| LBS | AA sequence | # species | Species |
A249 | GPISVAMDASH | 2 | Mus musculus, Rattus norvegicus | A249 | GPISVAIDAGH | 2 | Canis lupus familiaris, Bos taurus | A249 | GPISVAMDAGH | 1 | Homo sapiens | A249 | QPISVMISAAN | 1 | Arabidopsis thaliana | A249 | QPVSVAIEASS | 1 | Arabidopsis thaliana | A249 | QPISIAIEAGG | 1 | Arabidopsis thaliana | A249 | QPVSVAIDAGG | 1 | Oryza sativa subsp. japonica | A249 | QPVSVAIEAGG | 1 | Oryza sativa subsp. japonica | A249 | GPVSVAIDASH | 1 | Drosophila melanogaster | C138 | GQCGSCWAFSA | 6 | Oryza sativa subsp. japonica, Oryza sativa subsp. japonica, Mus musculus, Rattus norvegicus, Canis lupus familiaris, Bos taurus | C138 | KQCGSCWAFSA | 1 | Homo sapiens | C138 | GECGSCWAFAA | 1 | Arabidopsis thaliana | C138 | GNCGSCWAFSA | 1 | Arabidopsis thaliana | C138 | GGCGSCWAFST | 1 | Arabidopsis thaliana | C138 | GHCGSCWAFSS | 1 | Drosophila melanogaster | C178 | QGNQGCNGGLM | 3 | Mus musculus, Rattus norvegicus, Bos taurus | C178 | QGNQGCNGGFM | 1 | Homo sapiens | C178 | NDNFGCAGGGA | 1 | Arabidopsis thaliana | C178 | FVNAGCDGGIM | 1 | Arabidopsis thaliana | C178 | Y-NEGCNGGLM | 1 | Arabidopsis thaliana | C178 | GQNSGCNGGIM | 1 | Oryza sativa subsp. japonica | C178 | GQNSGCNGGLM | 1 | Oryza sativa subsp. japonica | C178 | YGNNGCNGGLM | 1 | Drosophila melanogaster | C178 | QGNEGCNGGLM | 1 | Canis lupus familiaris | D276 | SSKDLDHGVLV | 3 | Rattus norvegicus, Canis lupus familiaris, Bos taurus | D276 | SSKNLDHGVLV | 1 | Homo sapiens | D276 | SNLWGDHNVLI | 1 | Arabidopsis thaliana | D276 | GI-SLDHGVVV | 1 | Arabidopsis thaliana | D276 | GT-QLDHGVVA | 1 | Arabidopsis thaliana | D276 | GT-NLDHGVVA | 1 | Oryza sativa subsp. japonica | D276 | GT-SLDHGVVA | 1 | Oryza sativa subsp. japonica | D276 | DAQNLDHGVLV | 1 | Drosophila melanogaster | D276 | SSKNLDHGVLL | 1 | Mus musculus | F182 | GCNGGLMDNAF | 3 | Drosophila melanogaster, Canis lupus familiaris, Bos taurus | F182 | GCNGGLMDFAF | 2 | Mus musculus, Rattus norvegicus | F182 | GCNGGFMARAF | 1 | Homo sapiens | F182 | GCAGGGAVWAF | 1 | Arabidopsis thaliana | F182 | GCDGGIMNYAF | 1 | Arabidopsis thaliana | F182 | GCNGGLMDYAF | 1 | Arabidopsis thaliana | F182 | GCNGGIMDDAF | 1 | Oryza sativa subsp. japonica | F182 | GCNGGLMDDAF | 1 | Oryza sativa subsp. japonica | G136 | NQGQCGSCWAF | 6 | Oryza sativa subsp. japonica, Oryza sativa subsp. japonica, Mus musculus, Rattus norvegicus, Canis lupus familiaris, Bos taurus | G136 | NQKQCGSCWAF | 1 | Homo sapiens | G136 | RQGECGSCWAF | 1 | Arabidopsis thaliana | G136 | DQGNCGSCWAF | 1 | Arabidopsis thaliana | G136 | DQGGCGSCWAF | 1 | Arabidopsis thaliana | G136 | DQGHCGSCWAF | 1 | Drosophila melanogaster | G180 | NQGCNGGLMDF | 2 | Mus musculus, Rattus norvegicus | G180 | NQGCNGGFMAR | 1 | Homo sapiens | G180 | NFGCAGGGAVW | 1 | Arabidopsis thaliana | G180 | NAGCDGGIMNY | 1 | Arabidopsis thaliana | G180 | NEGCNGGLMDY | 1 | Arabidopsis thaliana | G180 | NSGCNGGIMDD | 1 | Oryza sativa subsp. japonica | G180 | NSGCNGGLMDD | 1 | Oryza sativa subsp. japonica | G180 | NNGCNGGLMDN | 1 | Drosophila melanogaster | G180 | NEGCNGGLMDN | 1 | Canis lupus familiaris | G180 | NQGCNGGLMDN | 1 | Bos taurus | G181 | QGCNGGLMDFA | 2 | Mus musculus, Rattus norvegicus | G181 | QGCNGGFMARA | 1 | Homo sapiens | G181 | FGCAGGGAVWA | 1 | Arabidopsis thaliana | G181 | AGCDGGIMNYA | 1 | Arabidopsis thaliana | G181 | EGCNGGLMDYA | 1 | Arabidopsis thaliana | G181 | SGCNGGIMDDA | 1 | Oryza sativa subsp. japonica | G181 | SGCNGGLMDDA | 1 | Oryza sativa subsp. japonica | G181 | NGCNGGLMDNA | 1 | Drosophila melanogaster | G181 | EGCNGGLMDNA | 1 | Canis lupus familiaris | G181 | QGCNGGLMDNA | 1 | Bos taurus | H277 | SKDLDHGVLVV | 3 | Rattus norvegicus, Canis lupus familiaris, Bos taurus | H277 | SKNLDHGVLVV | 1 | Homo sapiens | H277 | NLWGDHNVLIV | 1 | Arabidopsis thaliana | H277 | I-SLDHGVVVV | 1 | Arabidopsis thaliana | H277 | T-QLDHGVVAV | 1 | Arabidopsis thaliana | H277 | T-NLDHGVVAV | 1 | Oryza sativa subsp. japonica | H277 | T-SLDHGVVAV | 1 | Oryza sativa subsp. japonica | H277 | AQNLDHGVLVV | 1 | Drosophila melanogaster | H277 | SKNLDHGVLLV | 1 | Mus musculus | L275 | CSSKDLDHGVL | 3 | Rattus norvegicus, Canis lupus familiaris, Bos taurus | L275 | CSSKNLDHGVL | 2 | Homo sapiens, Mus musculus | L275 | CSNLWGDHNVL | 1 | Arabidopsis thaliana | L275 | CGI-SLDHGVV | 1 | Arabidopsis thaliana | L275 | CGT-QLDHGVV | 1 | Arabidopsis thaliana | L275 | CGT-NLDHGVV | 1 | Oryza sativa subsp. japonica | L275 | CGT-SLDHGVV | 1 | Oryza sativa subsp. japonica | L275 | CDAQNLDHGVL | 1 | Drosophila melanogaster | N179 | GNQGCNGGLMD | 3 | Mus musculus, Rattus norvegicus, Bos taurus | N179 | GNQGCNGGFMA | 1 | Homo sapiens | N179 | DNFGCAGGGAV | 1 | Arabidopsis thaliana | N179 | VNAGCDGGIMN | 1 | Arabidopsis thaliana | N179 | -NEGCNGGLMD | 1 | Arabidopsis thaliana | N179 | QNSGCNGGIMD | 1 | Oryza sativa subsp. japonica | N179 | QNSGCNGGLMD | 1 | Oryza sativa subsp. japonica | N179 | GNNGCNGGLMD | 1 | Drosophila melanogaster | N179 | GNEGCNGGLMD | 1 | Canis lupus familiaris | Q132 | TPVKNQGQCGS | 4 | Mus musculus, Rattus norvegicus, Canis lupus familiaris, Bos taurus | Q132 | APVKNQGQCGS | 2 | Oryza sativa subsp. japonica, Oryza sativa subsp. japonica | Q132 | TPVKNQKQCGS | 1 | Homo sapiens | Q132 | PRVKRQGECGS | 1 | Arabidopsis thaliana | Q132 | VSVKDQGNCGS | 1 | Arabidopsis thaliana | Q132 | AEVKDQGGCGS | 1 | Arabidopsis thaliana | Q132 | TAVKDQGHCGS | 1 | Drosophila melanogaster | Q176 | RPQGNQGCNGG | 1 | Homo sapiens | Q176 | RGNDNFGCAGG | 1 | Arabidopsis thaliana | Q176 | RGFVNAGCDGG | 1 | Arabidopsis thaliana | Q176 | TSY-NEGCNGG | 1 | Arabidopsis thaliana | Q176 | RNGQNSGCNGG | 1 | Oryza sativa subsp. japonica | Q176 | TNGQNSGCNGG | 1 | Oryza sativa subsp. japonica | Q176 | TKYGNNGCNGG | 1 | Drosophila melanogaster | Q176 | HAQGNQGCNGG | 1 | Mus musculus | Q176 | HDQGNQGCNGG | 1 | Rattus norvegicus | Q176 | RAQGNEGCNGG | 1 | Canis lupus familiaris | Q176 | RAQGNQGCNGG | 1 | Bos taurus | W139 | QCGSCWAFSAT | 3 | Homo sapiens, Canis lupus familiaris, Bos taurus | W139 | QCGSCWAFSAV | 2 | Oryza sativa subsp. japonica, Oryza sativa subsp. japonica | W139 | QCGSCWAFSAS | 2 | Mus musculus, Rattus norvegicus | W139 | ECGSCWAFAAT | 1 | Arabidopsis thaliana | W139 | NCGSCWAFSAV | 1 | Arabidopsis thaliana | W139 | GCGSCWAFSTI | 1 | Arabidopsis thaliana | W139 | HCGSCWAFSST | 1 | Drosophila melanogaster | W303 | IVKNSWGPEWG | 2 | Canis lupus familiaris, Bos taurus | W303 | LVKNSWGPEWG | 1 | Homo sapiens | W303 | LIRNSWGPEWG | 1 | Arabidopsis thaliana | W303 | IIRNSWGLNWG | 1 | Arabidopsis thaliana | W303 | IVRNSWGKSWG | 1 | Arabidopsis thaliana | W303 | TVRNSWGPDWG | 1 | Oryza sativa subsp. japonica | W303 | IVRNSWGPKWG | 1 | Oryza sativa subsp. japonica | W303 | LVKNSWGTTWG | 1 | Drosophila melanogaster | W303 | LVKNSWGSEWG | 1 | Mus musculus | W303 | LVKNSWGKEWG | 1 | Rattus norvegicus |
 Gene summary
Gene summary  Gene ontology having evidence of Inferred from Direct Assay (IDA) from Entrez
Gene ontology having evidence of Inferred from Direct Assay (IDA) from Entrez  Cancer type specific mutLBS sorted by frequency
Cancer type specific mutLBS sorted by frequency Relative protein structure stability change (ΔΔE) using Mupro 1.1
Relative protein structure stability change (ΔΔE) using Mupro 1.1  : nsSNV at non-LBS
: nsSNV at non-LBS : nsSNV at LBS
: nsSNV at LBS
 nsSNVs sorted by the relative stability change of protein structure by each mutation
nsSNVs sorted by the relative stability change of protein structure by each mutation  Structure image for CTSV from PDB
Structure image for CTSV from PDB Differential gene expression between mutated and non-mutated LBS samples in all 16 major cancer types
Differential gene expression between mutated and non-mutated LBS samples in all 16 major cancer types Differential co-expressed gene network based on protein-protein interaction data (CePIN)
Differential co-expressed gene network based on protein-protein interaction data (CePIN) Gene level disease information (DisGeNet)
Gene level disease information (DisGeNet)  Mutation level pathogenic information (ClinVar annotation)
Mutation level pathogenic information (ClinVar annotation)  Gene expression profile of anticancer drug treated cell-lines (CCLE)
Gene expression profile of anticancer drug treated cell-lines (CCLE)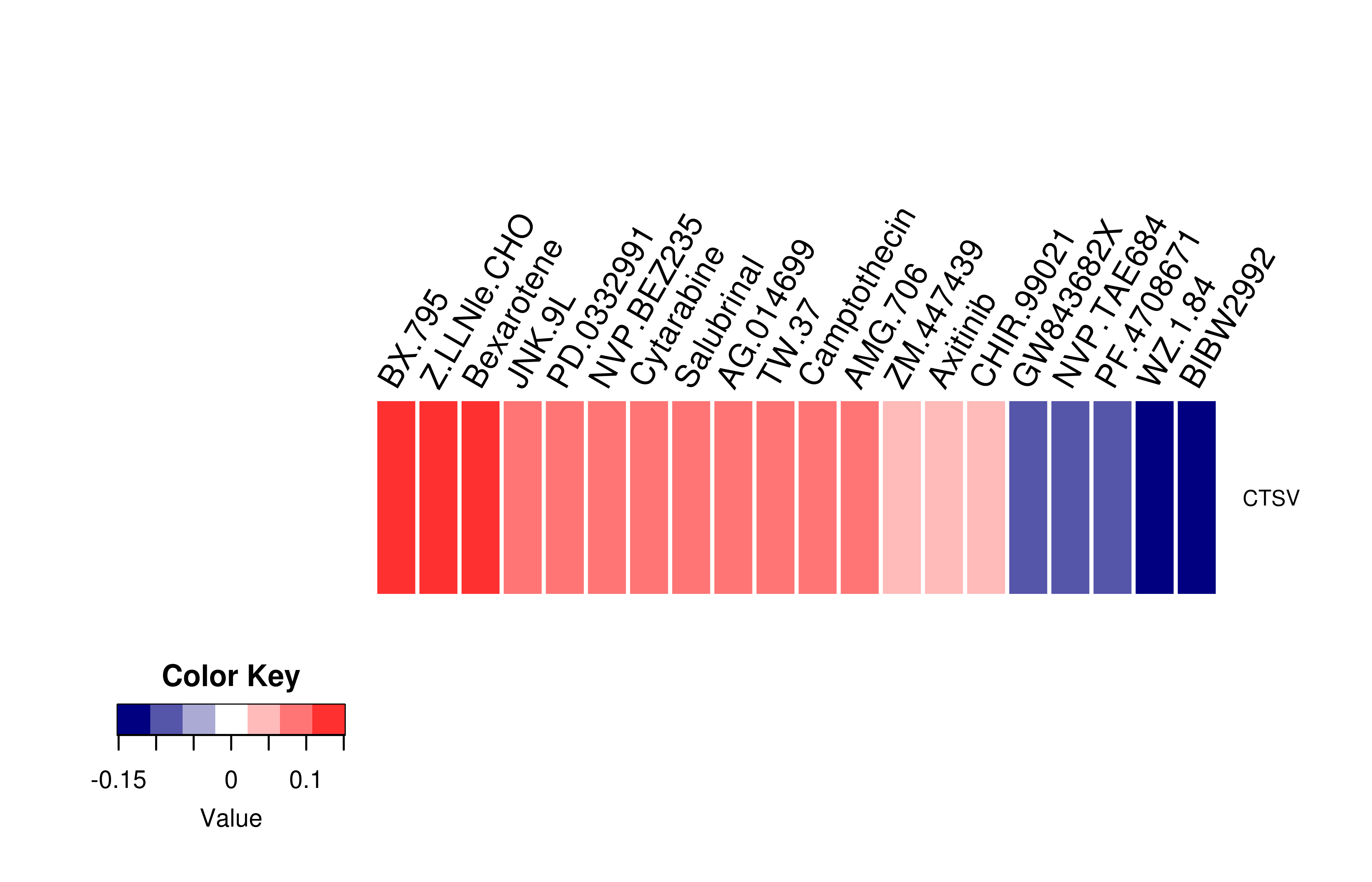
 Gene-centered drug-gene interaction network
Gene-centered drug-gene interaction network 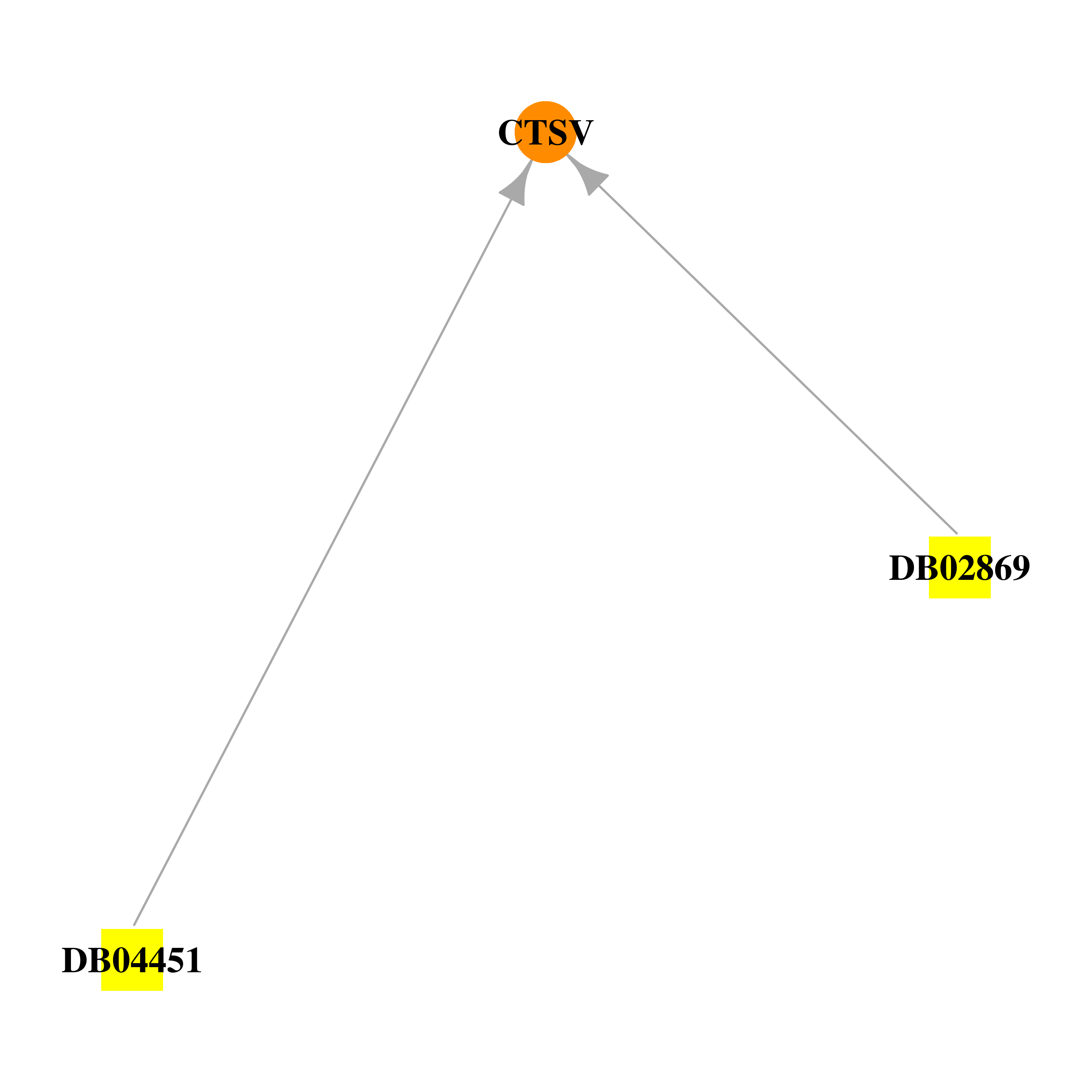
 Drug information targeting mutLBSgene (Approved drugs only)
Drug information targeting mutLBSgene (Approved drugs only)
 Gene-centered ligand-gene interaction network
Gene-centered ligand-gene interaction network 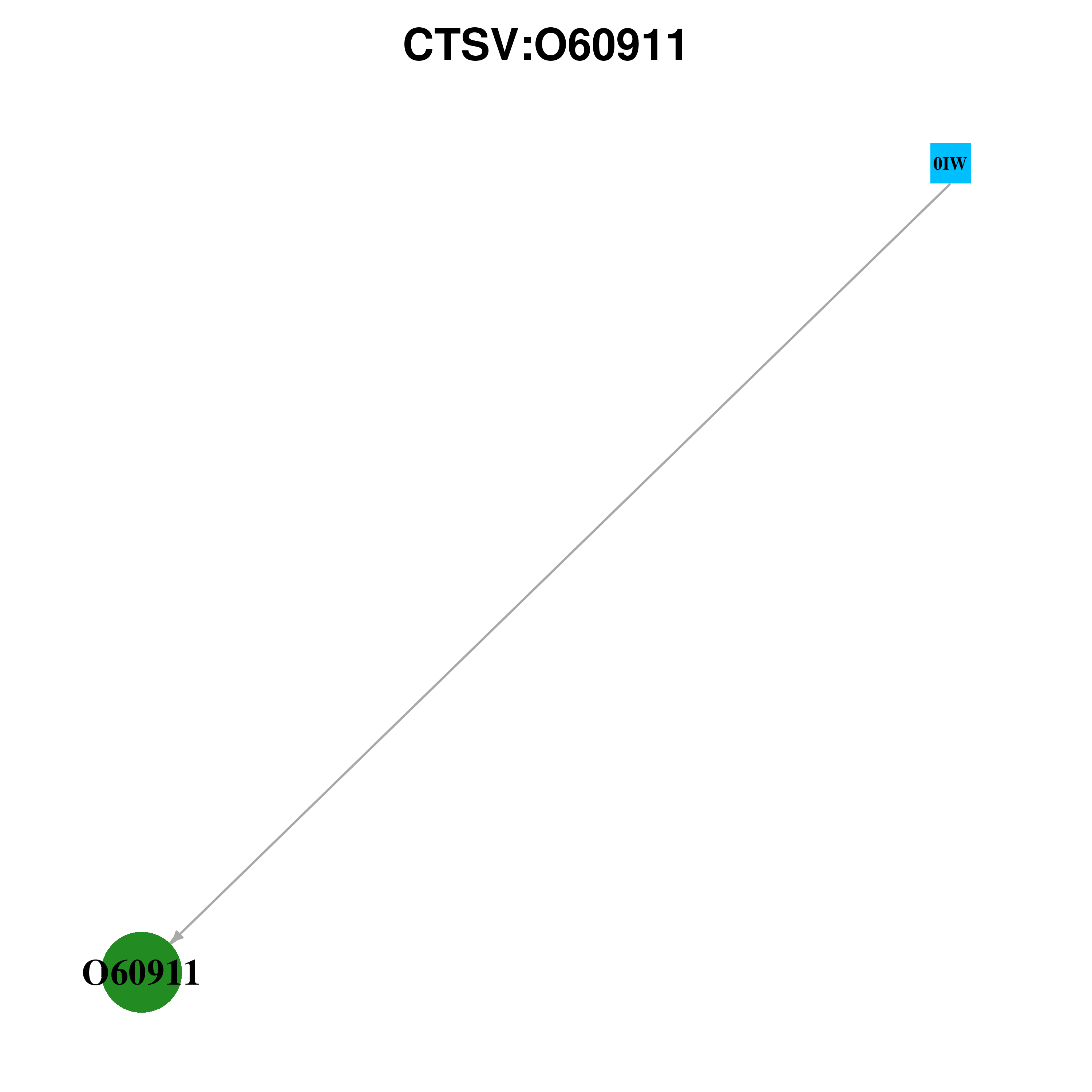
 Ligands binding to mutated ligand binding site of CTSV go to BioLip
Ligands binding to mutated ligand binding site of CTSV go to BioLip Multiple alignments for O60911 in multiple species
Multiple alignments for O60911 in multiple species 
