|
||||||||||||||||||||
| |
| Phenotypic Information (metabolism pathway, cancer, disease, phenome) |
| |
| |
| Gene-Gene Network Information: Co-Expression Network, Interacting Genes & KEGG |
| |
|
| Gene Summary for EIF5A2 |
| Basic gene info. | Gene symbol | EIF5A2 |
| Gene name | eukaryotic translation initiation factor 5A2 | |
| Synonyms | EIF-5A2|eIF5AII | |
| Cytomap | UCSC genome browser: 3q26.2 | |
| Genomic location | chr3 :170606203-170626426 | |
| Type of gene | protein-coding | |
| RefGenes | NM_020390.5, | |
| Ensembl id | ENSG00000163577 | |
| Description | eIF-5A-2eukaryotic initiation factor 5Aeukaryotic translation initiation factor 5A-2 | |
| Modification date | 20141207 | |
| dbXrefs | MIM : 605782 | |
| HGNC : HGNC | ||
| Ensembl : ENSG00000163577 | ||
| HPRD : 10426 | ||
| Vega : OTTHUMG00000158958 | ||
| Protein | UniProt: Q9GZV4 go to UniProt's Cross Reference DB Table | |
| Expression | CleanEX: HS_EIF5A2 | |
| BioGPS: 56648 | ||
| Gene Expression Atlas: ENSG00000163577 | ||
| The Human Protein Atlas: ENSG00000163577 | ||
| Pathway | NCI Pathway Interaction Database: EIF5A2 | |
| KEGG: EIF5A2 | ||
| REACTOME: EIF5A2 | ||
| ConsensusPathDB | ||
| Pathway Commons: EIF5A2 | ||
| Metabolism | MetaCyc: EIF5A2 | |
| HUMANCyc: EIF5A2 | ||
| Regulation | Ensembl's Regulation: ENSG00000163577 | |
| miRBase: chr3 :170,606,203-170,626,426 | ||
| TargetScan: NM_020390 | ||
| cisRED: ENSG00000163577 | ||
| Context | iHOP: EIF5A2 | |
| cancer metabolism search in PubMed: EIF5A2 | ||
| UCL Cancer Institute: EIF5A2 | ||
| Assigned class in ccmGDB | A - This gene has a literature evidence and it belongs to cancer gene. | |
| References showing role of EIF5A2 in cancer cell metabolism | 1. Zhu W, Cai MY, Tong ZT, Dong SS, Mai SJ, et al. (2012) Overexpression of EIF5A2 promotes colorectal carcinoma cell aggressiveness by upregulating MTA1 through C-myc to induce epithelial-mesenchymaltransition. Gut 61: 562-575. doi: 10.1136/gutjnl-2011-300207. go to article 2. Wei JH, Cao JZ, Zhang D, Liao B, Zhong WM, et al. (2014) EIF5A2 predicts outcome in localised invasive bladder cancer and promotes bladder cancer cell aggressiveness in vitro and in vivo. Br J Cancer 110: 1767-1777. doi: 10.1038/bjc.2014.52. pmid: 3974079. go to article 3. Li Y, Fu L, Li JB, Qin Y, Zeng TT, et al. (2014) Increased expression of EIF5A2, via hypoxia or gene amplification, contributes to metastasis and angiogenesis of esophageal squamous cell carcinoma. Gastroenterology 146: 1701-1713 e1709. doi: 10.1053/j.gastro.2014.02.029. go to article | |
| Top |
| Phenotypic Information for EIF5A2(metabolism pathway, cancer, disease, phenome) |
| Cancer | CGAP: EIF5A2 |
| Familial Cancer Database: EIF5A2 | |
| * This gene is included in those cancer gene databases. |
|
|
|
|
|
| . | ||||||||||||||
Oncogene 1 | Significant driver gene in | |||||||||||||||||||
| cf) number; DB name 1 Oncogene; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 2 Tumor Suppressor gene; https://bioinfo.uth.edu/TSGene/, 3 Cancer Gene Census; http://www.nature.com/nrc/journal/v4/n3/abs/nrc1299.html, 4 CancerGenes; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 5 Network of Cancer Gene; http://ncg.kcl.ac.uk/index.php, 1Therapeutic Vulnerabilities in Cancer; http://cbio.mskcc.org/cancergenomics/statius/ |
| REACTOME_METABOLISM_OF_PROTEINS | |
| OMIM | 605782; gene. |
| Orphanet | |
| Disease | KEGG Disease: EIF5A2 |
| MedGen: EIF5A2 (Human Medical Genetics with Condition) | |
| ClinVar: EIF5A2 | |
| Phenotype | MGI: EIF5A2 (International Mouse Phenotyping Consortium) |
| PhenomicDB: EIF5A2 | |
| Mutations for EIF5A2 |
| * Under tables are showing count per each tissue to give us broad intuition about tissue specific mutation patterns.You can go to the detailed page for each mutation database's web site. |
| There's no structural variation information in COSMIC data for this gene. |
| * From mRNA Sanger sequences, Chitars2.0 arranged chimeric transcripts. This table shows EIF5A2 related fusion information. |
| ID | Head Gene | Tail Gene | Accession | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a |
| AF262027 | EIF5A2 | 1 | 1431 | 3 | 170610215 | 170625632 | RAD23B | 1428 | 2216 | 9 | 110093684 | 110094473 | |
| Top |
| There's no copy number variation information in COSMIC data for this gene. |
| Top |
|
 |
| Top |
| Stat. for Non-Synonymous SNVs (# total SNVs=13) | (# total SNVs=2) |
 |  |
(# total SNVs=0) | (# total SNVs=0) |
| Top |
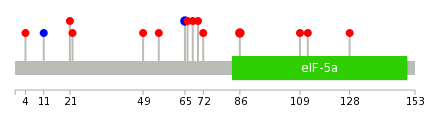
| * When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site,primary_histology,mutation(aa),pubmedID. |
| GRCh37 position | Mutation(aa) | Unique sampleID count |
| chr3:170624791-170624791 | p.R86I | 2 |
| chr3:170612080-170612080 | p.? | 2 |
| chr3:170624853-170624853 | p.T65T | 2 |
| chr3:170612158-170612158 | p.R109C | 1 |
| chr3:170625530-170625530 | p.C22W | 1 |
| chr3:170612213-170612213 | p.? | 1 |
| chr3:170625534-170625534 | p.Q21L | 1 |
| chr3:170625563-170625563 | p.D11D | 1 |
| chr3:170624834-170624834 | p.I72V | 1 |
| chr3:170625586-170625586 | p.E4K | 1 |
| Top |
|
 |
| Point Mutation/ Tissue ID | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 |
| # sample | 2 | 1 | 1 | 1 | 1 | 3 | ||||||||||||||
| # mutation | 2 | 1 | 1 | 1 | 1 | 3 | ||||||||||||||
| nonsynonymous SNV | 2 | 1 | 1 | 1 | 1 | 2 | ||||||||||||||
| synonymous SNV | 1 |
| cf) Tissue ID; Tissue type (1; BLCA[Bladder Urothelial Carcinoma], 2; BRCA[Breast invasive carcinoma], 3; CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], 4; COAD[Colon adenocarcinoma], 5; GBM[Glioblastoma multiforme], 6; Glioma Low Grade, 7; HNSC[Head and Neck squamous cell carcinoma], 8; KICH[Kidney Chromophobe], 9; KIRC[Kidney renal clear cell carcinoma], 10; KIRP[Kidney renal papillary cell carcinoma], 11; LAML[Acute Myeloid Leukemia], 12; LUAD[Lung adenocarcinoma], 13; LUSC[Lung squamous cell carcinoma], 14; OV[Ovarian serous cystadenocarcinoma ], 15; PAAD[Pancreatic adenocarcinoma], 16; PRAD[Prostate adenocarcinoma], 17; SKCM[Skin Cutaneous Melanoma], 18:STAD[Stomach adenocarcinoma], 19:THCA[Thyroid carcinoma], 20:UCEC[Uterine Corpus Endometrial Carcinoma]) |
| Top |
| * We represented just top 10 SNVs. When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site, primary_histology, mutation(aa), pubmedID. |
| Genomic Position | Mutation(aa) | Unique sampleID count |
| chr3:170624845 | p.G66D | 1 |
| chr3:170624851 | p.G49R | 1 |
| chr3:170625451 | p.D11D | 1 |
| chr3:170625563 | p.E4K | 1 |
| chr3:170625586 | p.N128I | 1 |
| chr3:170612100 | p.L112I | 1 |
| chr3:170612149 | p.R109C | 1 |
| chr3:170612158 | p.R86I | 1 |
| chr3:170624791 | p.K68T | 1 |
| * Copy number data were extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered on Jan-05-2015. Function ProcessCNAData in TCGA-Assembler package was used to obtain gene-level copy number value which is calculated as the average copy number of the genomic region of a gene. |
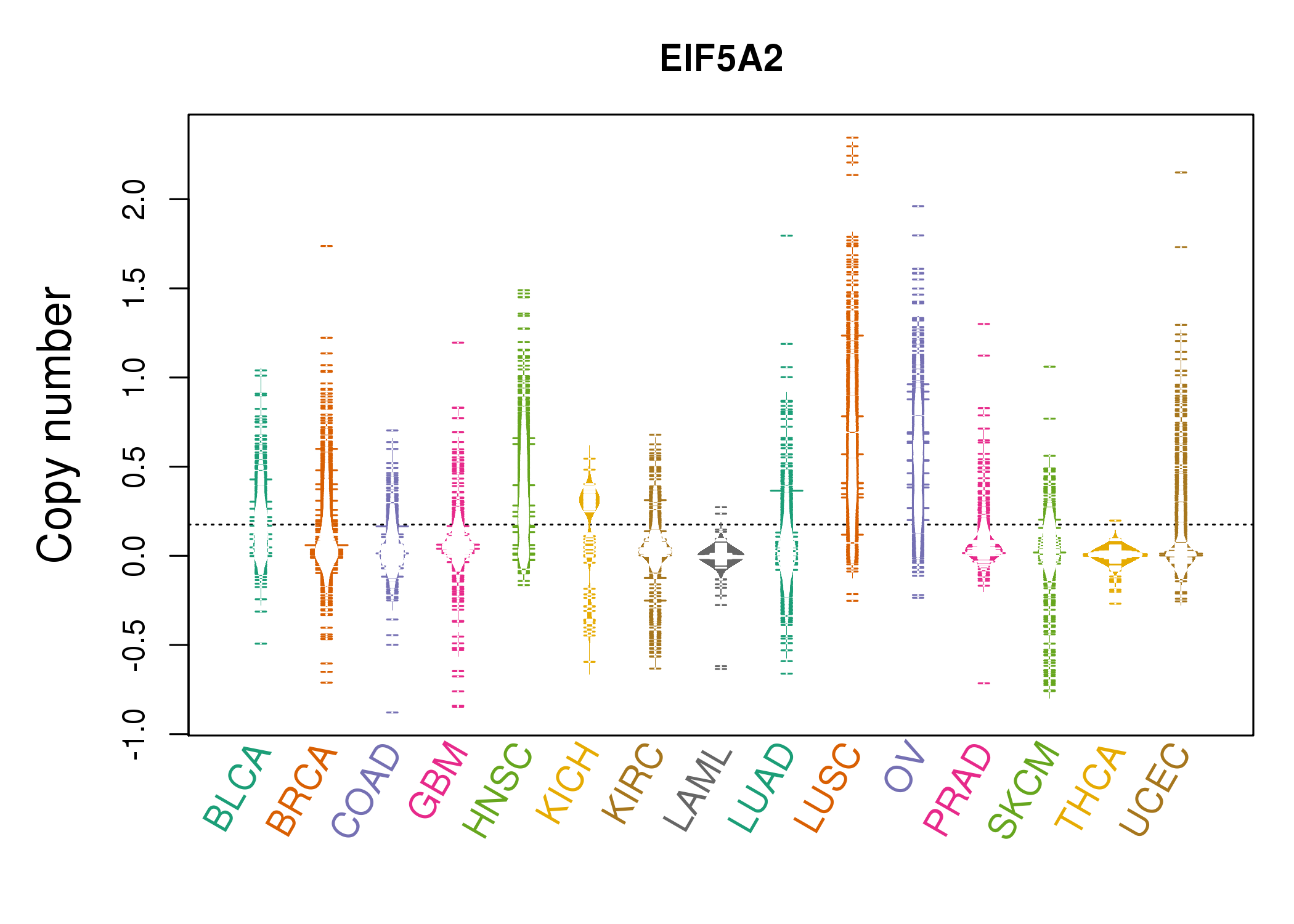 |
| cf) Tissue ID[Tissue type]: BLCA[Bladder Urothelial Carcinoma], BRCA[Breast invasive carcinoma], CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], COAD[Colon adenocarcinoma], GBM[Glioblastoma multiforme], Glioma Low Grade, HNSC[Head and Neck squamous cell carcinoma], KICH[Kidney Chromophobe], KIRC[Kidney renal clear cell carcinoma], KIRP[Kidney renal papillary cell carcinoma], LAML[Acute Myeloid Leukemia], LUAD[Lung adenocarcinoma], LUSC[Lung squamous cell carcinoma], OV[Ovarian serous cystadenocarcinoma ], PAAD[Pancreatic adenocarcinoma], PRAD[Prostate adenocarcinoma], SKCM[Skin Cutaneous Melanoma], STAD[Stomach adenocarcinoma], THCA[Thyroid carcinoma], UCEC[Uterine Corpus Endometrial Carcinoma] |
| Top |
| Gene Expression for EIF5A2 |
| * CCLE gene expression data were extracted from CCLE_Expression_Entrez_2012-10-18.res: Gene-centric RMA-normalized mRNA expression data. |
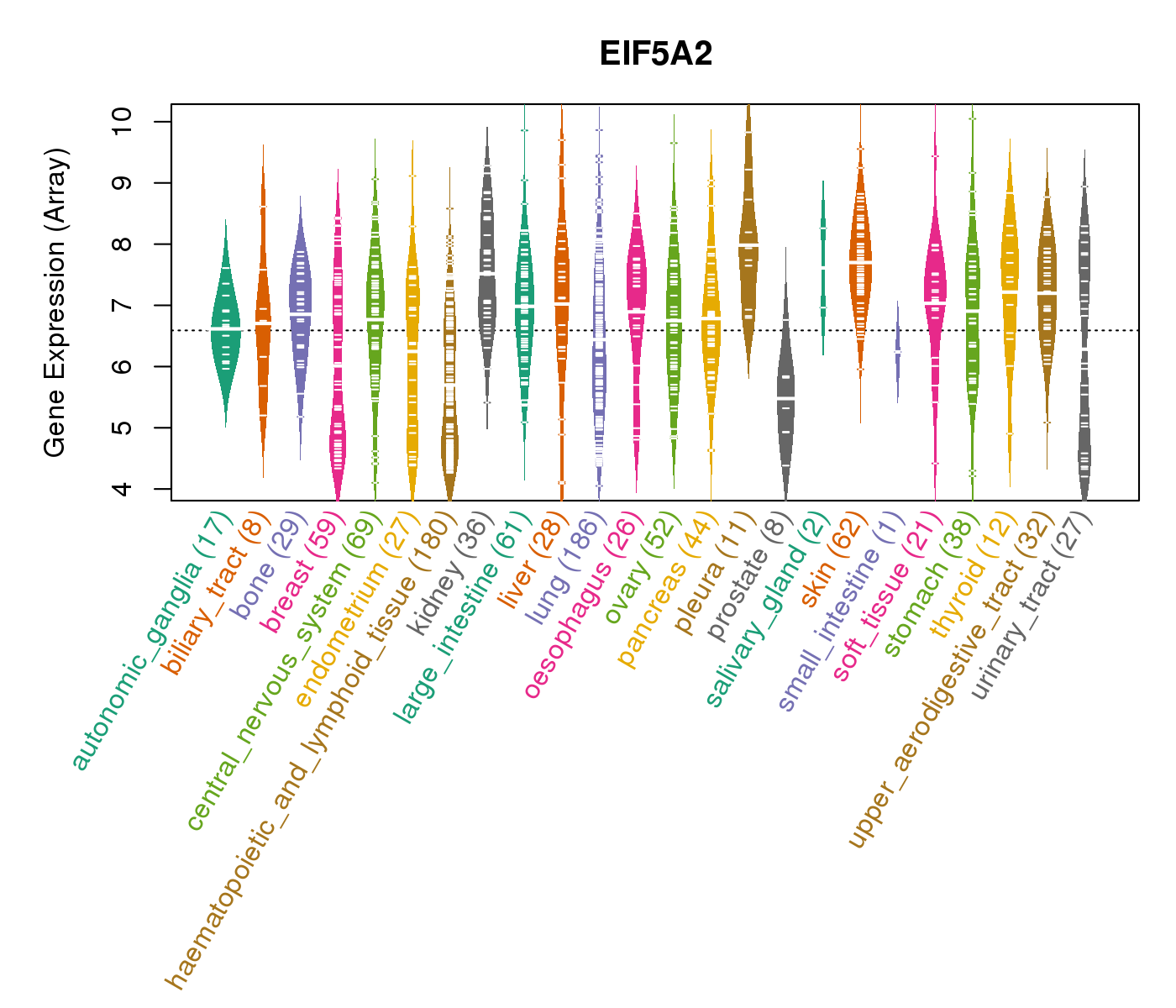 |
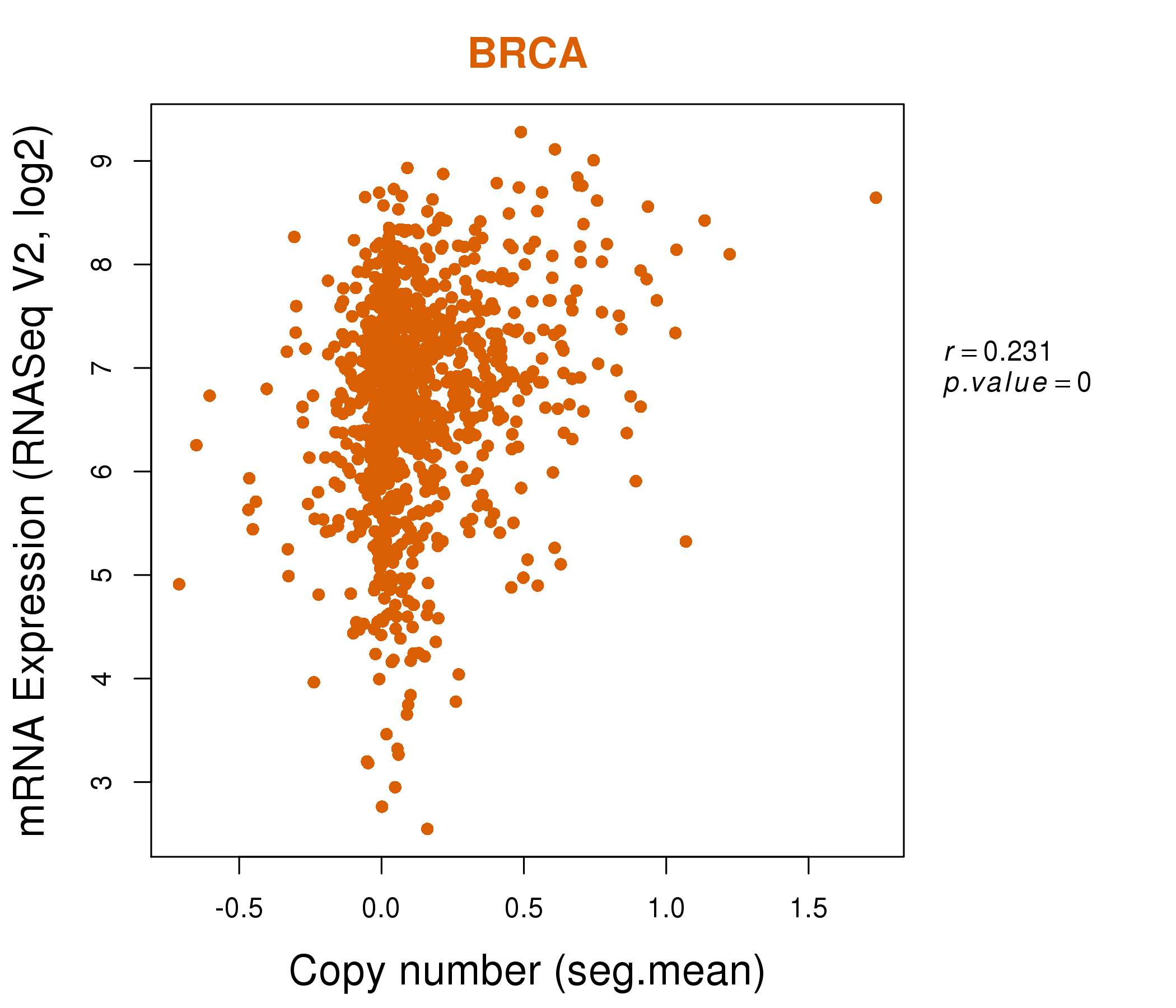
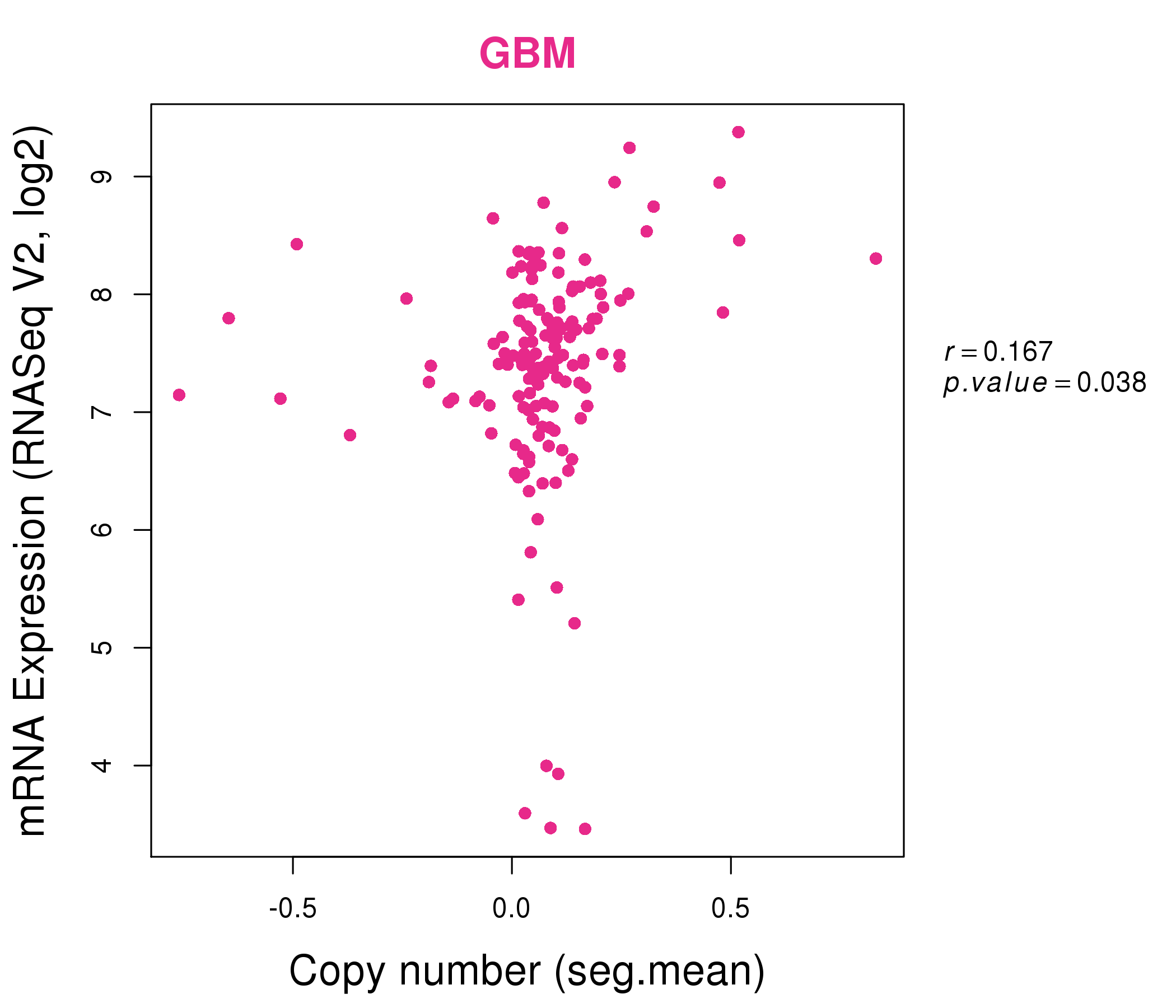
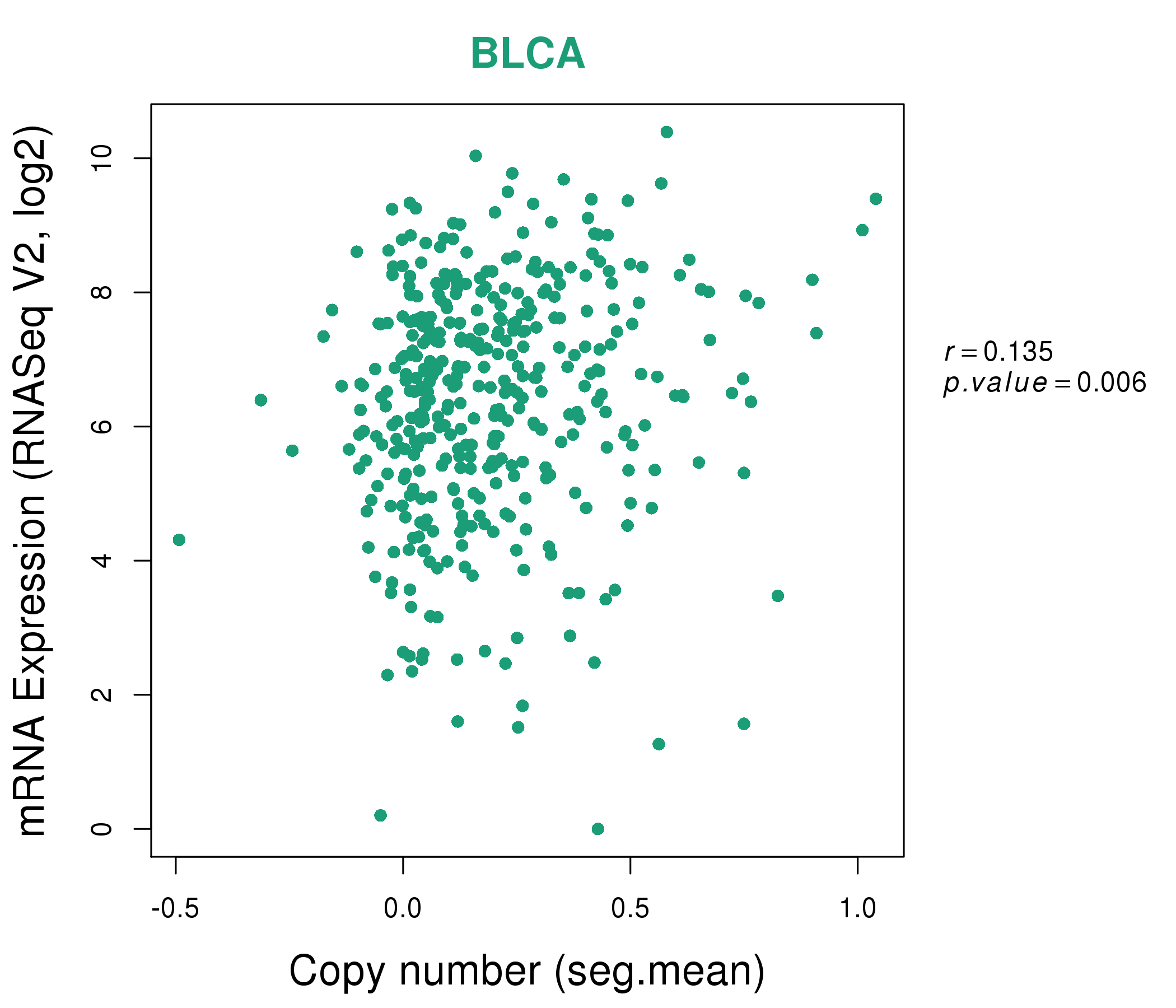
| * Normalized gene expression data of RNASeqV2 was extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered at Jan-05-2015. Only eight cancer types have enough normal control samples for differential expression analysis. (t test, adjusted p<0.05 (using Benjamini-Hochberg FDR)) |
 |
| Top |
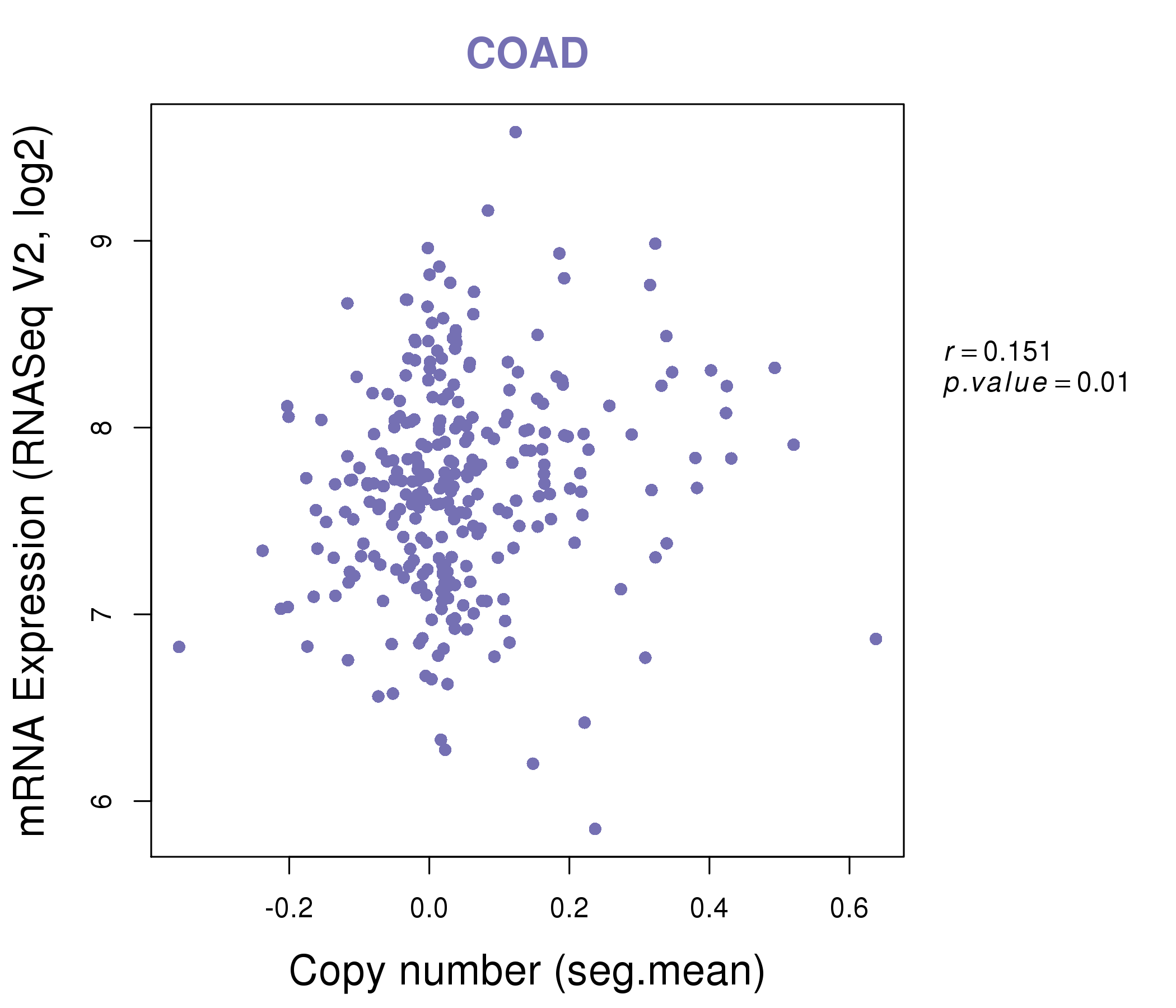
| * This plots show the correlation between CNV and gene expression. |
: Open all plots for all cancer types
 |
|
 |
|
| Top |
| Gene-Gene Network Information |
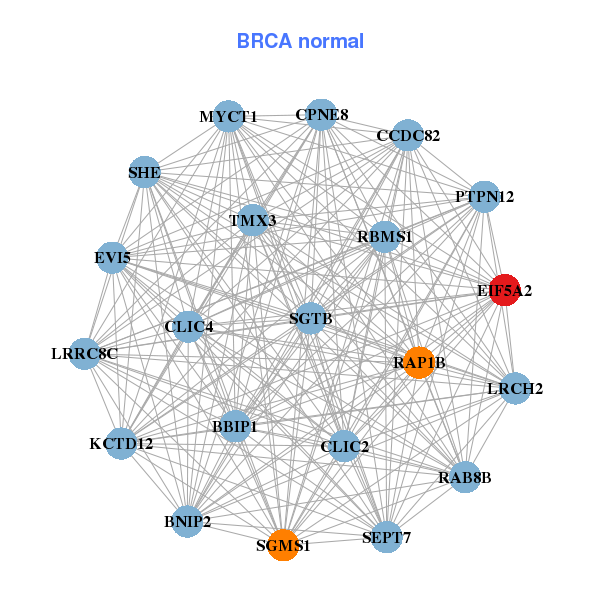
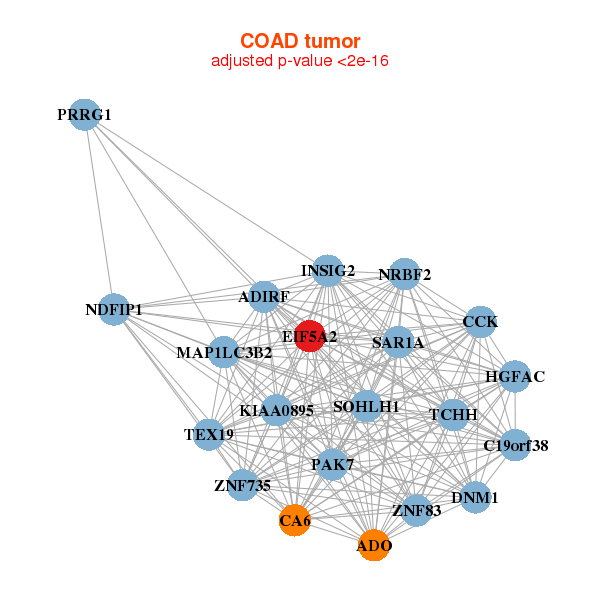
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
 |
|
| CALD1,CLIC4,CSGALNACT2,DACT1,DPYSL3,DSE,EIF5A2, FAP,FERMT2,FNDC3B,FSTL1,IKBIP,ITGB1,LATS2, LOX,PKD2,PLS3,RECK,SEC23A,TIMP2,TRPC1 | BNIP2,CCDC82,CLIC2,CLIC4,CPNE8,EIF5A2,EVI5, KCTD12,LRCH2,LRRC8C,MYCT1,BBIP1,PTPN12,RAB8B, RAP1B,RBMS1,SEPT7,SGMS1,SGTB,SHE,TMX3 |
 |
|
| ADO,ADIRF,C19orf38,CA6,CCK,DNM1,EIF5A2, HGFAC,INSIG2,KIAA0895,MAP1LC3B2,NDFIP1,NRBF2,PAK7, PRRG1,SAR1A,SOHLH1,TCHH,TEX19,ZNF735,ZNF83 | BLVRA,FAM229B,EIF5A2,FAM195B,FEZ1,GNB4,GUCY1B3, HDGFRP3,LOC399959,LOC644538,MRAS,SLC9B2,NME4,RAB23, RAB31,IFT22,RHOQ,RND2,SNCA,SPG20,ZNF25 |
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
| Top |
: Open all interacting genes' information including KEGG pathway for all interacting genes from DAVID
| Top |
| Pharmacological Information for EIF5A2 |
| There's no related Drug. |
| Top |
| Cross referenced IDs for EIF5A2 |
| * We obtained these cross-references from Uniprot database. It covers 150 different DBs, 18 categories. http://www.uniprot.org/help/cross_references_section |
: Open all cross reference information
|
Copyright © 2016-Present - The Univsersity of Texas Health Science Center at Houston @ |