|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |
| Phenotypic Information (metabolism pathway, cancer, disease, phenome) |
| |
| |
| Gene-Gene Network Information: Co-Expression Network, Interacting Genes & KEGG |
| |
|
| Gene Summary for PRKACA |
| Top |
| Phenotypic Information for PRKACA(metabolism pathway, cancer, disease, phenome) |
| Cancer | CGAP: PRKACA |
| Familial Cancer Database: PRKACA | |
| * This gene is included in those cancer gene databases. |
|
|
|
|
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Oncogene 1 | Significant driver gene in | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| cf) number; DB name 1 Oncogene; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 2 Tumor Suppressor gene; https://bioinfo.uth.edu/TSGene/, 3 Cancer Gene Census; http://www.nature.com/nrc/journal/v4/n3/abs/nrc1299.html, 4 CancerGenes; http://nar.oxfordjournals.org/content/35/suppl_1/D721.long, 5 Network of Cancer Gene; http://ncg.kcl.ac.uk/index.php, 1Therapeutic Vulnerabilities in Cancer; http://cbio.mskcc.org/cancergenomics/statius/ |
| REACTOME_INTEGRATION_OF_ENERGY_METABOLISM REACTOME_METABOLISM_OF_LIPIDS_AND_LIPOPROTEINS REACTOME_METABOLISM_OF_CARBOHYDRATES REACTOME_GLUCOSE_METABOLISM | |
| OMIM | 601639; gene. 615830; phenotype. |
| Orphanet | 401920; Fibrolamellar hepatocellular carcinoma. |
| Disease | KEGG Disease: PRKACA |
| MedGen: PRKACA (Human Medical Genetics with Condition) | |
| ClinVar: PRKACA | |
| Phenotype | MGI: PRKACA (International Mouse Phenotyping Consortium) |
| PhenomicDB: PRKACA | |
| Mutations for PRKACA |
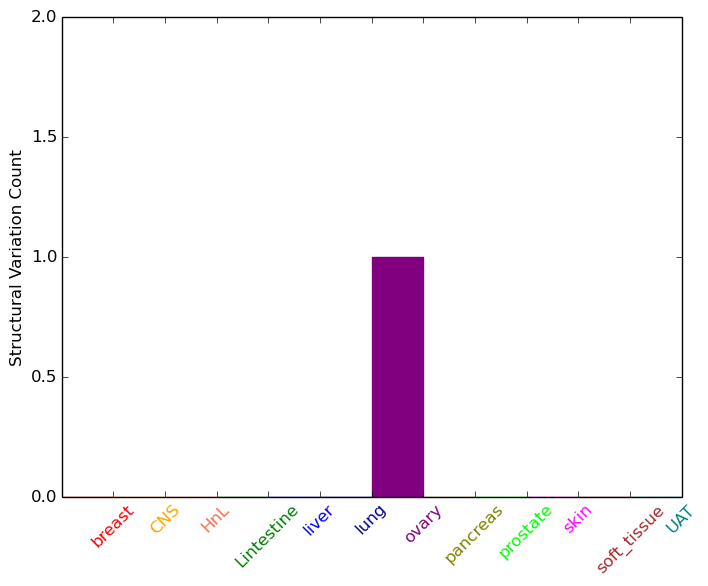
| * Under tables are showing count per each tissue to give us broad intuition about tissue specific mutation patterns.You can go to the detailed page for each mutation database's web site. |
| - Statistics for Tissue and Mutation type | Top |
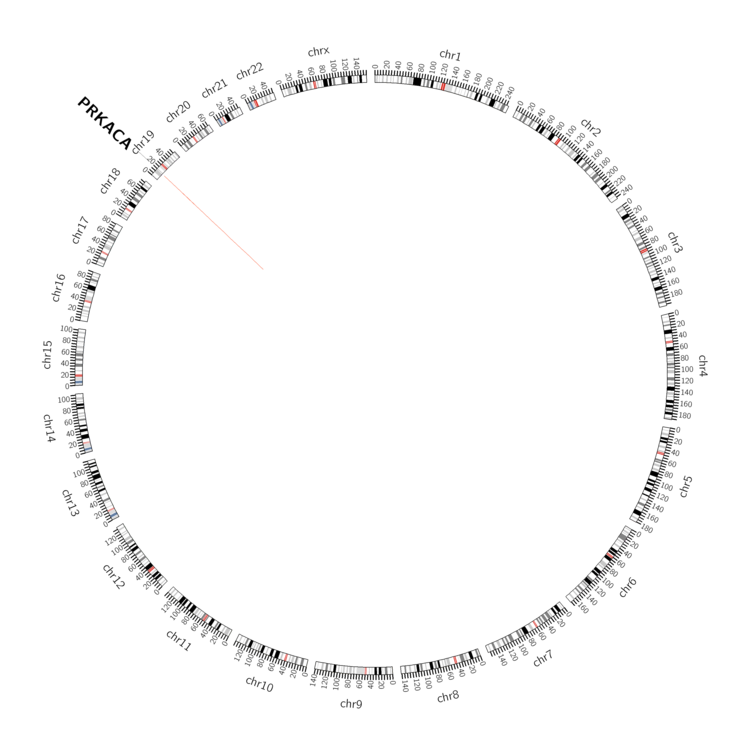 |
| - For Inter-chromosomal Variations |
| There's no inter-chromosomal structural variation. |
| - For Intra-chromosomal Variations |
| * Intra-chromosomal variantions includes 'intrachromosomal amplicon to amplicon', 'intrachromosomal amplicon to non-amplified dna', 'intrachromosomal deletion', 'intrachromosomal fold-back inversion', 'intrachromosomal inversion', 'intrachromosomal tandem duplication', 'Intrachromosomal unknown type', 'intrachromosomal with inverted orientation', 'intrachromosomal with non-inverted orientation'. |
 |
| Sample | Symbol_a | Chr_a | Start_a | End_a | Symbol_b | Chr_b | Start_b | End_b |
| ovary | PRKACA | chr19 | 14219322 | 14219342 | PRKACA | chr19 | 14224487 | 14224507 |
| cf) Tissue number; Tissue name (1;Breast, 2;Central_nervous_system, 3;Haematopoietic_and_lymphoid_tissue, 4;Large_intestine, 5;Liver, 6;Lung, 7;Ovary, 8;Pancreas, 9;Prostate, 10;Skin, 11;Soft_tissue, 12;Upper_aerodigestive_tract) |
| * From mRNA Sanger sequences, Chitars2.0 arranged chimeric transcripts. This table shows PRKACA related fusion information. |
| ID | Head Gene | Tail Gene | Accession | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a | Gene_a | qStart_a | qEnd_a | Chromosome_a | tStart_a | tEnd_a |
| DR006627 | PRKACA | 23 | 456 | 19 | 14217572 | 14228556 | ENTPD1 | 452 | 694 | 10 | 97627439 | 97627682 | |
| BQ278860 | PRKACA | 1 | 531 | 19 | 14203311 | 14203841 | NDOR1 | 530 | 898 | 9 | 140110420 | 140110940 | |
| Top |
| Mutation type/ Tissue ID | brca | cns | cerv | endome | haematopo | kidn | Lintest | liver | lung | ns | ovary | pancre | prost | skin | stoma | thyro | urina | |||
| Total # sample | 1 | |||||||||||||||||||
| GAIN (# sample) | 1 | |||||||||||||||||||
| LOSS (# sample) |
| cf) Tissue ID; Tissue type (1; Breast, 2; Central_nervous_system, 3; Cervix, 4; Endometrium, 5; Haematopoietic_and_lymphoid_tissue, 6; Kidney, 7; Large_intestine, 8; Liver, 9; Lung, 10; NS, 11; Ovary, 12; Pancreas, 13; Prostate, 14; Skin, 15; Stomach, 16; Thyroid, 17; Urinary_tract) |
| Top |
|
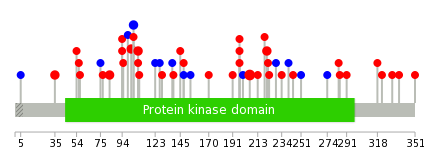 |
| Top |
| Stat. for Non-Synonymous SNVs (# total SNVs=35) | (# total SNVs=13) |
 | 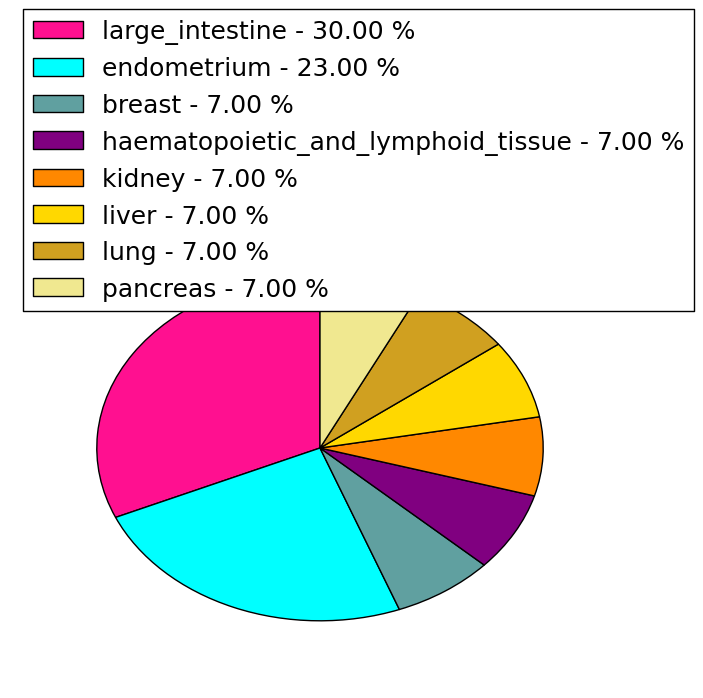 |
(# total SNVs=1) | (# total SNVs=1) |
 |  |
| Top |
| * When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site,primary_histology,mutation(aa),pubmedID. |
| GRCh37 position | Mutation(aa) | Unique sampleID count |
| chr19:14208416-14208416 | p.L206R | 4 |
| chr19:14213642-14213642 | p.E108Q | 3 |
| chr19:14208444-14208444 | p.W197G | 2 |
| chr19:14213716-14213716 | p.L83P | 2 |
| chr19:14208277-14208277 | p.D221N | 2 |
| chr19:14213652-14213652 | p.L104L | 2 |
| chr19:14213659-14213659 | p.P102L | 2 |
| chr19:14218165-14218165 | p.A35S | 2 |
| chr19:14208433-14208433 | p.C200C | 1 |
| chr19:14218178-14218179 | p.W31fs*15 | 1 |
| Top |
|
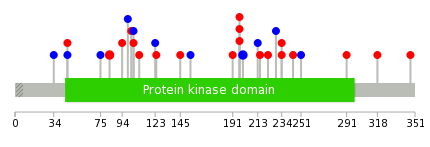 |
| Point Mutation/ Tissue ID | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 |
| # sample | 3 | 4 | 4 | 2 | 3 | 1 | 1 | 3 | 4 | 7 | ||||||||||
| # mutation | 3 | 3 | 4 | 2 | 3 | 1 | 1 | 3 | 4 | 8 | ||||||||||
| nonsynonymous SNV | 2 | 1 | 4 | 1 | 2 | 1 | 1 | 3 | 1 | 5 | ||||||||||
| synonymous SNV | 1 | 2 | 1 | 1 | 3 | 3 |
| cf) Tissue ID; Tissue type (1; BLCA[Bladder Urothelial Carcinoma], 2; BRCA[Breast invasive carcinoma], 3; CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], 4; COAD[Colon adenocarcinoma], 5; GBM[Glioblastoma multiforme], 6; Glioma Low Grade, 7; HNSC[Head and Neck squamous cell carcinoma], 8; KICH[Kidney Chromophobe], 9; KIRC[Kidney renal clear cell carcinoma], 10; KIRP[Kidney renal papillary cell carcinoma], 11; LAML[Acute Myeloid Leukemia], 12; LUAD[Lung adenocarcinoma], 13; LUSC[Lung squamous cell carcinoma], 14; OV[Ovarian serous cystadenocarcinoma ], 15; PAAD[Pancreatic adenocarcinoma], 16; PRAD[Prostate adenocarcinoma], 17; SKCM[Skin Cutaneous Melanoma], 18:STAD[Stomach adenocarcinoma], 19:THCA[Thyroid carcinoma], 20:UCEC[Uterine Corpus Endometrial Carcinoma]) |
| Top |
| * We represented just top 10 SNVs. When you move the cursor on each content, you can see more deailed mutation information on the Tooltip. Those are primary_site, primary_histology, mutation(aa), pubmedID. |
| Genomic Position | Mutation(aa) | Unique sampleID count |
| chr19:14208444 | p.A234T,PRKACA | 2 |
| chr19:14213716 | p.L83P,PRKACA | 2 |
| chr19:14208238 | p.C200C,PRKACA | 2 |
| chr19:14208433 | p.W197R,PRKACA | 2 |
| chr19:14213684 | p.I251I,PRKACA | 1 |
| chr19:14204497 | p.L104L,PRKACA | 1 |
| chr19:14208462 | p.P244L,PRKACA | 1 |
| chr19:14208185 | p.L104F,PRKACA | 1 |
| chr19:14208660 | p.P102L,PRKACA | 1 |
| chr19:14217584 | p.I229I,PRKACA | 1 |
| * Copy number data were extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered on Jan-05-2015. Function ProcessCNAData in TCGA-Assembler package was used to obtain gene-level copy number value which is calculated as the average copy number of the genomic region of a gene. |
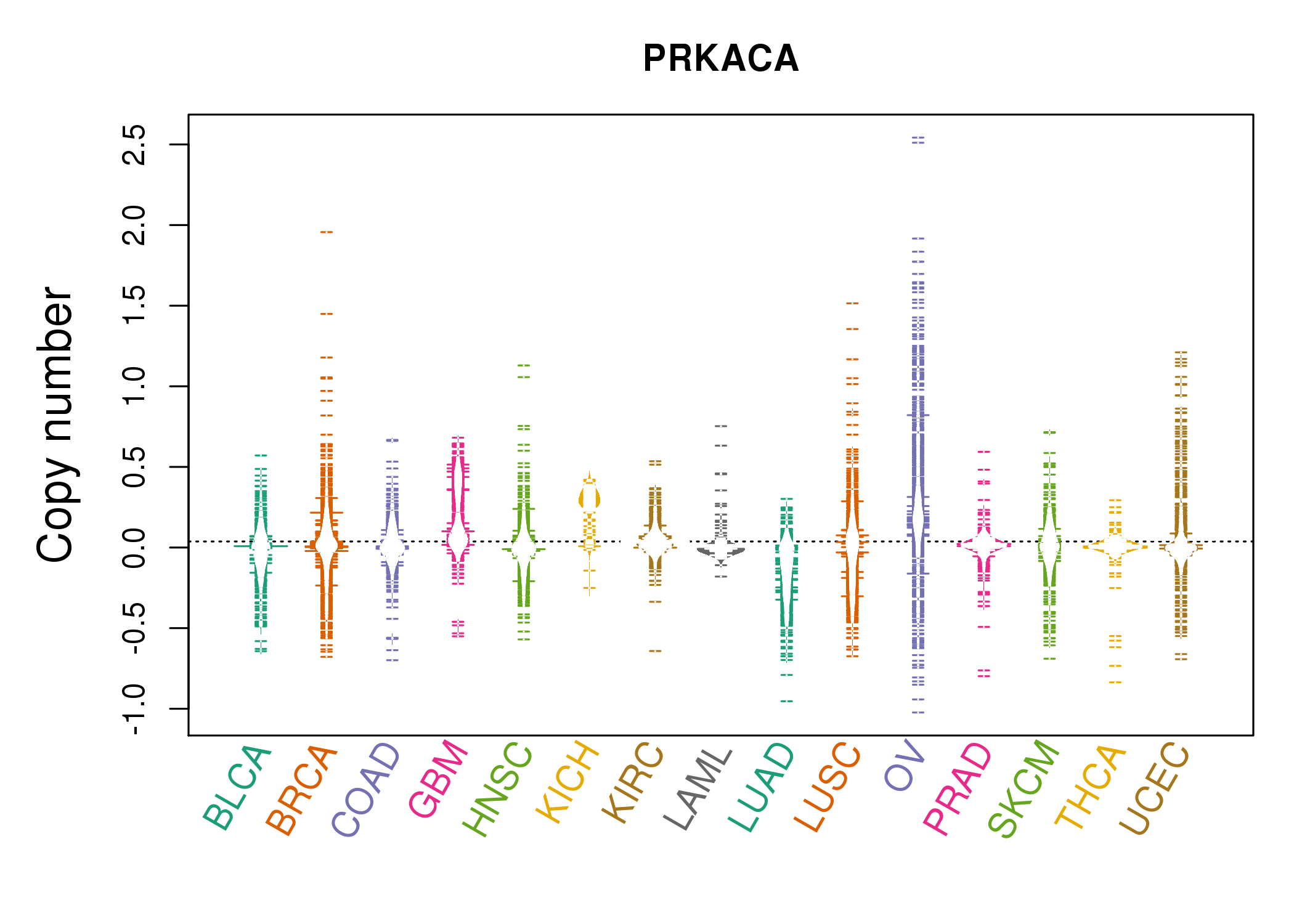 |
| cf) Tissue ID[Tissue type]: BLCA[Bladder Urothelial Carcinoma], BRCA[Breast invasive carcinoma], CESC[Cervical squamous cell carcinoma and endocervical adenocarcinoma], COAD[Colon adenocarcinoma], GBM[Glioblastoma multiforme], Glioma Low Grade, HNSC[Head and Neck squamous cell carcinoma], KICH[Kidney Chromophobe], KIRC[Kidney renal clear cell carcinoma], KIRP[Kidney renal papillary cell carcinoma], LAML[Acute Myeloid Leukemia], LUAD[Lung adenocarcinoma], LUSC[Lung squamous cell carcinoma], OV[Ovarian serous cystadenocarcinoma ], PAAD[Pancreatic adenocarcinoma], PRAD[Prostate adenocarcinoma], SKCM[Skin Cutaneous Melanoma], STAD[Stomach adenocarcinoma], THCA[Thyroid carcinoma], UCEC[Uterine Corpus Endometrial Carcinoma] |
| Top |
| Gene Expression for PRKACA |
| * CCLE gene expression data were extracted from CCLE_Expression_Entrez_2012-10-18.res: Gene-centric RMA-normalized mRNA expression data. |
 |
| * Normalized gene expression data of RNASeqV2 was extracted from TCGA using R package TCGA-Assembler. The URLs of all public data files on TCGA DCC data server were gathered at Jan-05-2015. Only eight cancer types have enough normal control samples for differential expression analysis. (t test, adjusted p<0.05 (using Benjamini-Hochberg FDR)) |
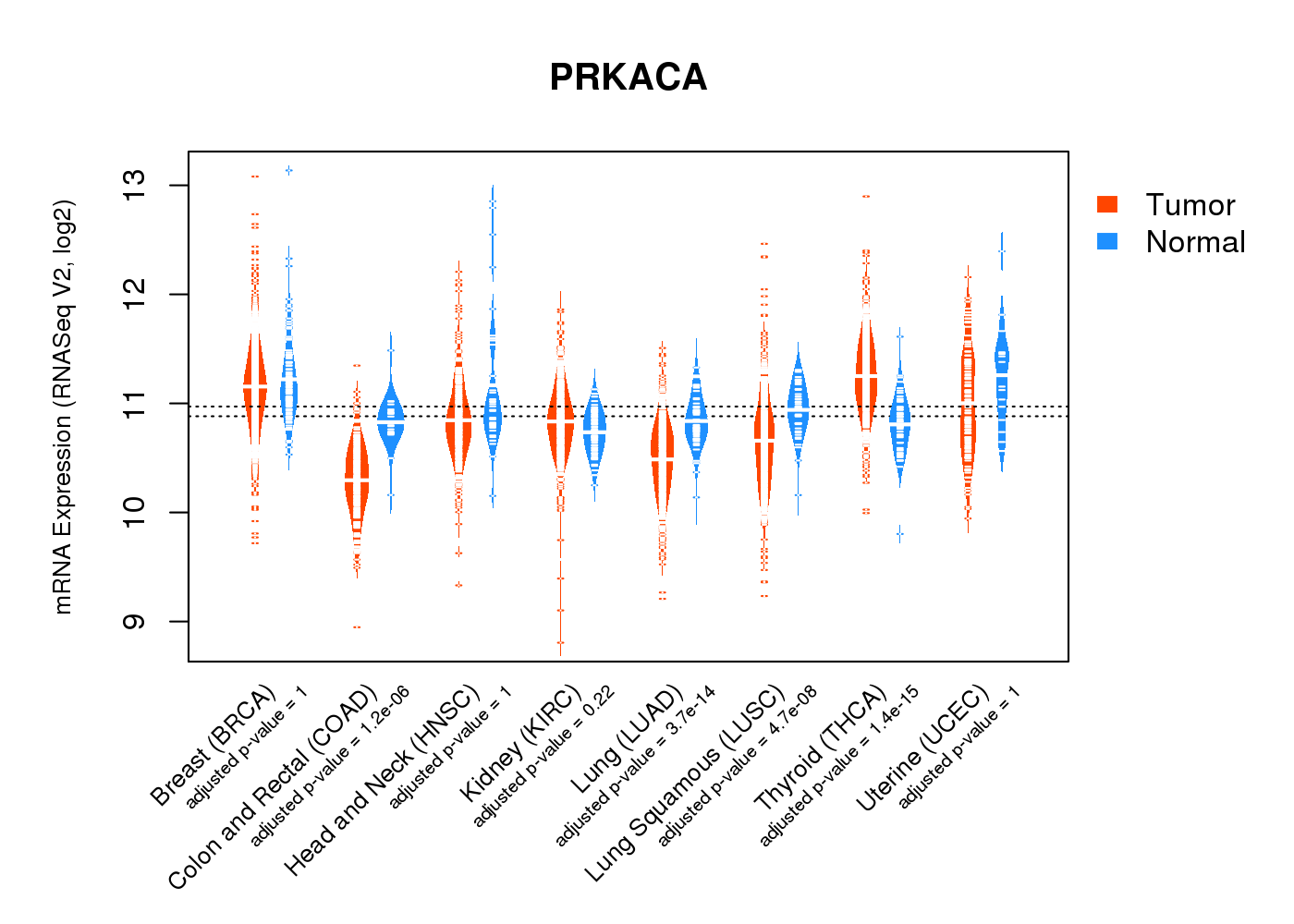 |
| Top |
| * This plots show the correlation between CNV and gene expression. |
: Open all plots for all cancer types
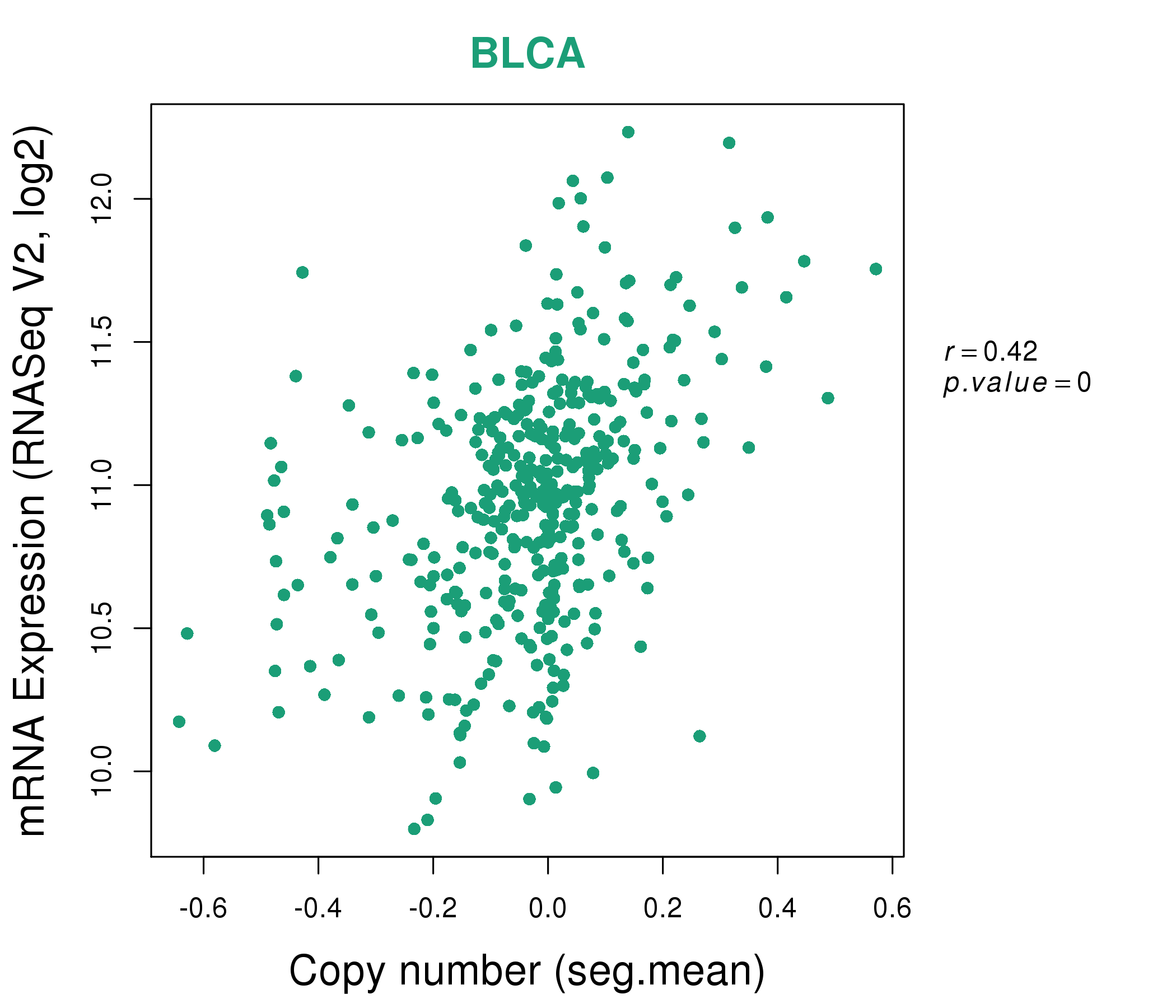 |
|
 |
|
| Top |
| Gene-Gene Network Information |
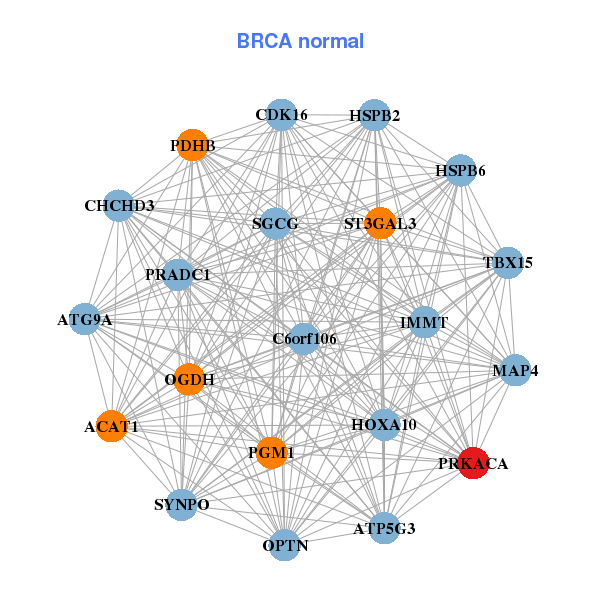
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
 |
| ||||
| AP1M1,ASF1B,BRD4,C19orf44,C19orf53,CC2D1A,CHERP, DCAF15,DDA1,DDX39A,FAM32A,LPHN1,MED26,NACC1, PRKACA,RAB8A,RAD23A,RAVER1,TBX22,TECR,WIZ | ACAT1,ATG9A,ATP5G3,PRADC1,C6orf106,CDK16,CHCHD3, HOXA10,HSPB2,HSPB6,IMMT,MAP4,OGDH,OPTN, PDHB,PGM1,PRKACA,SGCG,ST3GAL3,SYNPO,TBX15 | ||||
 |
| ||||
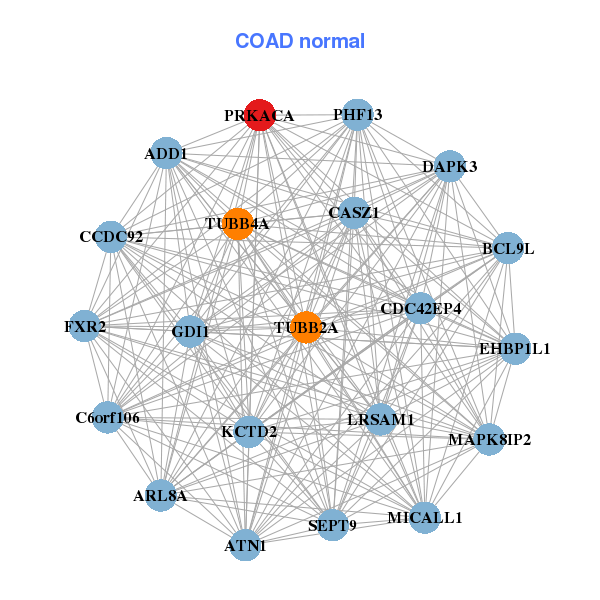
| ABHD8,AP1M1,ATN1,DVL3,EHD2,EPS15L1,FAM127C, MARCH2,MARK4,MLLT1,NFIX,PACS1,PALM2-AKAP2,PIP5K1C, PRAF2,PRKACA,INAFM1,TCEAL6,TCF7L1,WIZ,ZNF358 | ADD1,ARL8A,ATN1,BCL9L,C6orf106,CASZ1,CCDC92, CDC42EP4,DAPK3,EHBP1L1,FXR2,GDI1,KCTD2,LRSAM1, MAPK8IP2,MICALL1,PHF13,PRKACA,SEPT9,TUBB2A,TUBB4A |
| * Co-Expression network figures were drawn using R package igraph. Only the top 20 genes with the highest correlations were shown. Red circle: input gene, orange circle: cell metabolism gene, sky circle: other gene |
: Open all plots for all cancer types
| Top |
: Open all interacting genes' information including KEGG pathway for all interacting genes from DAVID
| Top |
| Pharmacological Information for PRKACA |
| DB Category | DB Name | DB's ID and Url link |
| Chemistry | BindingDB | P17612; -. |
| Chemistry | ChEMBL | CHEMBL2094138; -. |
| Chemistry | GuidetoPHARMACOLOGY | 1476; -. |
| Organism-specific databases | PharmGKB | PA33748; -. |
| Organism-specific databases | CTD | 5566; -. |
| * Gene Centered Interaction Network. |
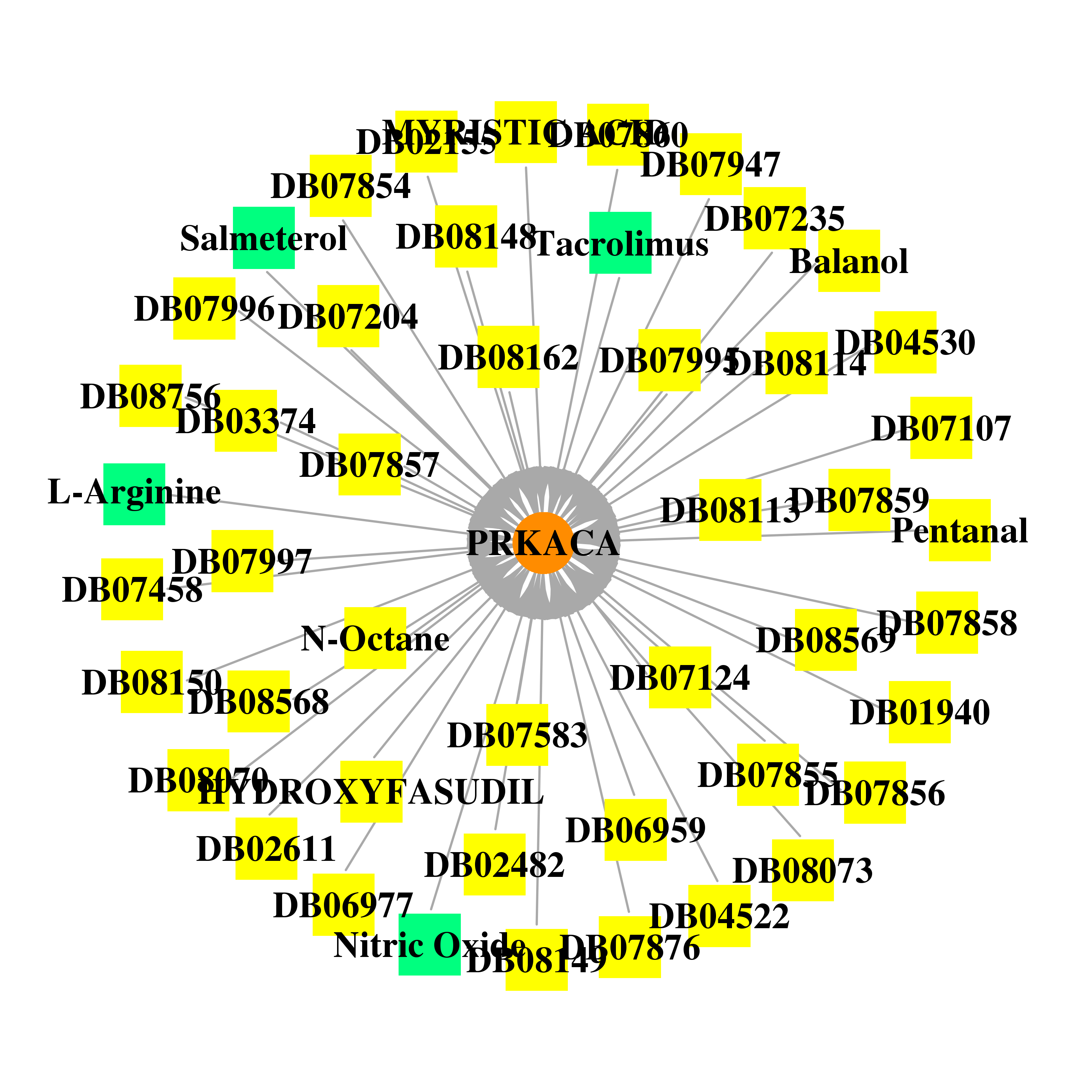 |
| * Drug Centered Interaction Network. |
| DrugBank ID | Target Name | Drug Groups | Generic Name | Drug Centered Network | Drug Structure |
| DB01919 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Pentanal |  |  |
| DB01940 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Balanol Analog 2 |  | 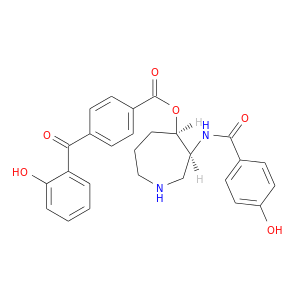 |
| DB02155 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Balanol Analog 8 |  |  |
| DB02440 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | N-Octane |  | 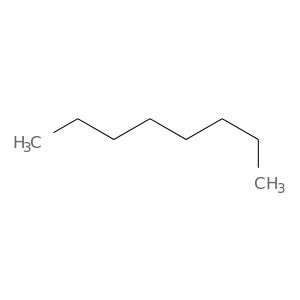 |
| DB02482 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Phosphonothreonine |  |  |
| DB02611 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Balanol Analog 1 |  |  |
| DB03374 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 3,5-Diiodotyrosine |  | 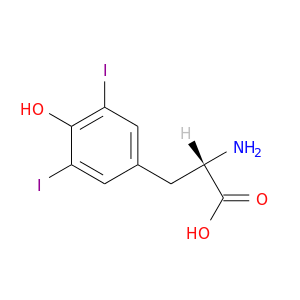 |
| DB04098 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Balanol |  |  |
| DB04522 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | Phosphonoserine |  |  |
| DB04530 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | S,S-(2-Hydroxyethyl)Thiocysteine | 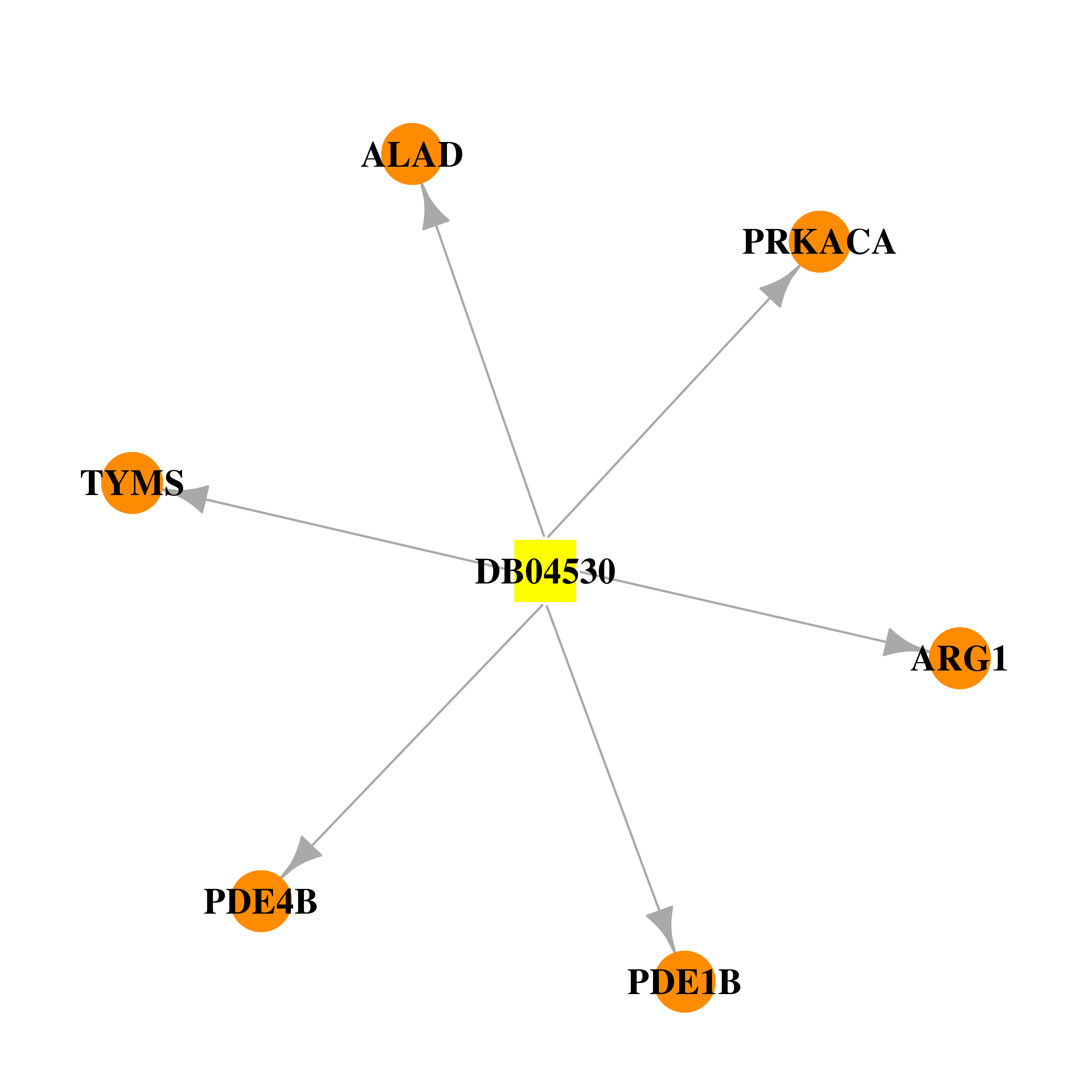 | 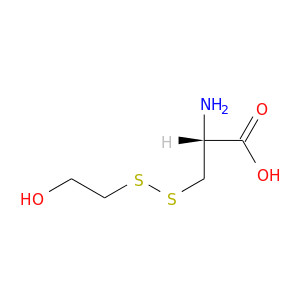 |
| DB04707 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | HYDROXYFASUDIL | 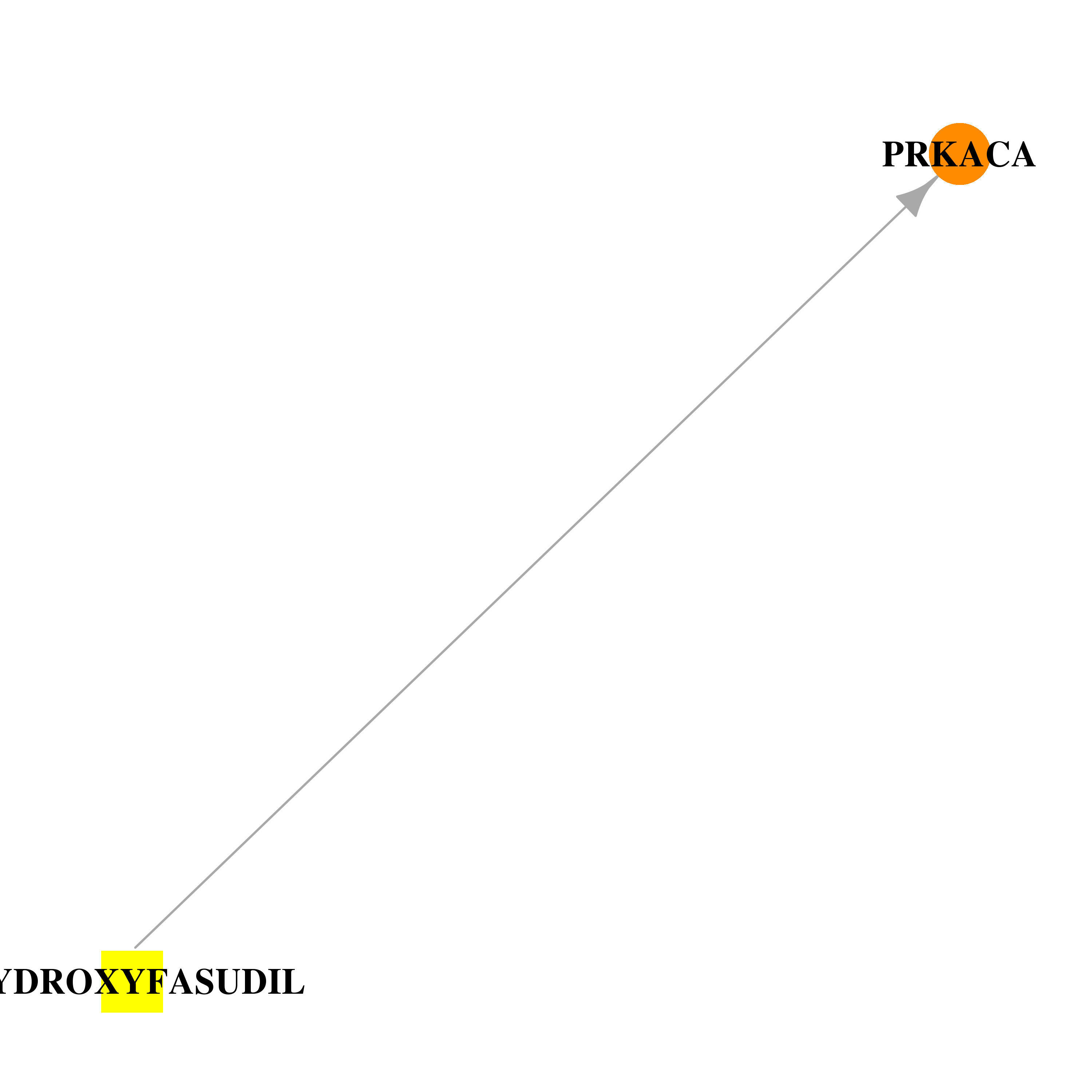 | 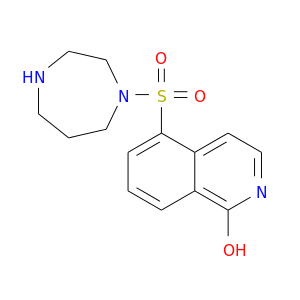 |
| DB06959 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (1S)-2-(1H-INDOL-3-YL)-1-{[(5-ISOQUINOLIN-6-YLPYRIDIN-3-YL)OXY]METHYL}ETHYLAMINE |  |  |
| DB06977 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2S)-1-{[5-(1H-INDAZOL-5-YL)PYRIDIN-3-YL]OXY}-3-[(7AS)-7AH-INDOL-3-YL]PROPAN-2-AMINE |  | 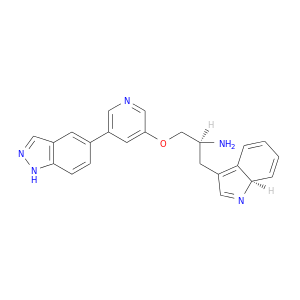 |
| DB07107 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (1S)-2-(1H-INDOL-3-YL)-1-[({5-[(E)-2-PYRIDIN-4-YLVINYL]PYRIDIN-3-YL}OXY)METHYL]ETHYLAMINE |  |  |
| DB07124 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2S)-1-(6H-INDOL-3-YL)-3-{[5-(7H-PYRAZOLO[3,4-C]PYRIDIN-5-YL)PYRIDIN-3-YL]OXY}PROPAN-2-AMINE | 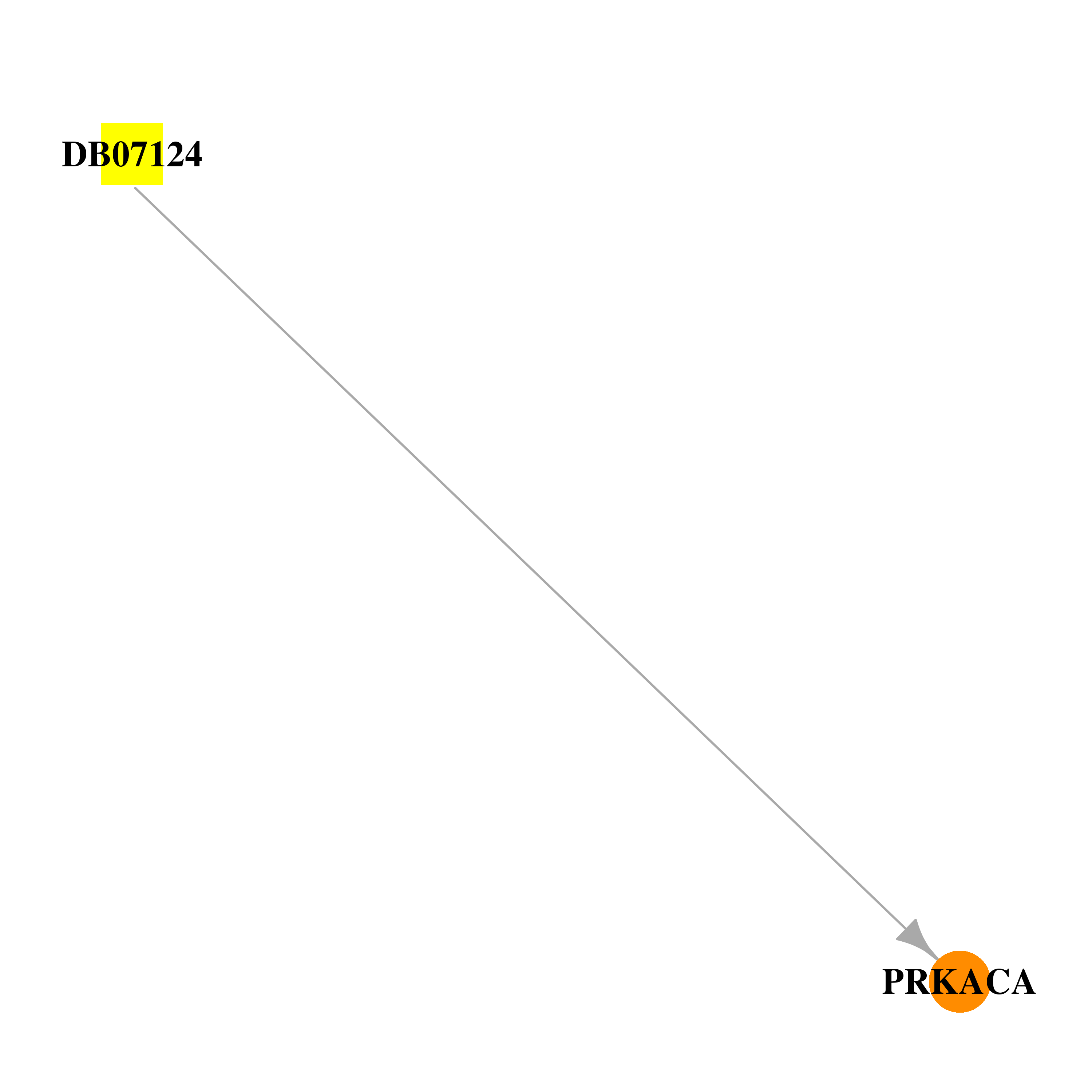 |  |
| DB07204 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (1S)-1-(1H-INDOL-3-YLMETHYL)-2-(2-PYRIDIN-4-YL-[1,7]NAPHTYRIDIN-5-YLOXY)-EHYLAMINE |  |  |
| DB07235 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | N-[(1S)-2-AMINO-1-(2,4-DICHLOROBENZYL)ETHYL]-5-[2-(METHYLAMINO)PYRIMIDIN-4-YL]THIOPHENE-2-CARBOXAMIDE |  | 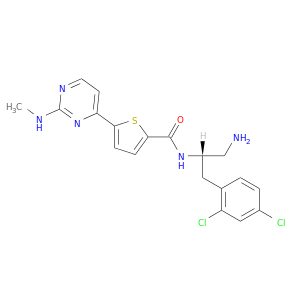 |
| DB07458 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 3-(1H-INDOL-3-YL)-4-{1-[2-(1-METHYLPYRROLIDIN-2-YL)ETHYL]-1H-INDOL-3-YL}-1H-PYRROLE-2,5-DIONE |  | 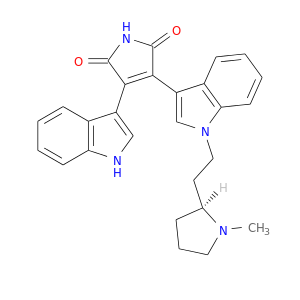 |
| DB07583 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (4R,2S)-5'-(4-(4-CHLOROBENZYLOXY)PYRROLIDIN-2-YLMETHANESULFONYL)ISOQUINOLINE |  | 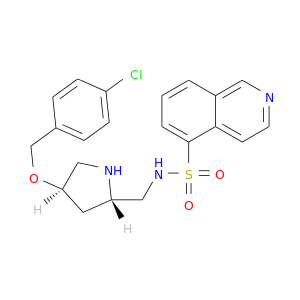 |
| DB07854 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | N-METHYL-1-[4-(9H-PURIN-6-YL)PHENYL]METHANAMINE |  | 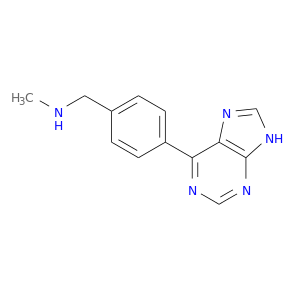 |
| DB07855 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (S)-1-PHENYL-1-[4-(9H-PURIN-6-YL)PHENYL]METHANAMINE |  | 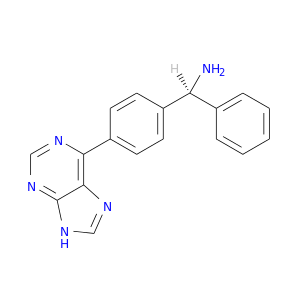 |
| DB07856 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 6-{4-[4-(4-CHLOROPHENYL)PIPERIDIN-4-YL]PHENYL}-9H-PURINE |  | 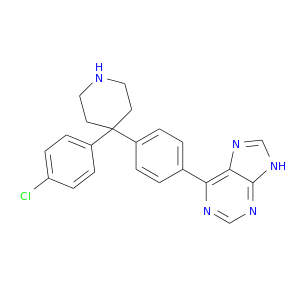 |
| DB07857 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2R)-2-(4-CHLOROPHENYL)-2-[4-(1H-PYRAZOL-4-YL)PHENYL]ETHANAMINE |  | 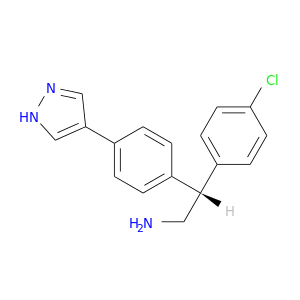 |
| DB07858 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2S)-2-(4-CHLOROPHENYL)-2-[4-(1H-PYRAZOL-4-YL)PHENYL]ETHANAMINE |  |  |
| DB07859 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 4-(4-CHLOROPHENYL)-4-[4-(1H-PYRAZOL-4-YL)PHENYL]PIPERIDINE | 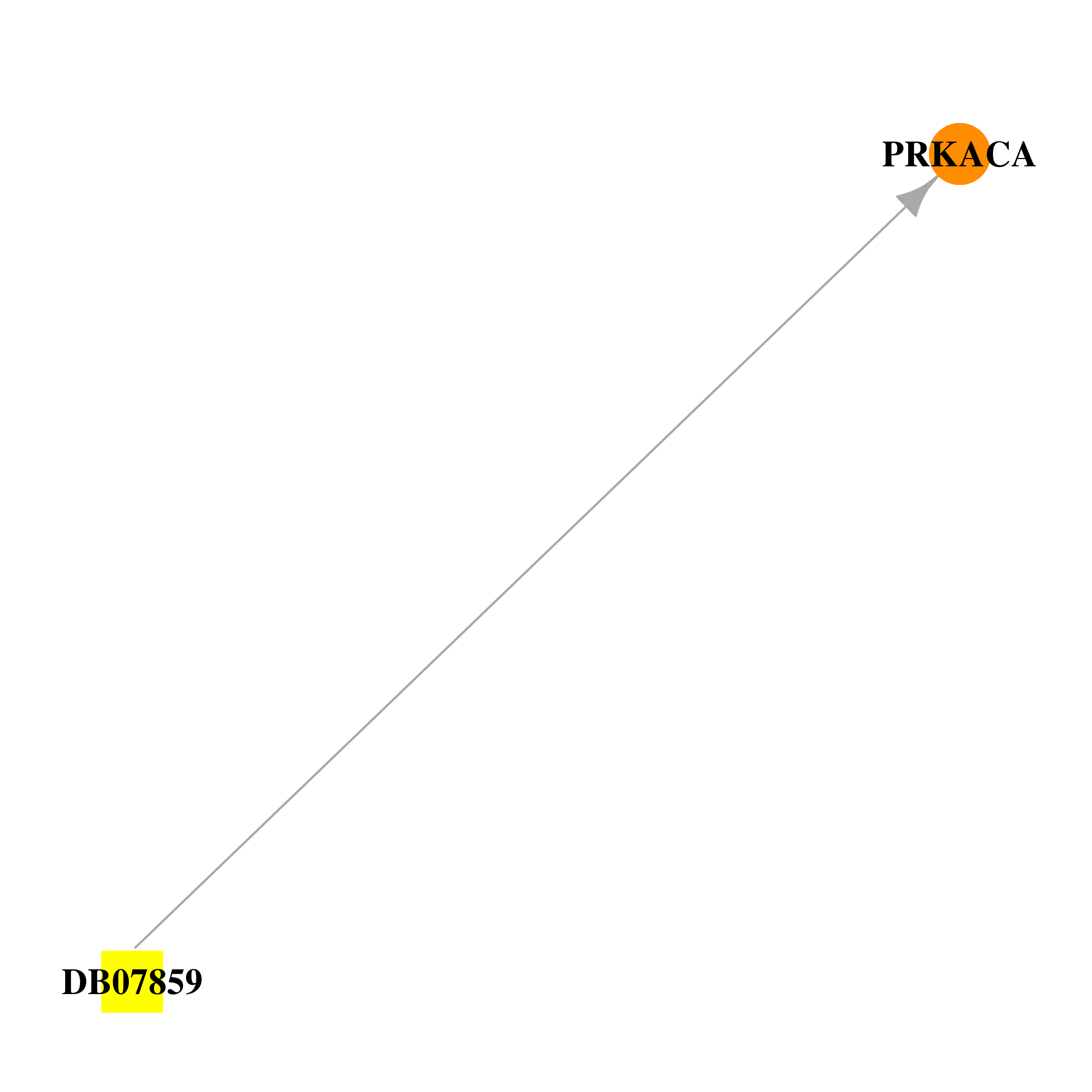 | 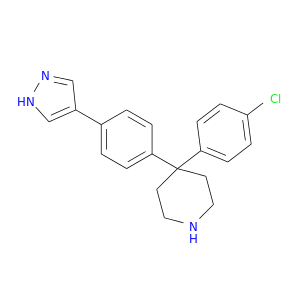 |
| DB07860 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2R)-2-(4-CHLOROPHENYL)-2-PHENYLETHANAMINE |  |  |
| DB07876 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (S)-2-METHYL-1-[(4-METHYL-5-ISOQUINOLINE)SULFONYL]-HOMOPIPERAZINE |  | 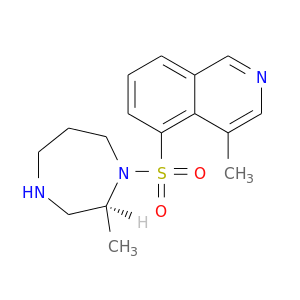 |
| DB07947 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | ISOQUINOLINE-5-SULFONIC ACID (2-(2-(4-CHLOROBENZYLOXY)ETHYLAMINO)ETHYL)AMIDE |  |  |
| DB07995 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | N-[2-(4-BROMOCINNAMYLAMINO)ETHYL]-5-ISOQUINOLINE SULFONAMIDE |  | 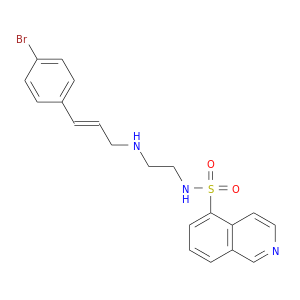 |
| DB07996 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 1-(5-ISOQUINOLINESULFONYL)-2-METHYLPIPERAZINE |  |  |
| DB07997 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | N-[2-(METHYLAMINO)ETHYL]-5-ISOQUINOLINESULFONAMIDE |  | 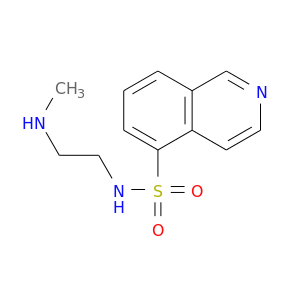 |
| DB08070 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 2-[4-(3-METHYL-1H-PYRAZOL-4-YL)PHENYL]ETHANAMINE |  |  |
| DB08073 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2S)-1-(1H-INDOL-3-YL)-3-{[5-(3-METHYL-1H-INDAZOL-5-YL)PYRIDIN-3-YL]OXY}PROPAN-2-AMINE |  |  |
| DB08113 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 3-pyridin-4-yl-1H-indazole |  |  |
| DB08114 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 5-benzyl-1,3-thiazol-2-amine |  |  |
| DB08148 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 1-[4-(4-chlorophenyl)-1-(7H-pyrrolo[2,3-d]pyrimidin-4-yl)piperidin-4-yl]methanamine |  |  |
| DB08149 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 1-[4-(4-chlorobenzyl)-1-(7H-pyrrolo[2,3-d]pyrimidin-4-yl)piperidin-4-yl]methanamine |  |  |
| DB08150 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 4-(4-chlorobenzyl)-1-(7H-pyrrolo[2,3-d]pyrimidin-4-yl)piperidin-4-aminium |  |  |
| DB08162 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 5-(1,4-DIAZEPAN-1-SULFONYL)ISOQUINOLINE |  |  |
| DB08231 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | MYRISTIC ACID |  | 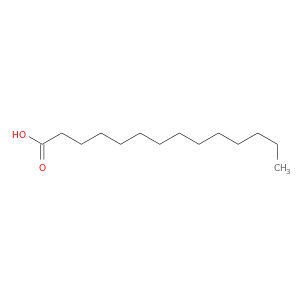 |
| DB08568 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (2S)-1-{[5-(3-METHYL-1H-INDAZOL-5-YL)PYRIDIN-3-YL]OXY}-3-PHENYLPROPAN-2-AMINE |  | 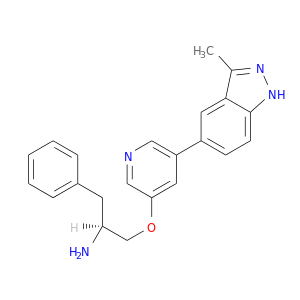 |
| DB08569 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | 3-PYRIDIN-4-YL-2,4-DIHYDRO-INDENO[1,2-.C.] PYRAZOLE |  | 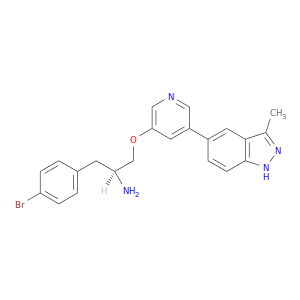 |
| DB08756 | protein kinase, cAMP-dependent, catalytic, alpha | experimental | (R)-TRANS-4-(1-AMINOETHYL)-N-(4-PYRIDYL) CYCLOHEXANECARBOXAMIDE |  | 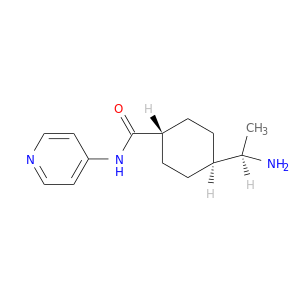 |
| DB00938 | protein kinase, cAMP-dependent, catalytic, alpha | approved | Salmeterol | 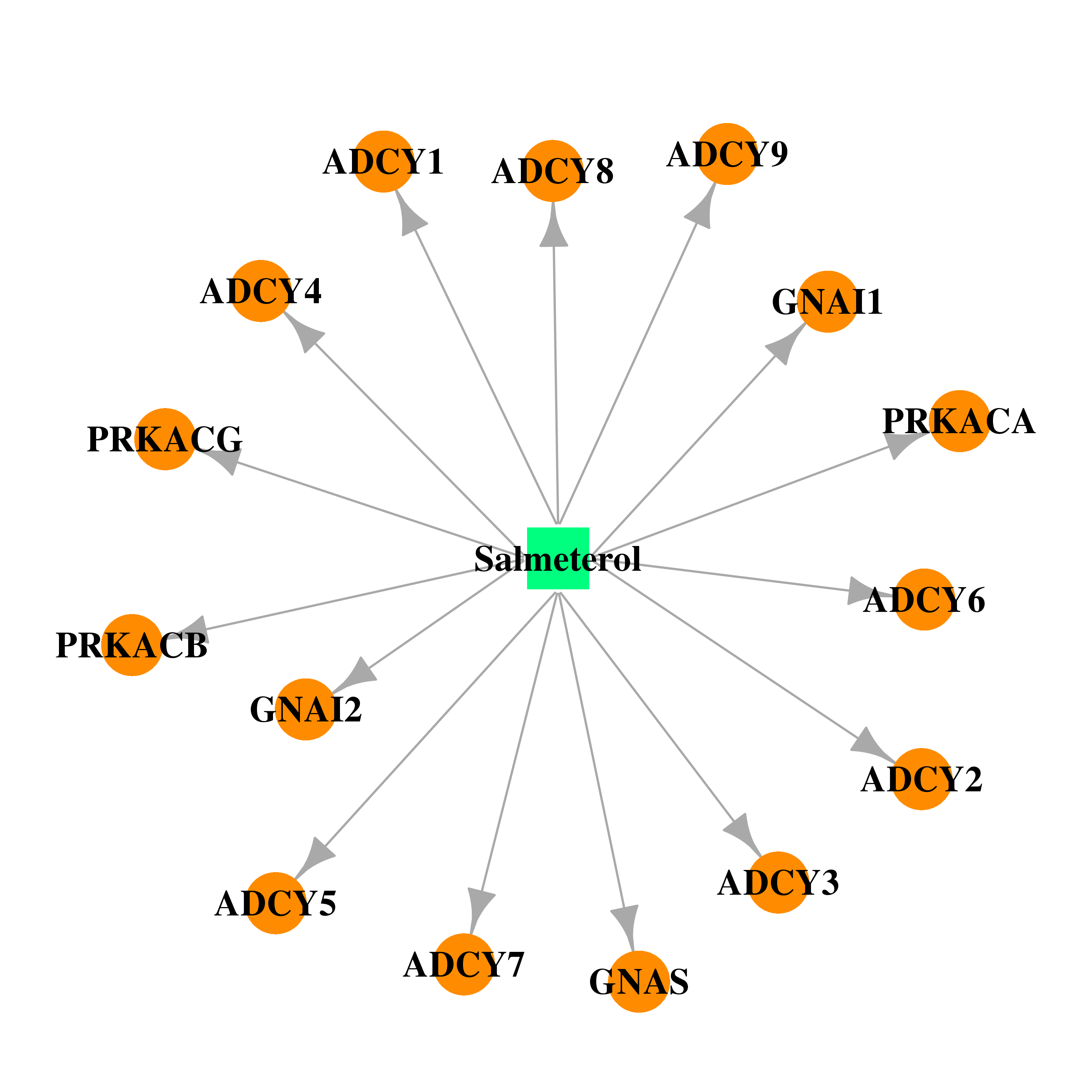 | 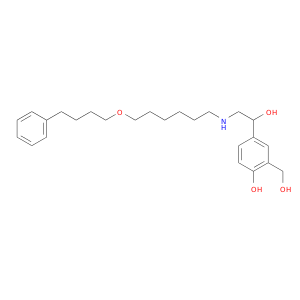 |
| DB00125 | protein kinase, cAMP-dependent, catalytic, alpha | approved; nutraceutical | L-Arginine | 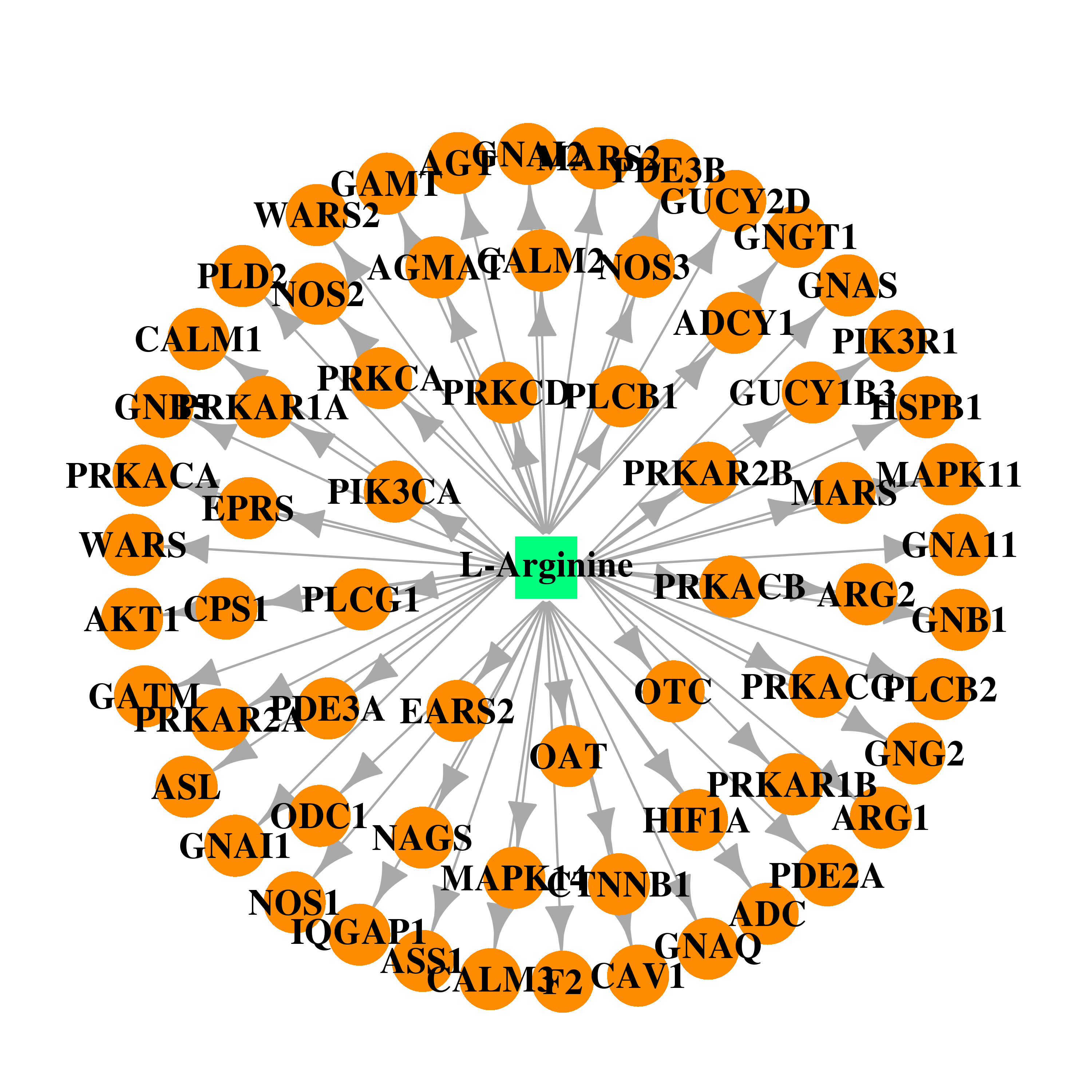 | 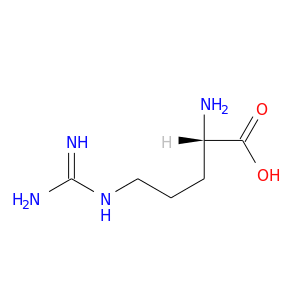 |
| DB00435 | protein kinase, cAMP-dependent, catalytic, alpha | approved | Nitric Oxide | 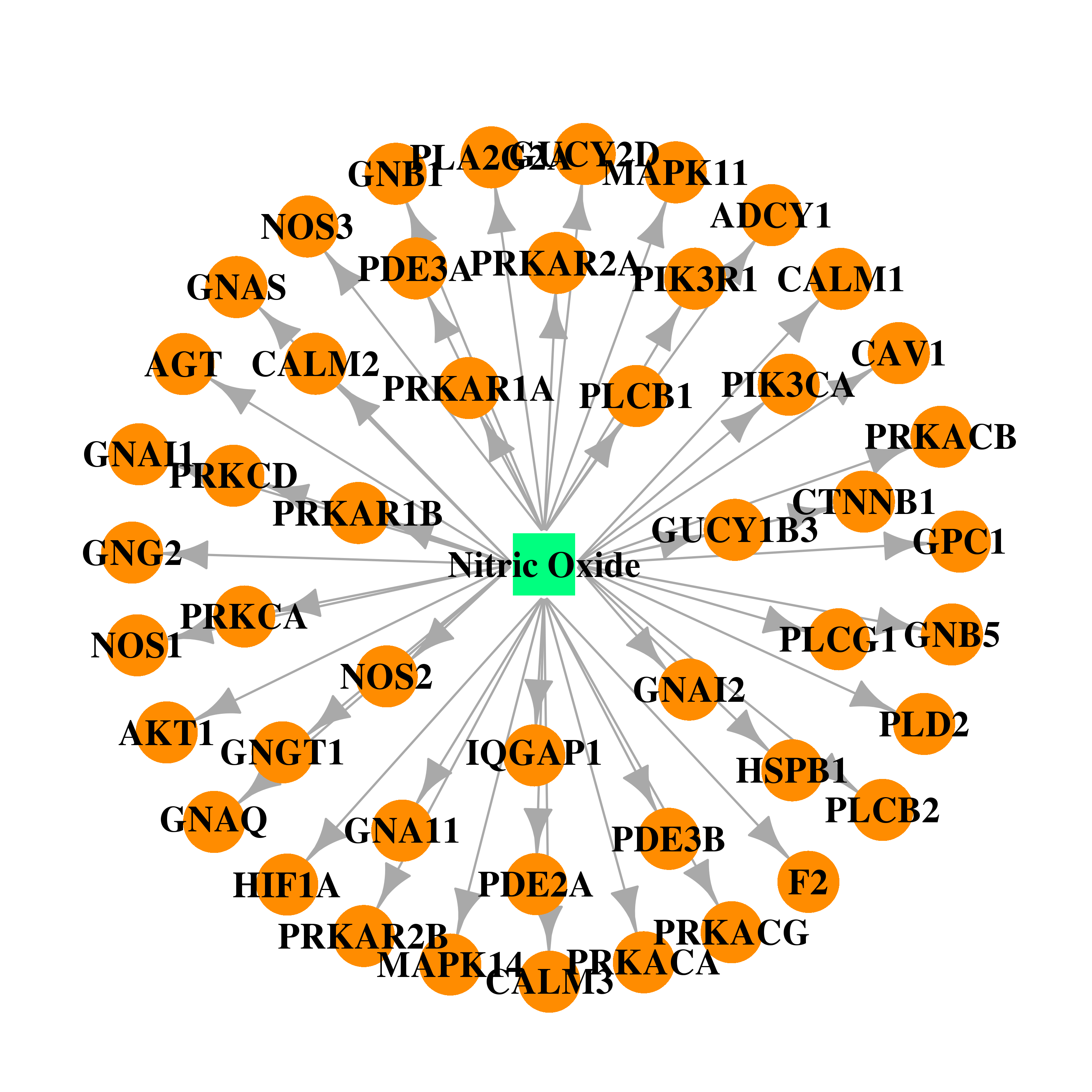 |  |
| DB00864 | protein kinase, cAMP-dependent, catalytic, alpha | approved; investigational | Tacrolimus |  |  |
| Top |
| Cross referenced IDs for PRKACA |
| * We obtained these cross-references from Uniprot database. It covers 150 different DBs, 18 categories. http://www.uniprot.org/help/cross_references_section |
: Open all cross reference information
|
Copyright © 2016-Present - The Univsersity of Texas Health Science Center at Houston @ |