBL | CHEMBL3172; -. |
| Organism-specific databases | PharmGKB | PA145; -. |
| Organism-specific databases | PharmGKB | PA145; -. |
| Organism-specific databases | CTD | 1806; -. |
| Organism-specific databases | CTD | 1806; -. |
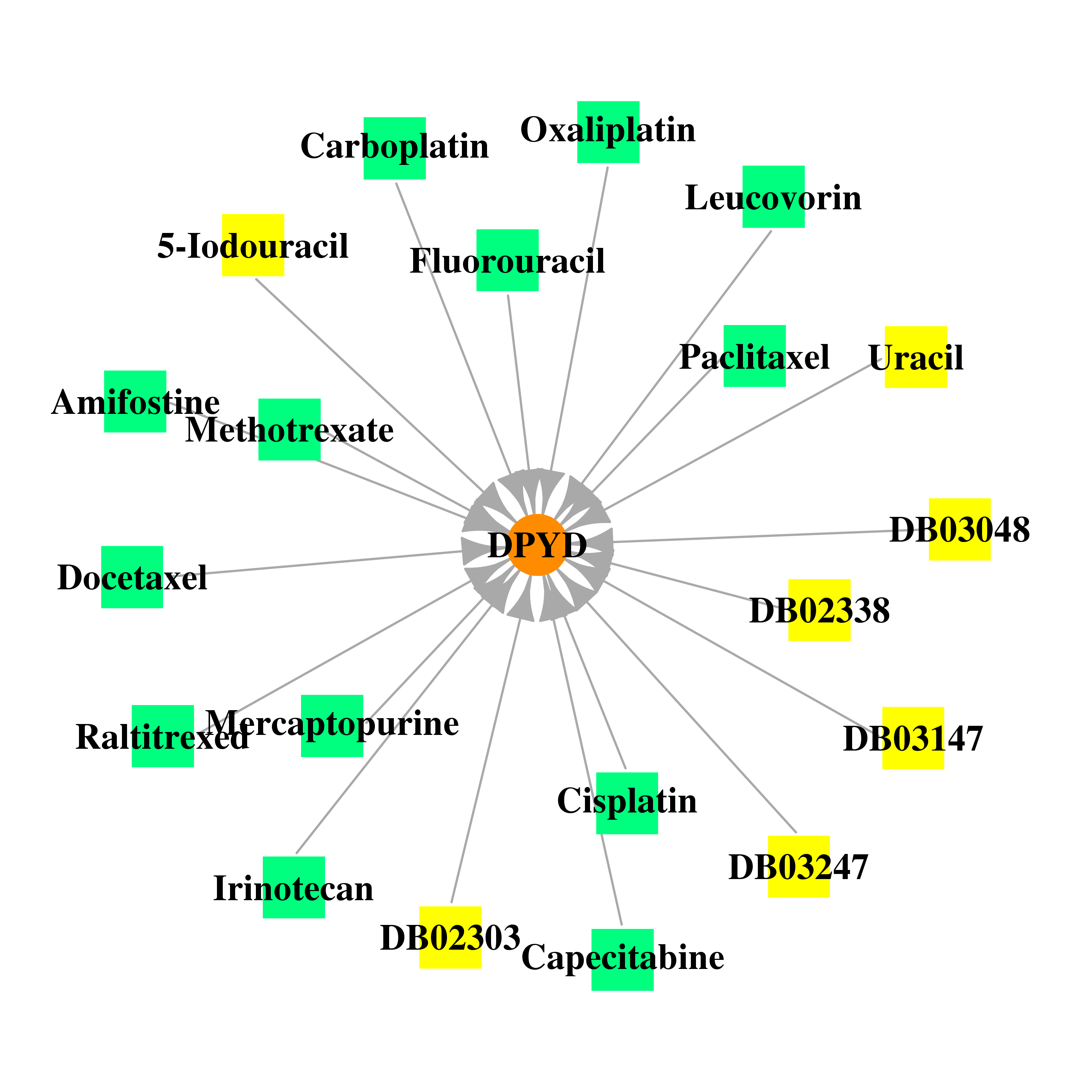
| * Gene Centered Interaction Network. |
 |
| * Drug Centered Interaction Network. |
| DrugBank ID | Target Name | Drug Groups | Generic Name | Drug Centered Network | Drug Structure |
| DB02303 | dihydropyrimidine dehydrogenase | experimental | (5s)-5-Iododihydro-2,4(1h,3h)-Pyrimidinedione | 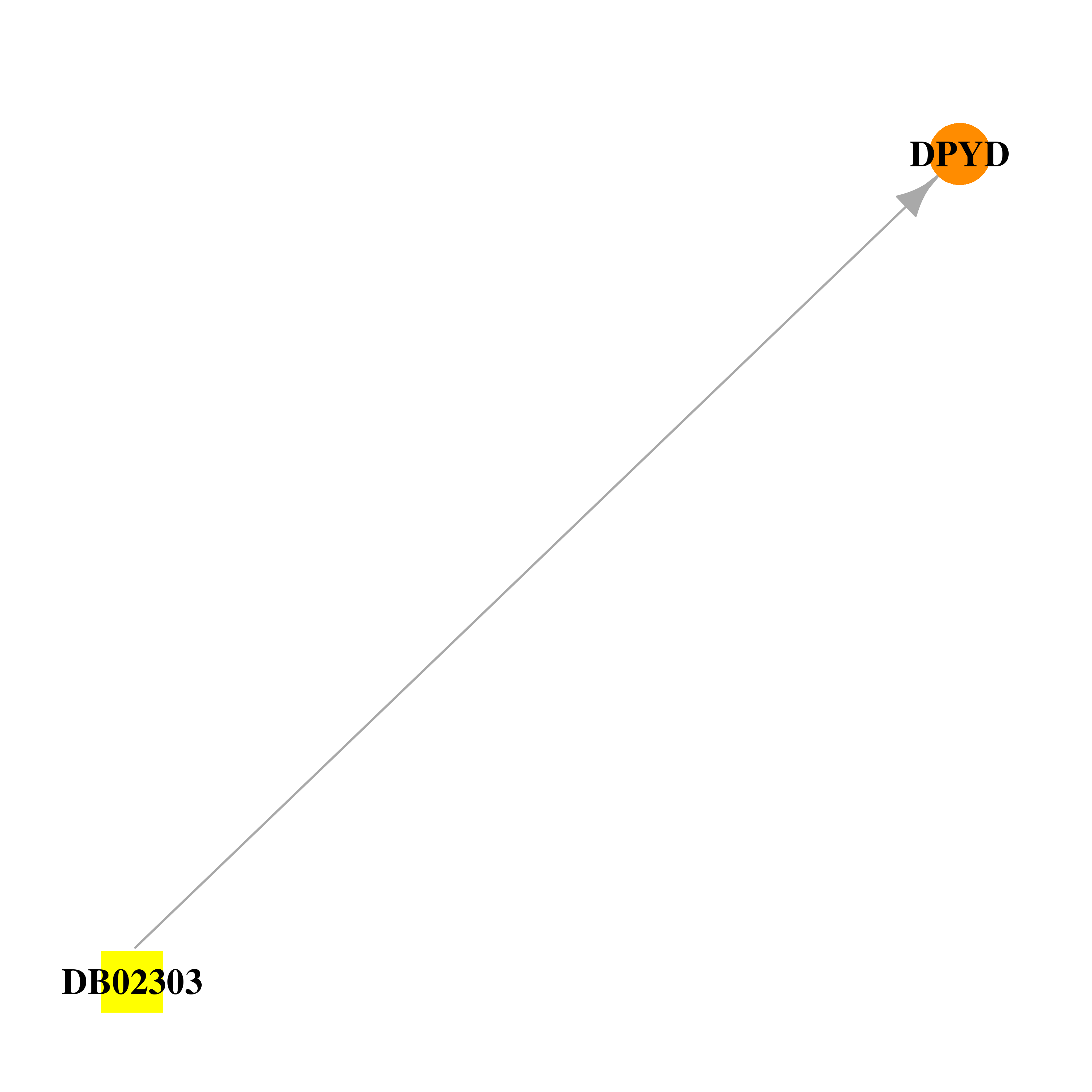 | 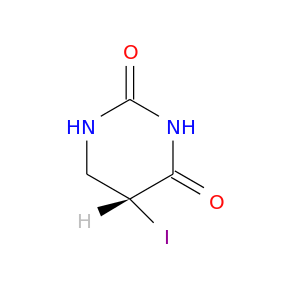 |
| DB02338 | dihydropyrimidine dehydrogenase | experimental | Nadph Dihydro-Nicotinamide-Adenine-Dinucleotidephosphate | 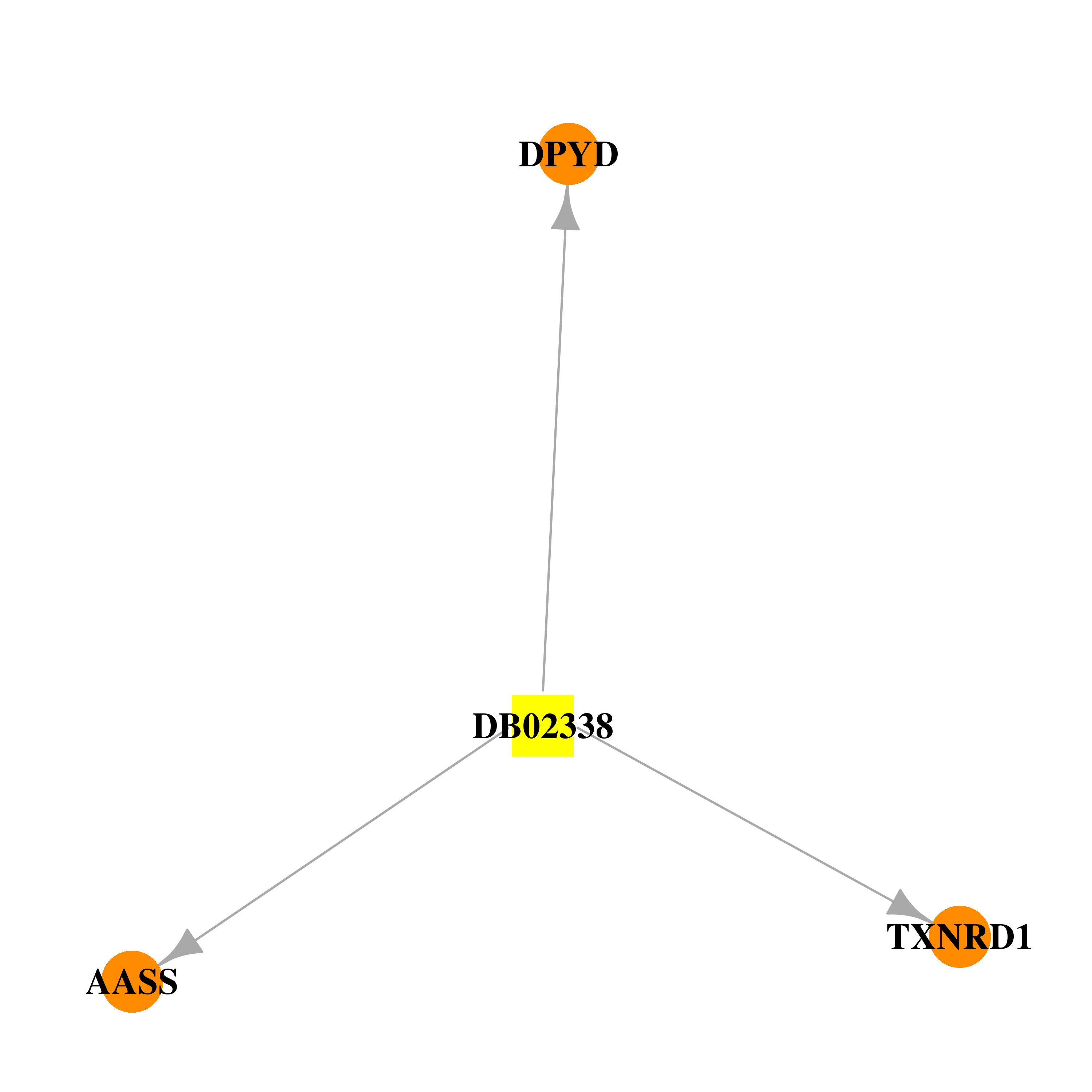 | 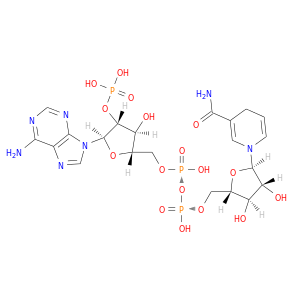 |
| DB03048 | dihydropyrimidine dehydrogenase | experimental | 6-Carboxymethyluracil | 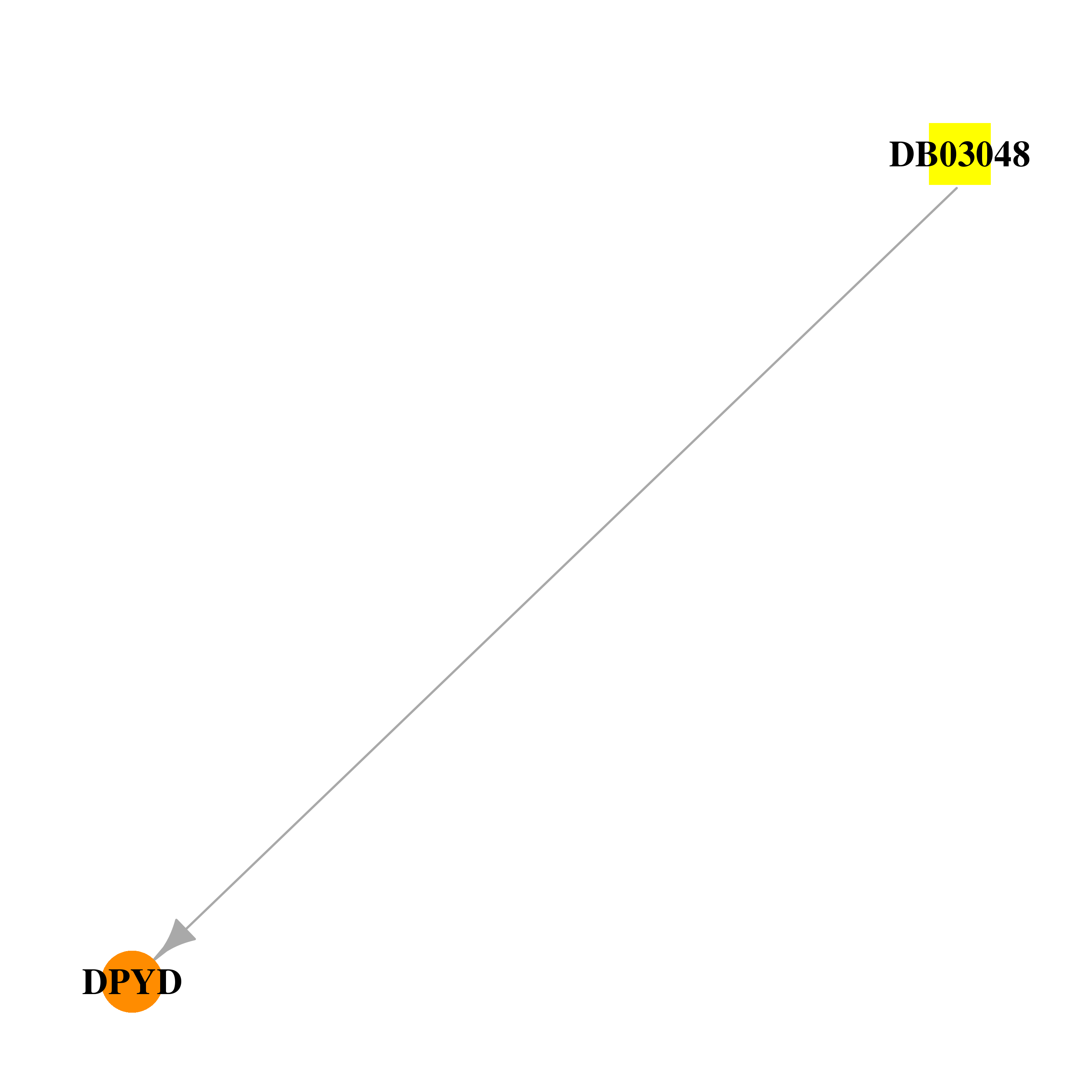 | 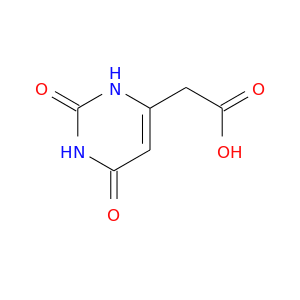 |
| DB03147 | dihydropyrimidine dehydrogenase | experimental | Flavin-Adenine Dinucleotide | 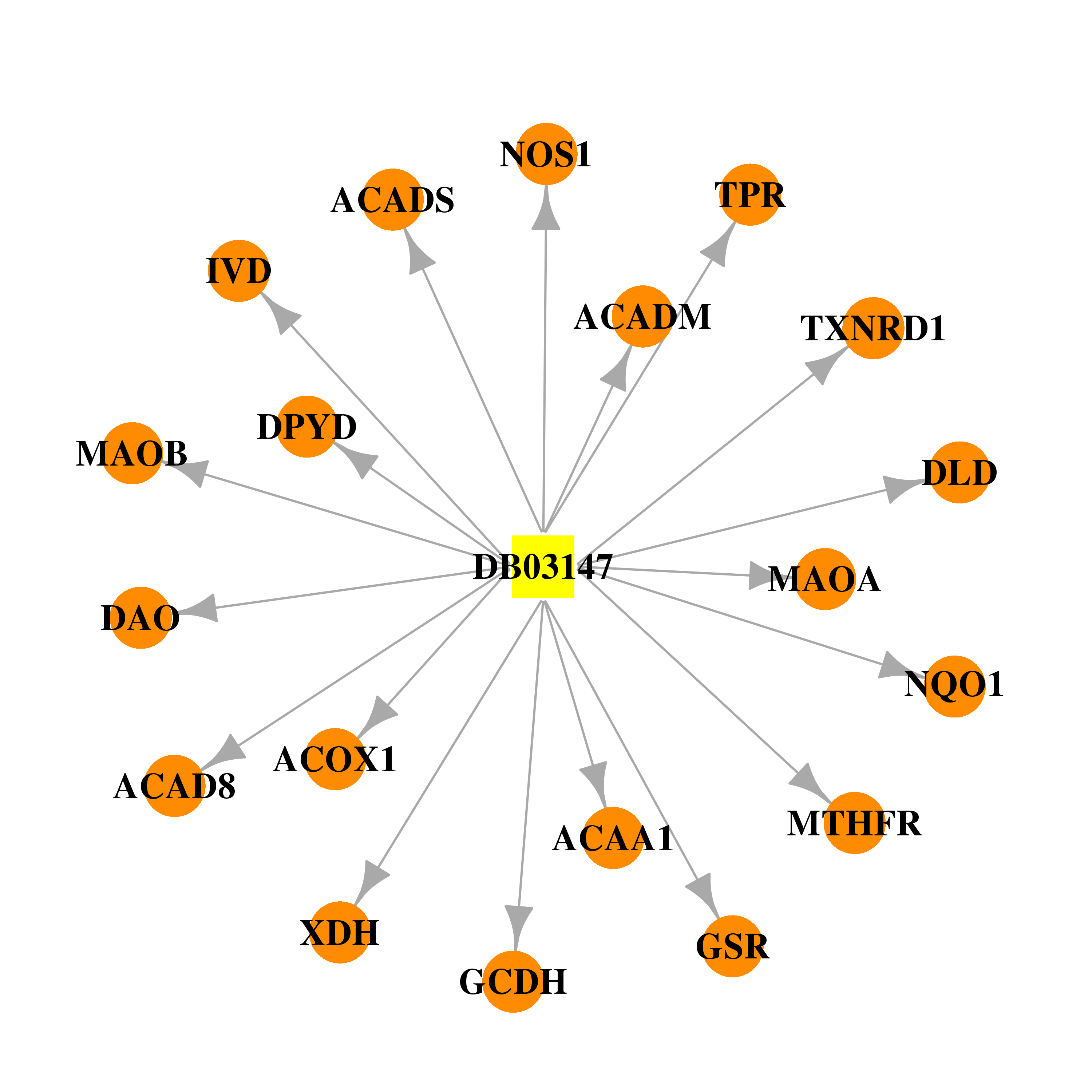 |  |
| DB03247 | dihydropyrimidine dehydrogenase | experimental | Riboflavin Monophosphate |  | 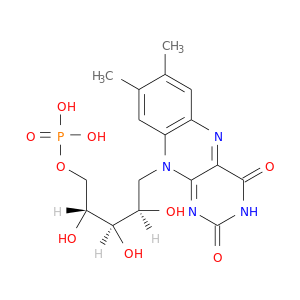 |
| DB03419 | dihydropyrimidine dehydrogenase | experimental | Uracil | 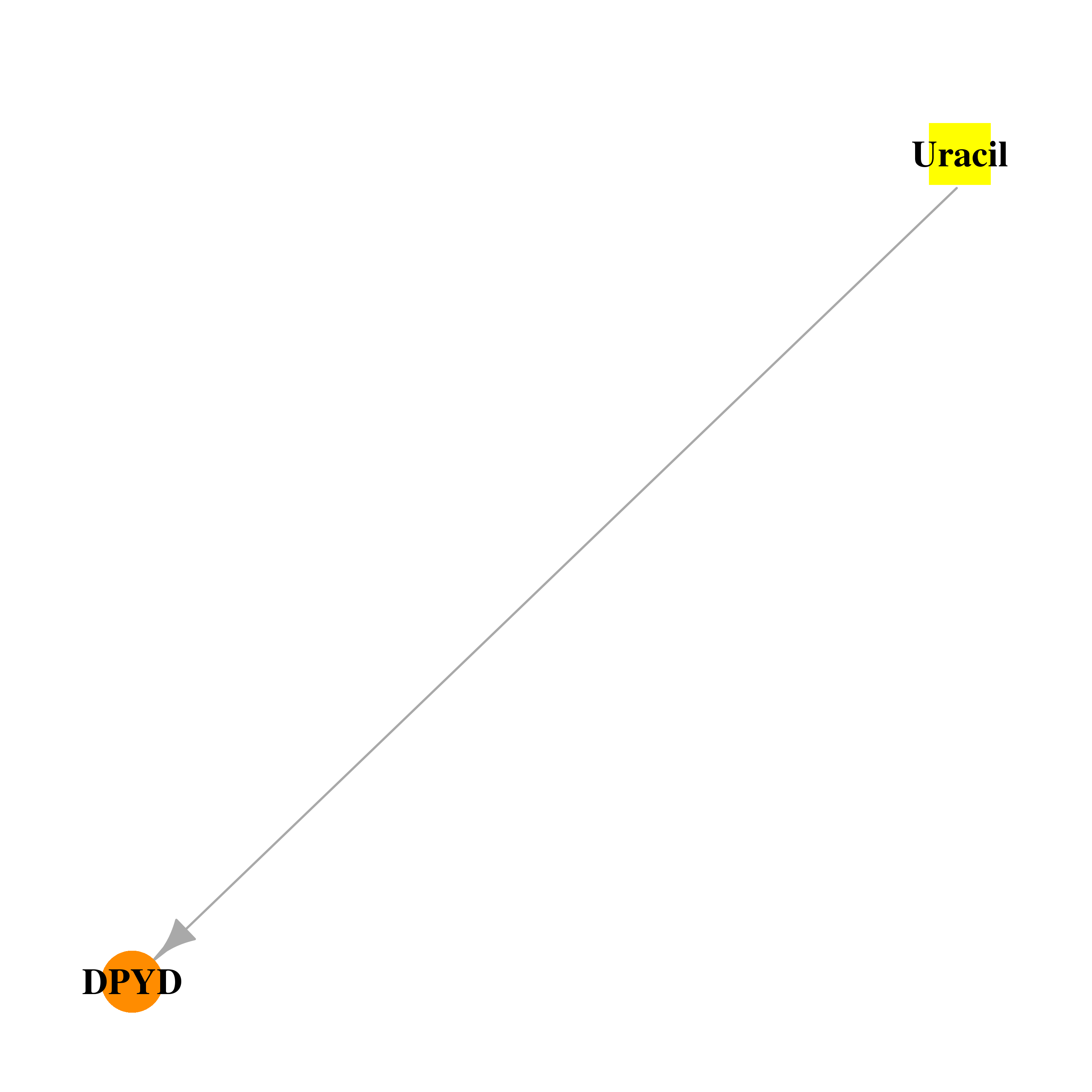 | 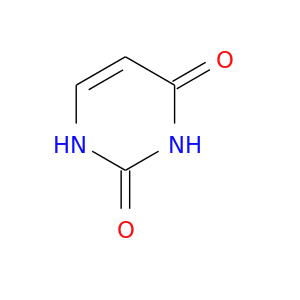 |
| DB03554 | dihydropyrimidine dehydrogenase | experimental | 5-Iodouracil | 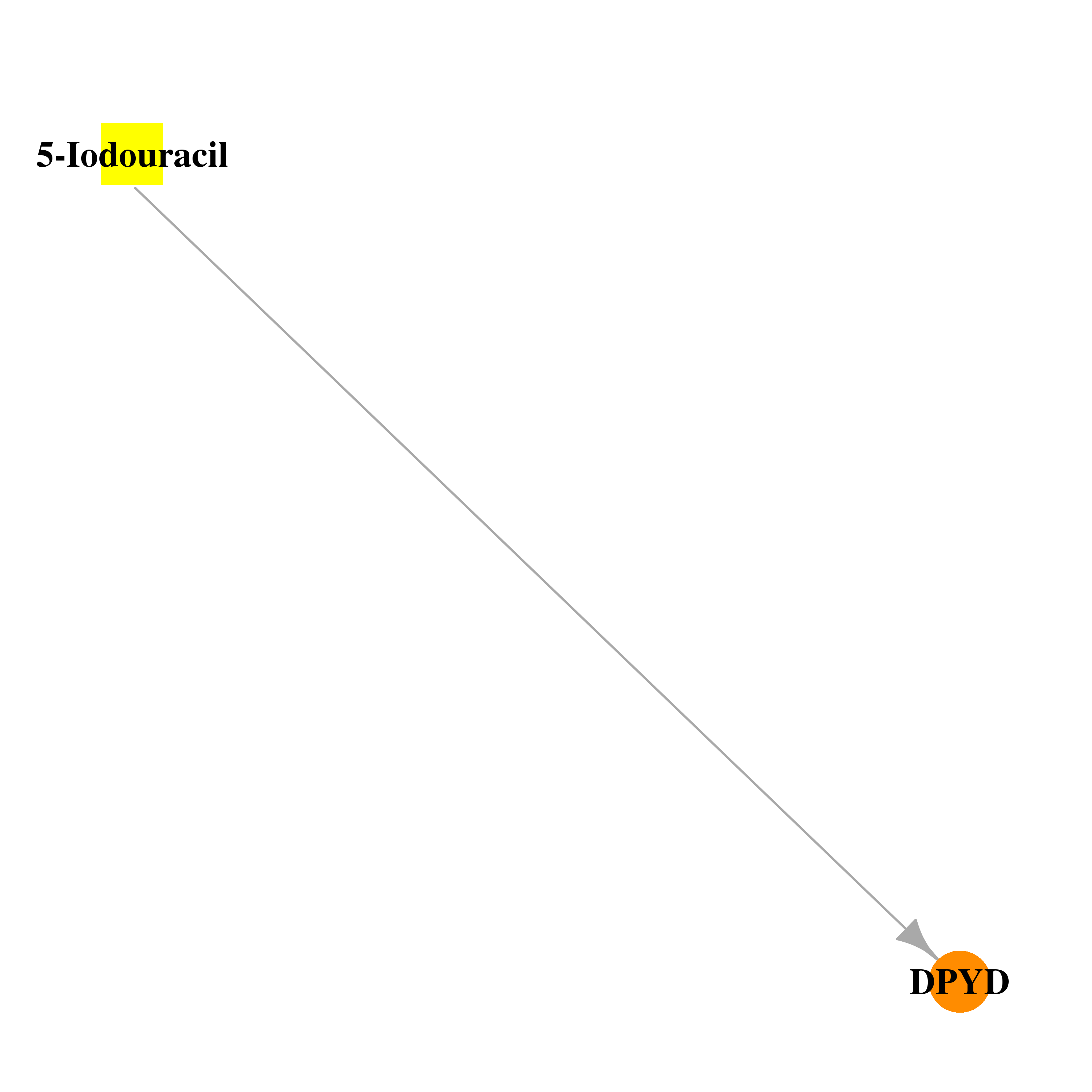 | 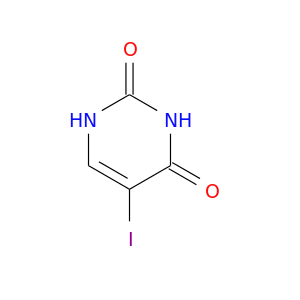 |
| DB01248 | dihydropyrimidine dehydrogenase | approved; investigational | Docetaxel | 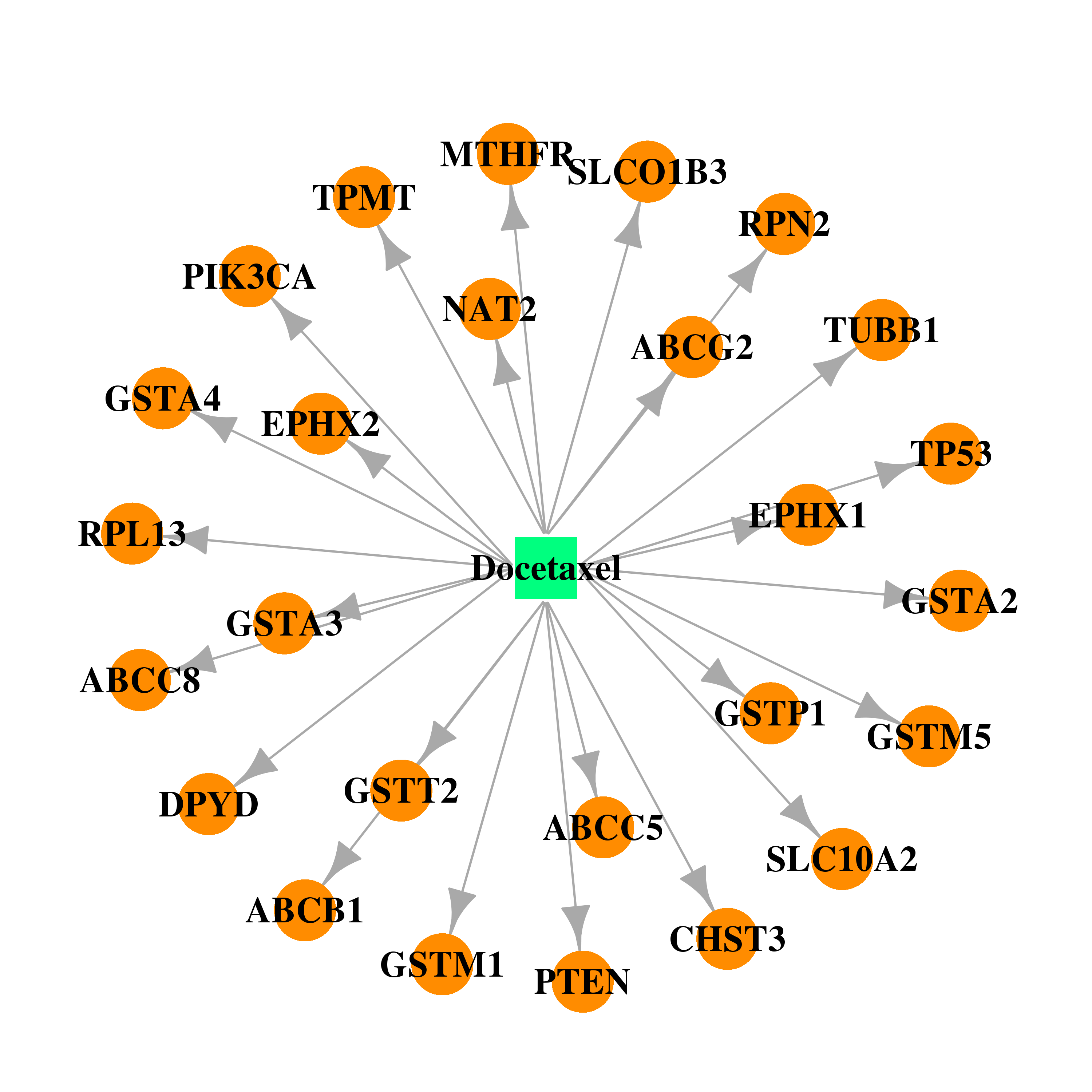 | 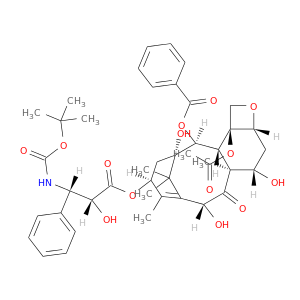 |
| DB00515 | dihydropyrimidine dehydrogenase | approved | Cisplatin |  |  |
| DB00650 | dihydropyrimidine dehydrogenase | approved | Leucovorin | 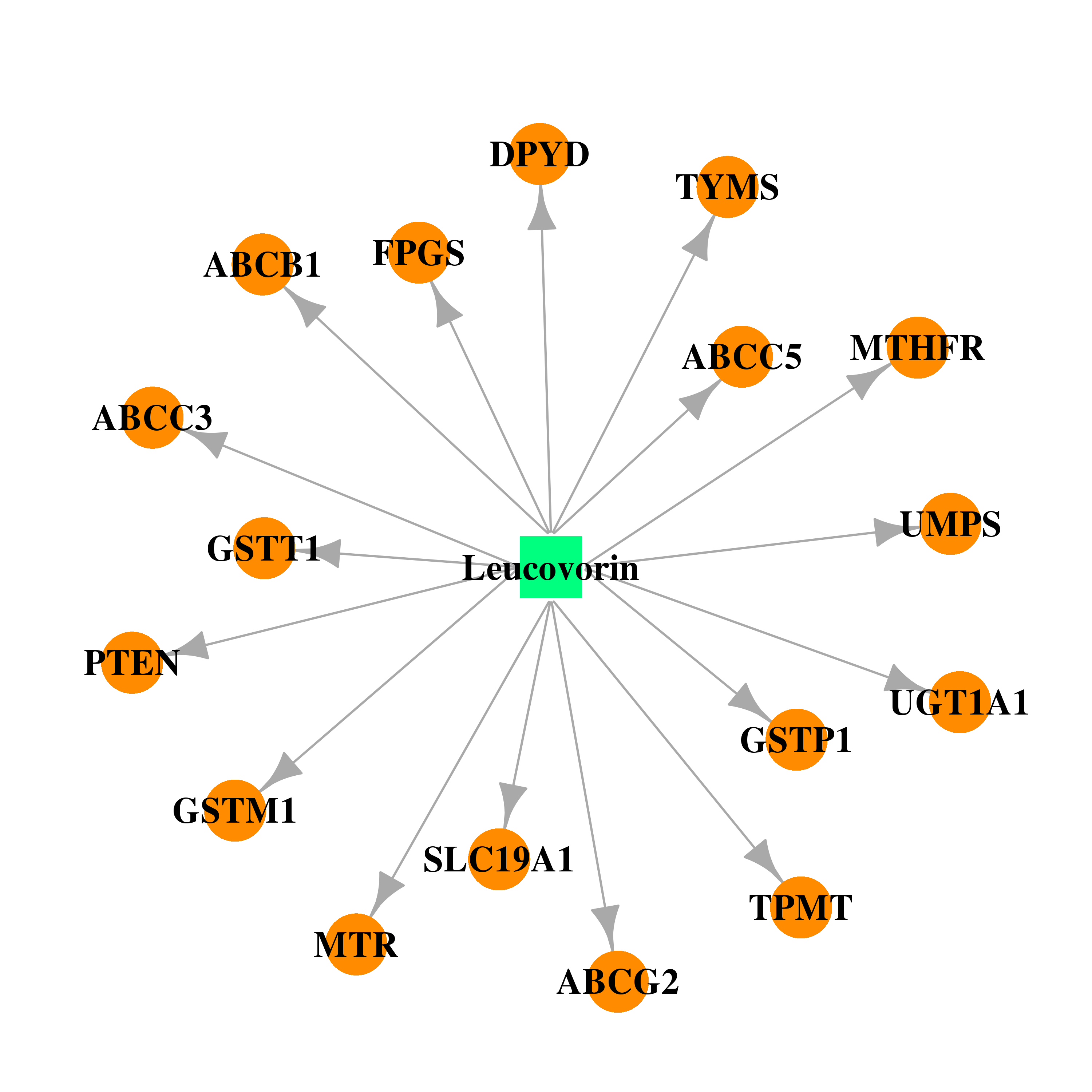 |  |
| DB00526 | dihydropyrimidine dehydrogenase | approved; investigational | Oxaliplatin | 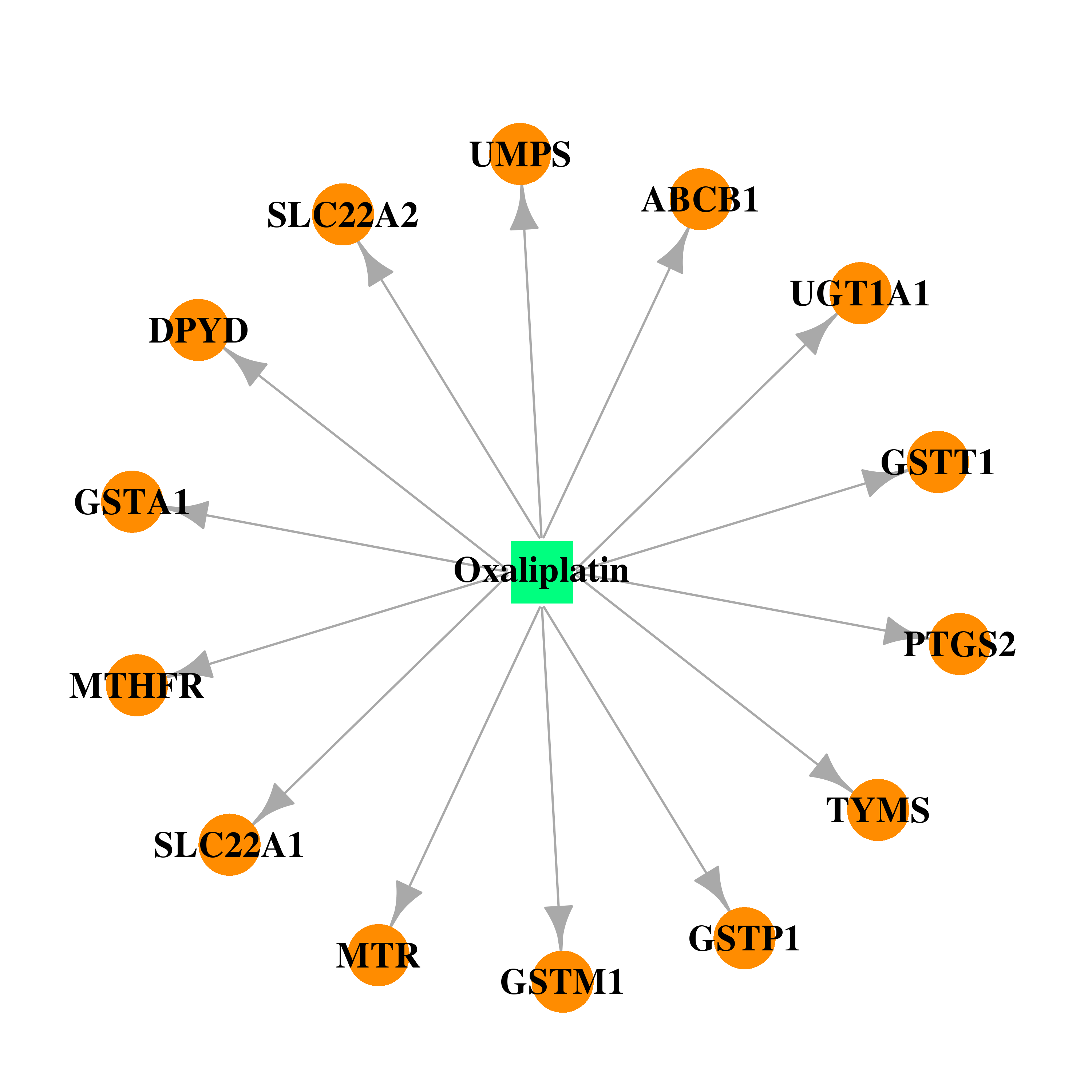 |  |
| DB00293 | dihydropyrimidine dehydrogenase | approved; investigational | Raltitrexed | 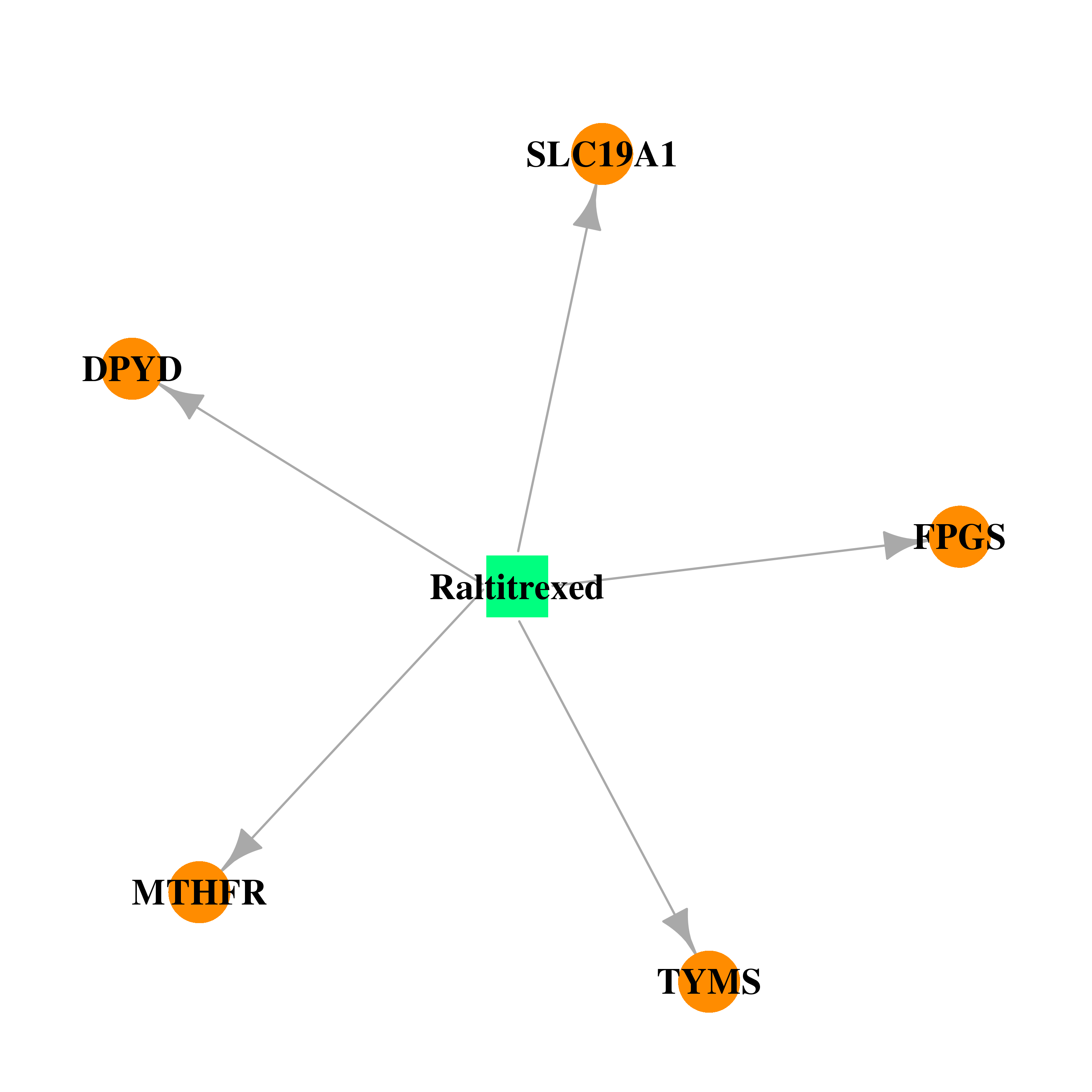 |  |
| DB00563 | dihydropyrimidine dehydrogenase | approved | Methotrexate | 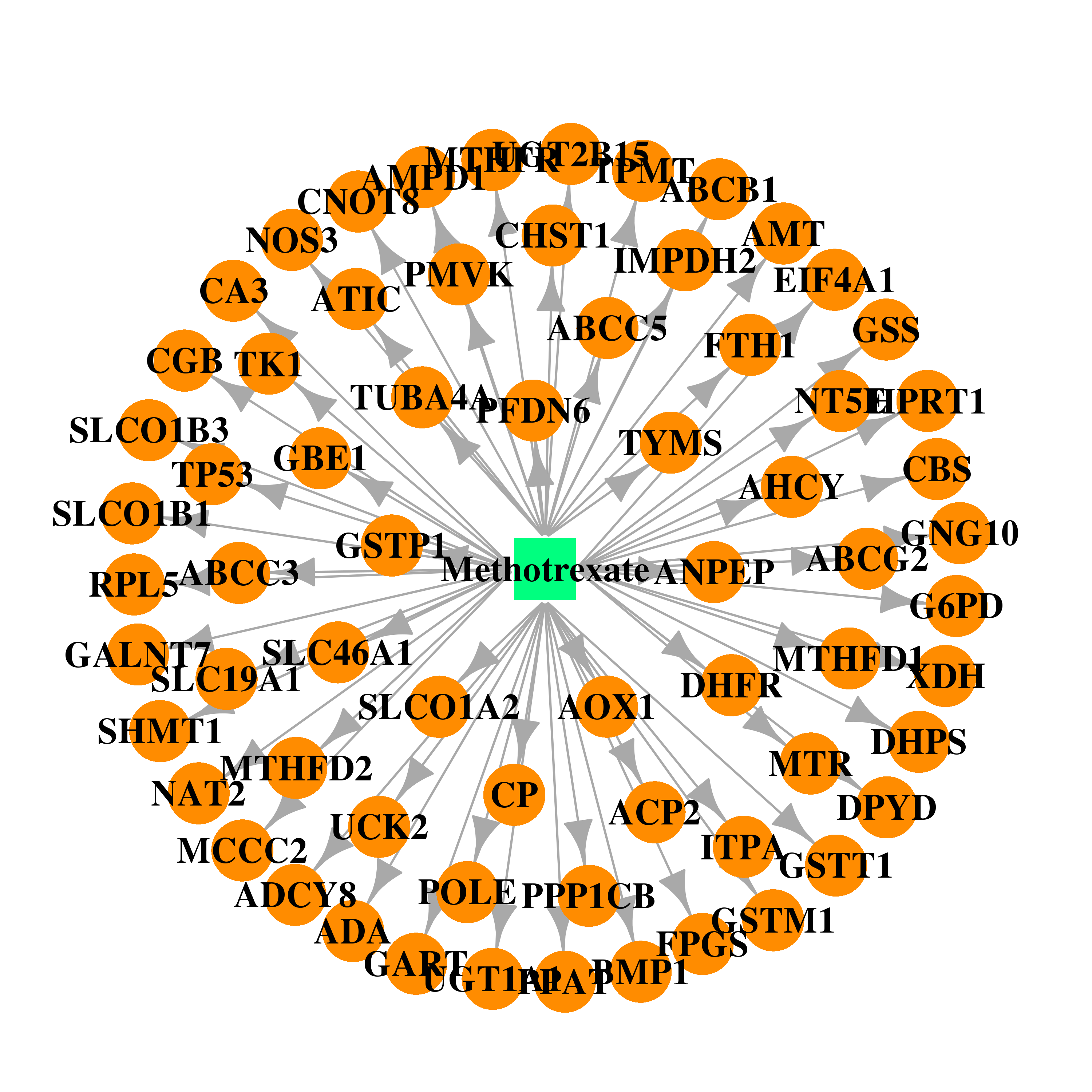 | 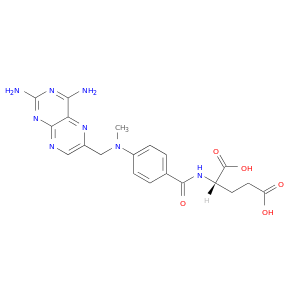 |
| DB01101 | dihydropyrimidine dehydrogenase | approved; investigational | Capecitabine |  | 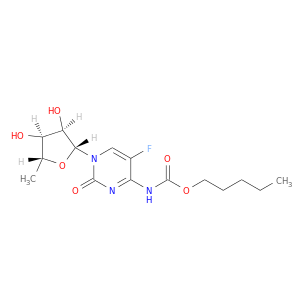 |
| DB00762 | dihydropyrimidine dehydrogenase | approved; investigational | Irinotecan | 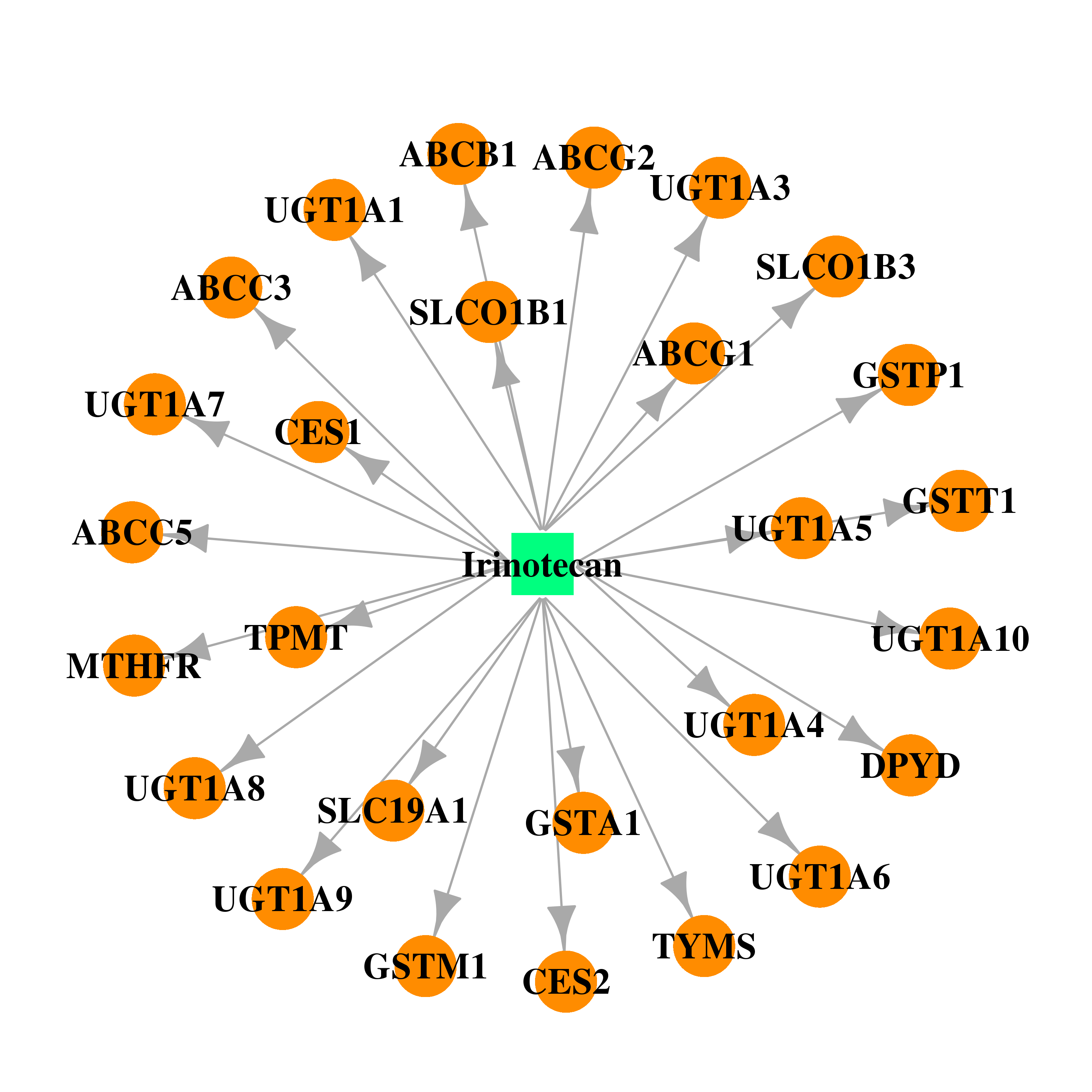 | 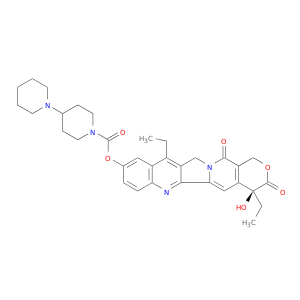 |
| DB01033 | dihydropyrimidine dehydrogenase | approved | Mercaptopurine | 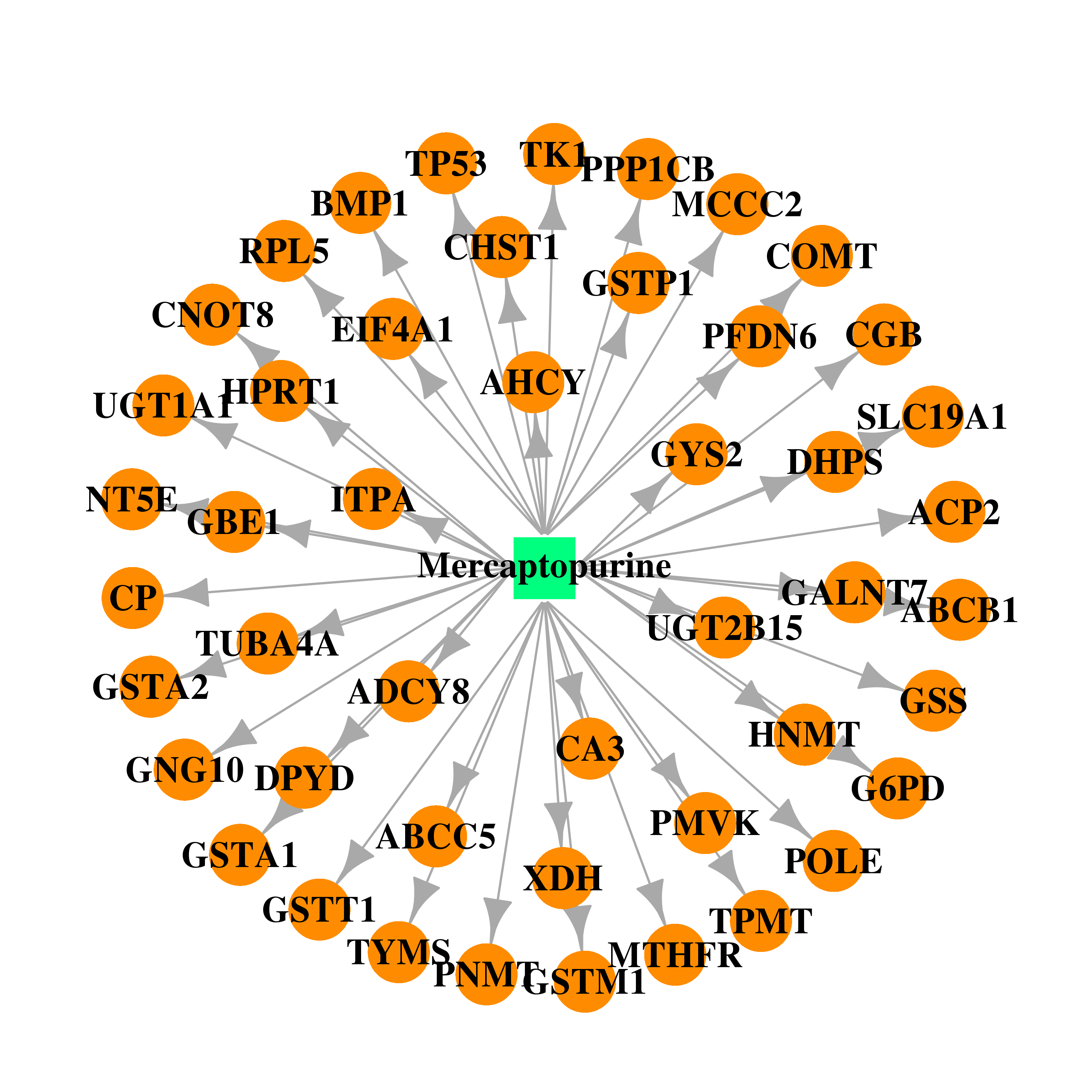 | 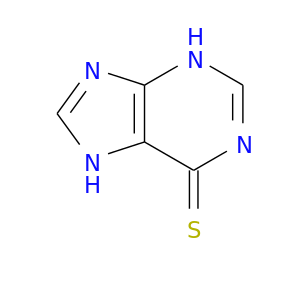 |
| DB00544 | dihydropyrimidine dehydrogenase | approved | Fluorouracil |  |  |
| DB01143 | dihydropyrimidine dehydrogenase | approved; investigational | Amifostine | 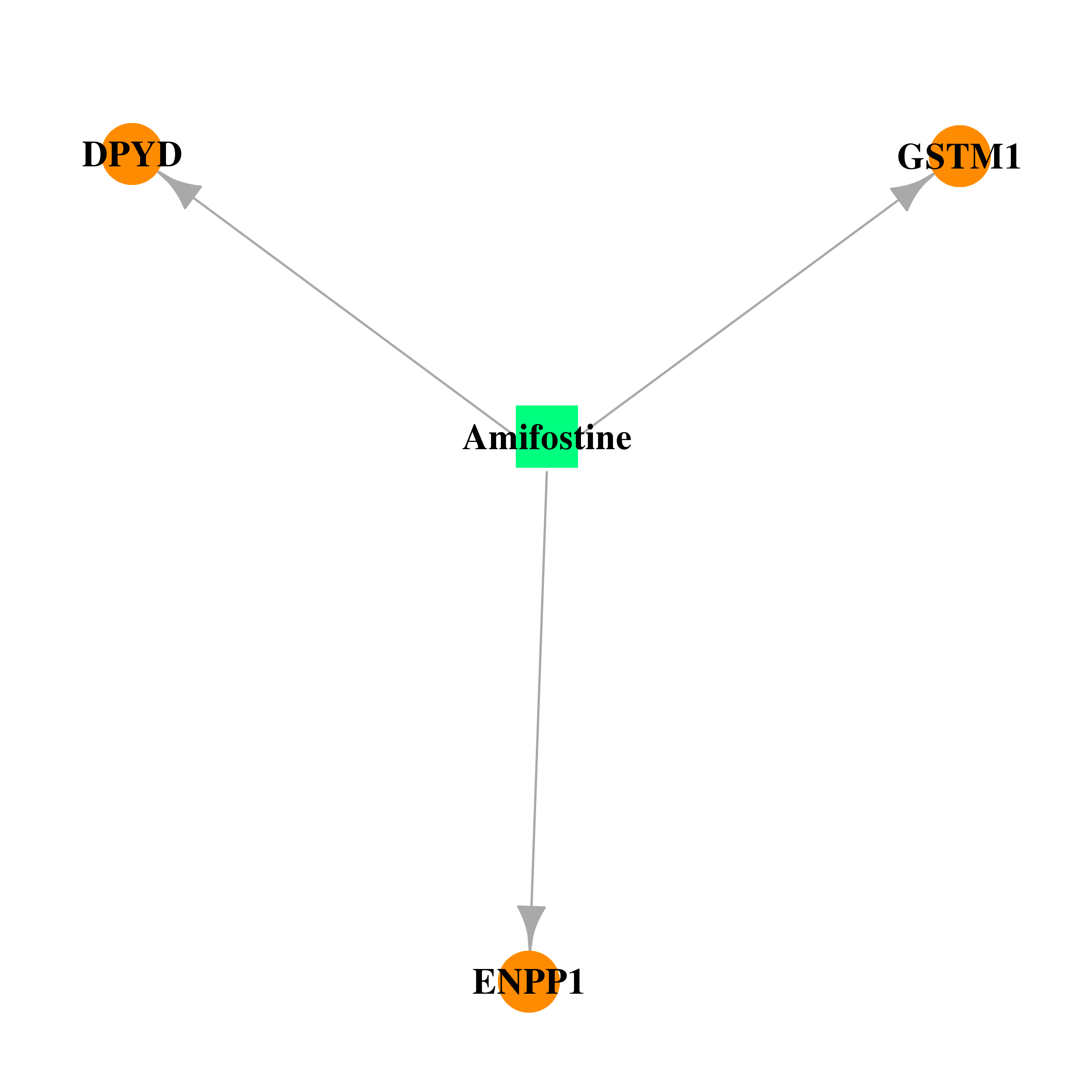 | 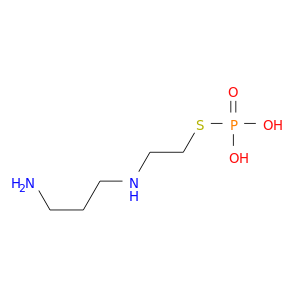 |
| DB00958 | dihydropyrimidine dehydrogenase | approved | Carboplatin | 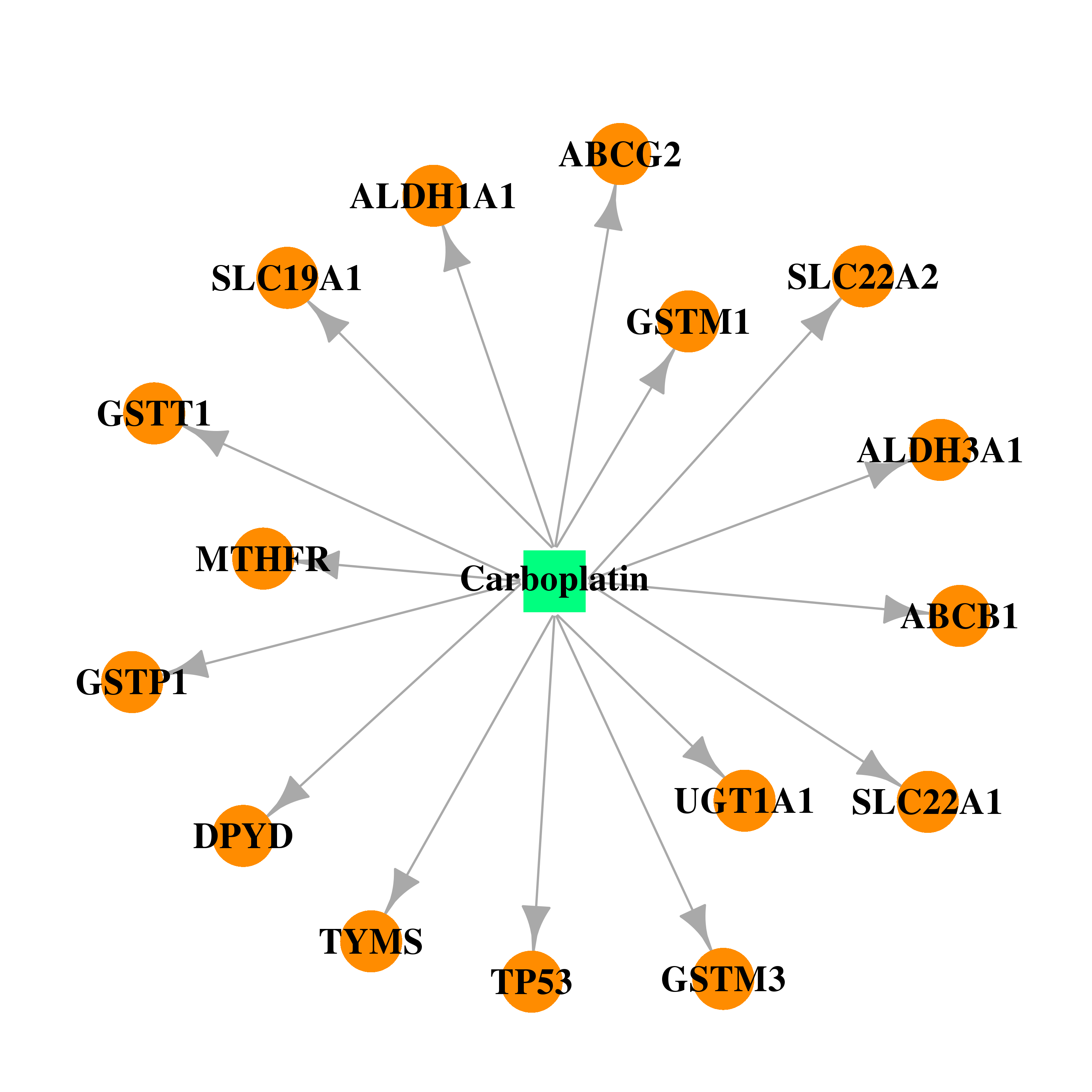 | 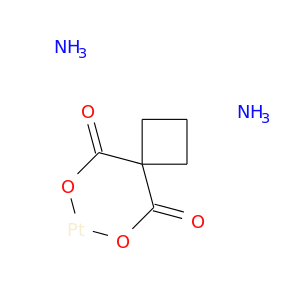 |
| DB01229 | dihydropyrimidine dehydrogenase | approved | Paclitaxel | 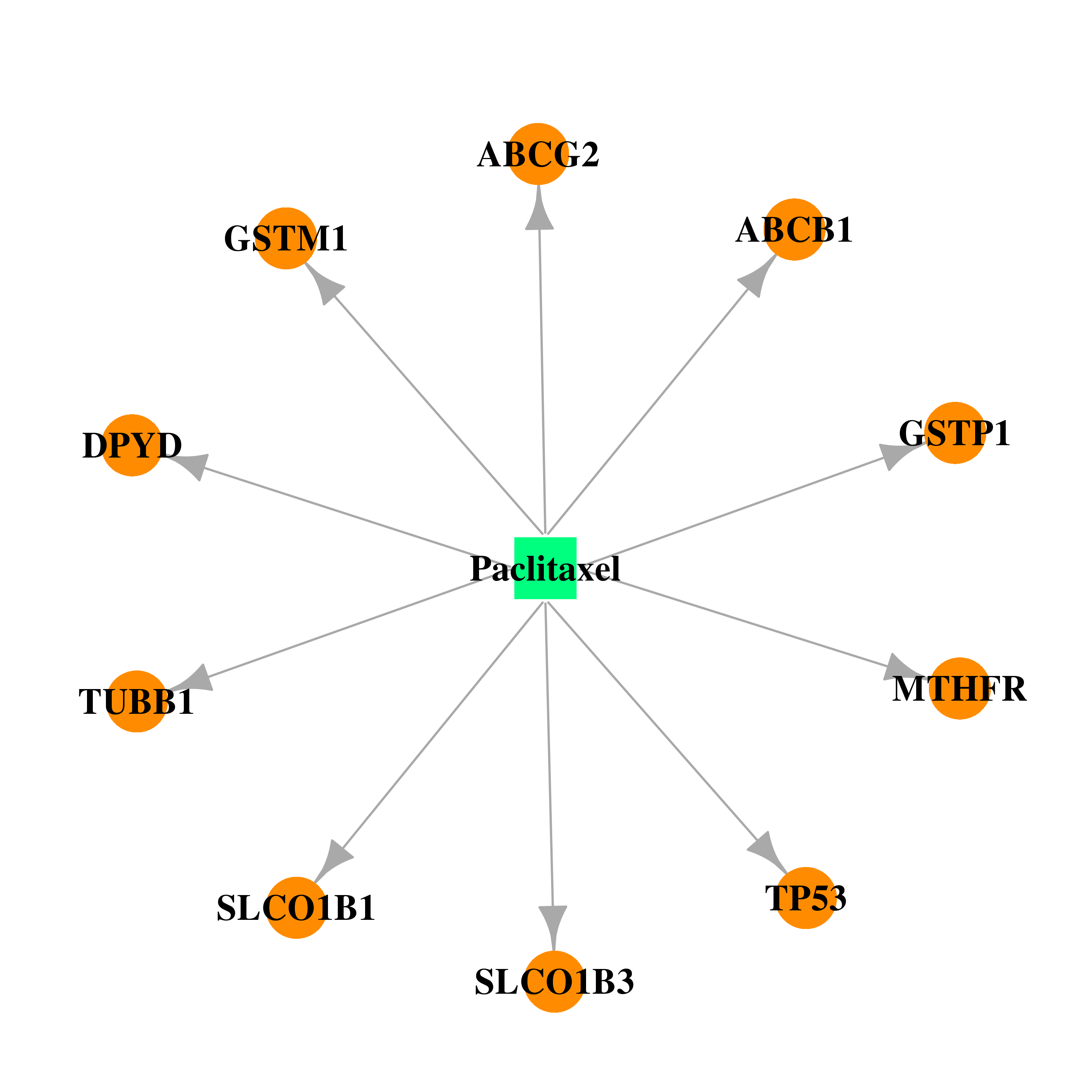 | 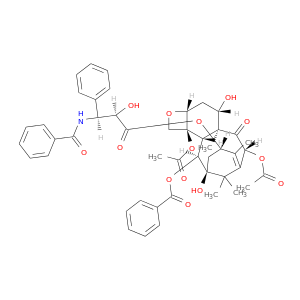 |
| Top |
| Cross referenced IDs for DPYD |
| * We obtained these cross-references from Uniprot database. It covers 150 different DBs, 18 categories. http://www.uniprot.org/help/cross_references_section |
: Open all cross reference information
|
Copyright © 2016-Present - The Univsersity of Texas Health Science Center at Houston @ |