|
mutLBSgeneDB |
| |
| |
| |
| |
| |
| |
|
| Gene summary for FGFR2 |
 Gene summary Gene summary |
| Basic gene Info. | Gene symbol | FGFR2 |
| Gene name | fibroblast growth factor receptor 2 | |
| Synonyms | BBDS|BEK|BFR-1|CD332|CEK3|CFD1|ECT1|JWS|K-SAM|KGFR|TK14|TK25 | |
| Cytomap | UCSC genome browser: 10q26 | |
| Type of gene | protein-coding | |
| RefGenes | NM_000141.4, NM_001144913.1,NM_001144914.1,NM_001144915.1,NM_001144916.1, NM_001144917.1,NM_001144918.1,NM_001144919.1,NM_022970.3, NM_023029.2,NR_073009.1,NM_022971.1,NM_022972.1, NM_022973.1,NM_022974.1,NM_022975.2,NM_022976.1, NM_023028.1,NM_023030.1, | |
| Description | BEK fibroblast growth factor receptorFGF receptorFGFR-2FGFR2-AHCYL1 fusion kinase proteinbacteria-expressed kinasehydroxyaryl-protein kinasekeratinocyte growth factor receptorprotein tyrosine kinase, receptor like 14soluble FGFR4 variant 4 | |
| Modification date | 20141222 | |
| dbXrefs | MIM : 176943 | |
| HGNC : HGNC | ||
| Ensembl : ENSG00000066468 | ||
| HPRD : 01492 | ||
| Vega : OTTHUMG00000019175 | ||
| Protein | UniProt: P21802 go to UniProt's Cross Reference DB Table | |
| Expression | CleanEX: HS_FGFR2 | |
| BioGPS: 2263 | ||
| Pathway | NCI Pathway Interaction Database: FGFR2 | |
| KEGG: FGFR2 | ||
| REACTOME: FGFR2 | ||
| Pathway Commons: FGFR2 | ||
| Context | iHOP: FGFR2 | |
| ligand binding site mutation search in PubMed: FGFR2 | ||
| UCL Cancer Institute: FGFR2 | ||
| Assigned class in mutLBSgeneDB | A: This gene has a literature evidence and it belongs to targetable_mutLBSgenes. | |
| References showing study about ligand binding site mutation for FGFR2. | 1. "Wilkie AO, Oldridge M, Tang Z, Maxson RE Jr. Craniosynostosis and related limb anomalies. Novartis Found Symp. 2001;232:122-33; discussion 133-43. Review. PubMed PMID: 11277076. " 11277076 2. "Lew ED, Bae JH, Rohmann E, Wollnik B, Schlessinger J. Structural basis for reduced FGFR2 activity in LADD syndrome: Implications for FGFR autoinhibition and activation. Proc Natl Acad Sci U S A. 2007 Dec 11;104(50):19802-7. Epub 2007 Dec 3. PubMed PMID: 18056630; PubMed Central PMCID: PMC2148379. " 18056630 3. "Martin LA, Assif N, Gilbert M, Wijewarnasuriya D, Seandel M. Enhanced fitness of adult spermatogonial stem cells bearing a paternal age-associated FGFR2 mutation. Stem Cell Reports. 2014 Aug 12;3(2):219-26. doi:10.1016/j.stemcr.2014.06.007. Epub 2014 Jul 17. PubMed PMID: 25254335; PubMed Central PMCID: PMC4176532. " 25254335 | |
 Gene ontology having evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology having evidence of Inferred from Direct Assay (IDA) from Entrez |
| GO ID | GO Term | PubMed ID | GO:0008284 | positive regulation of cell proliferation | 8663044 | GO:0008543 | fibroblast growth factor receptor signaling pathway | 15629145 | GO:0018108 | peptidyl-tyrosine phosphorylation | 15629145 | GO:0046777 | protein autophosphorylation | 15629145 |
| Top |
| Ligand binding site mutations for FGFR2 |
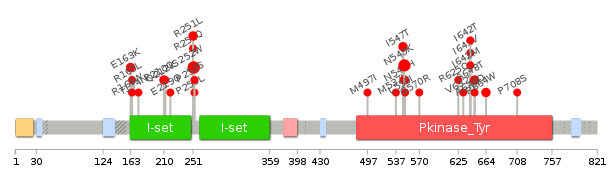
 Lollipop-style diagram of mutations at LBS in amino-acid sequence. Lollipop-style diagram of mutations at LBS in amino-acid sequence. We represented ligand binding site mutations only. (You can see big image via clicking.) |
 |
 Cancer type specific mutLBS sorted by frequency Cancer type specific mutLBS sorted by frequency |
| LBS | AAchange of nsSNV | Cancer type | # samples | R251 | S252W | UCEC | 9 | I548 | N549K | BRCA | 4 | I548 | N549K | UCEC | 4 | I548 | I547T | COAD | 2 | R251 | R251L | COAD | 2 | G646 | A648T | COAD | 2 | K164 | E163K | SKCM | 2 | I548 | N549H | UCEC | 2 | N631,L633 | V632I | BRCA | 1 | H254 | P253S | COAD | 1 | I548 | L550I | COAD | 1 | D644 | I642T | COAD | 1 | R664 | R664W | COAD | 1 | R210 | R210Q | COAD | 1 | V495 | M497I | COAD | 1 | D644 | I642V | COAD | 1 | I217 | E219G | COAD | 1 | D644 | I642M | COAD | 1 | G646 | A648D | COAD | 1 | H213 | Q212K | GBM | 1 | T174 | T174N | HNSC | 1 | K164 | R165L | LUAD | 1 | H254 | P253L | LUAD | 1 | D626 | R625Q | SKCM | 1 | R251 | R251Q | SKCM | 1 | V709 | P708S | SKCM | 1 | R210 | R210Q | SKCM | 1 | G570 | G570R | STAD | 1 | K164 | R165W | UCEC | 1 | M538 | M537I | UCEC | 1 | R664 | R664W | UCEC | 1 |
| cf) Cancer type abbreviation. BLCA: Bladder urothelial carcinoma, BRCA: Breast invasive carcinoma, CESC: Cervical squamous cell carcinoma and endocervical adenocarcinoma, COAD: Colon adenocarcinoma, GBM: Glioblastoma multiforme, LGG: Brain lower grade glioma, HNSC: Head and neck squamous cell carcinoma, KICH: Kidney chromophobe, KIRC: Kidney renal clear cell carcinoma, KIRP: Kidney renal papillary cell carcinoma, LAML: Acute myeloid leukemia, LUAD: Lung adenocarcinoma, LUSC: Lung squamous cell carcinoma, OV: Ovarian serous cystadenocarcinoma, PAAD: Pancreatic adenocarcinoma, PRAD: Prostate adenocarcinoma, SKCM: Skin cutaneous melanoma, STAD: Stomach adenocarcinoma, THCA: Thyroid carcinoma, UCEC: Uterine corpus endometrial carcinoma. |
 Clinical information for FGFR2 from My Cancer Genome. Clinical information for FGFR2 from My Cancer Genome. |
| The fibroblast growth factor receptor 2 gene (FGFR2) encodes one member of the FGFR tyrosine kinase (TK) family, which includes four kinases: FGFR1, 2, 3, and 4. FGFR TKs play crucial roles in development and have been shown in cancers to be deregulated by amplification, point mutation, or translocation (Turner and Grose 2010). Amplification or activation of FGFR2 has been reported in breast cancer and gastric cancer (Jain and Turner 2012). FGFR2 mutations have been observed in endometrial cancer and breast cancer (Dutt et al. 2008; Jain and Turner 2012;Reintjes et al. 2013).Turner, N. 2015. FGFR2. My Cancer Genomehttps://www.mycancergenome.org/content/gene/fgfr2/ (Updated December 2015) |
| Top |
| Protein structure related information for FGFR2 |
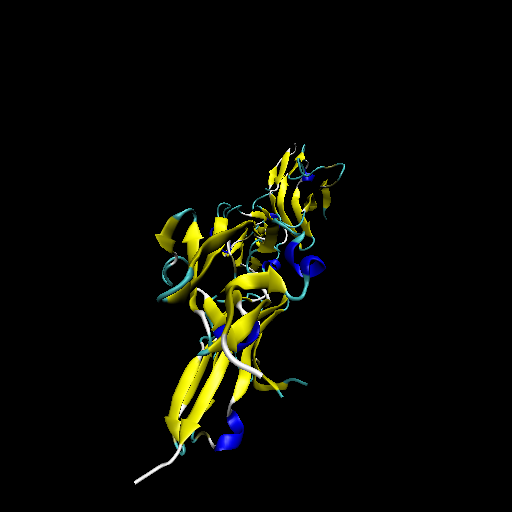
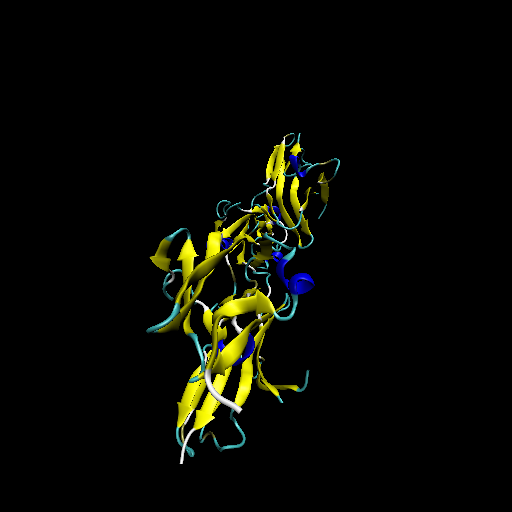
 Protein structure of wild type (WT) and mutant type (MT) of FGFR2 Protein structure of wild type (WT) and mutant type (MT) of FGFR2 |
| Wild type FGFR2 |
 |
| Mutant type FGFR2 |
 |
 Free energy of binding of drugs to wild type and mutant tpye of FGFR2 Free energy of binding of drugs to wild type and mutant tpye of FGFR2 |
| Gene symbol | Drug name | Free energy of binding (kcal/mol) of wild type | Free energy of binding (kcal/mol) of mutant type | FGFR2 | Ponatinib | -9.5 | -9.4 | FGFR2 | Regorafenib | -8.1 | -8.1 |
 Relative protein structure stability change (ΔΔE) using Mupro 1.1 Relative protein structure stability change (ΔΔE) using Mupro 1.1 Mupro score denotes assessment of the effect of mutations on thermodynamic stability. (ΔΔE<0: mutation decreases stability, ΔΔE>0: mutation increases stability) |
 : nsSNV at non-LBS : nsSNV at non-LBS : nsSNV at LBS : nsSNV at LBS |
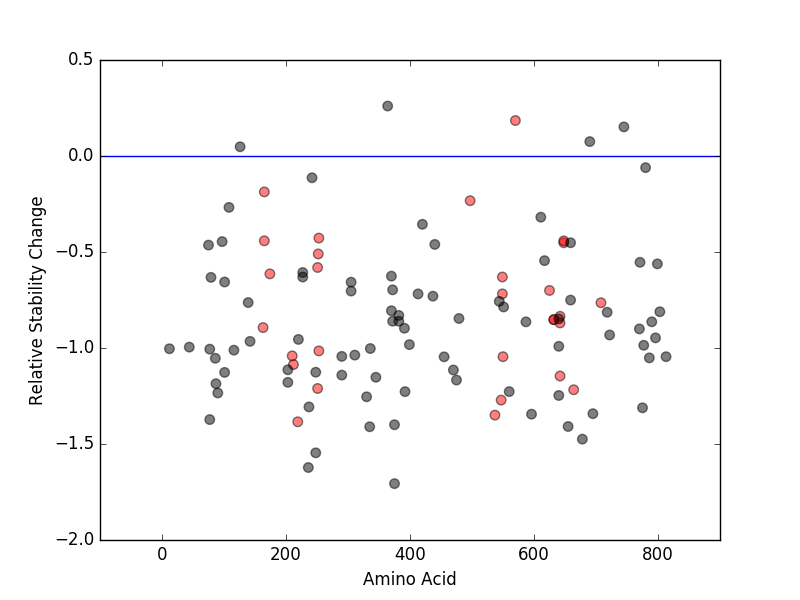 |
 nsSNVs sorted by the relative stability change of protein structure by each mutation nsSNVs sorted by the relative stability change of protein structure by each mutation Blue: mutations of positive stability change. and red : the most recurrent mutation for this gene. |
| LBS | AAchange of nsSNV | Relative stability change | G570 | G570R | 0.18475134 | I217 | E219G | -1.3843316 | M538 | M537I | -1.3495591 | I548 | I547T | -1.2713025 | R664 | R664W | -1.2174143 | R251 | R251Q | -1.2108418 | D644 | I642T | -1.1459779 | H213 | Q212K | -1.0862754 | I548 | L550I | -1.0452371 | R210 | R210Q | -1.0410815 | H254 | P253S | -1.015026 | K164 | E163K | -0.89351539 | D644 | I642V | -0.86954299 | N631 | V632I | -0.85241959 | L633 | V632I | -0.85241959 | D644 | I642M | -0.83491218 | V709 | P708S | -0.76431207 | I548 | N549H | -0.71820678 | D626 | R625Q | -0.7002287 | I548 | N549K | -0.63012981 | T174 | T174N | -0.61386992 | R251 | R251L | -0.58065701 | R251 | S252W | -0.51009314 | G646 | A648T | -0.45183309 | G646 | A648D | -0.44179852 | K164 | R165W | -0.44173214 | H254 | P253L | -0.42749447 | V495 | M497I | -0.23277115 | K164 | R165L | -0.18696067 |
| (MuPro1.1: Jianlin Cheng et al., Prediction of Protein Stability Changes for Single-Site Mutations Using Support Vector Machines, PROTEINS: Structure, Function, and Bioinformatics. 2006, 62:1125-1132) |
 Structure image for FGFR2 from PDB Structure image for FGFR2 from PDB |
| PDB ID | PDB title | PDB structure | 1GJO | THE FGFR2 TYROSINE KINASE DOMAIN | 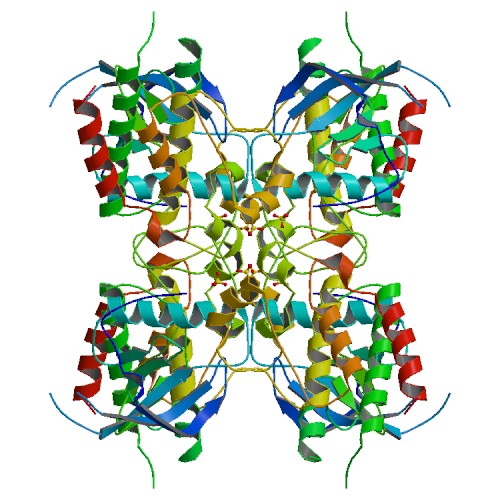 |
| Top |
| Differential gene expression and gene-gene network for FGFR2 |
 Differential gene expression between mutated and non-mutated LBS samples in all 16 major cancer types Differential gene expression between mutated and non-mutated LBS samples in all 16 major cancer types |
| FGFR2_COAD_DE |
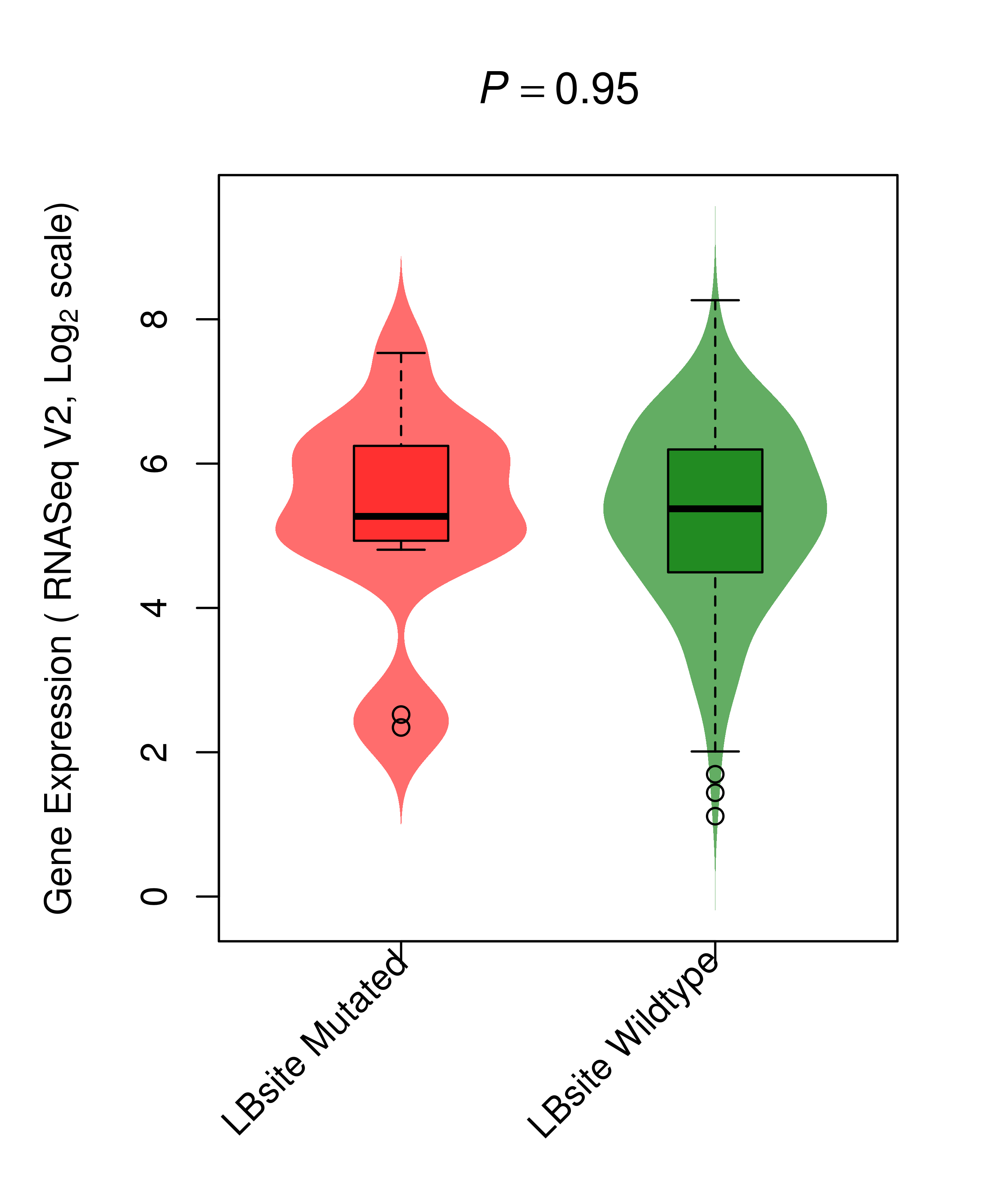 |
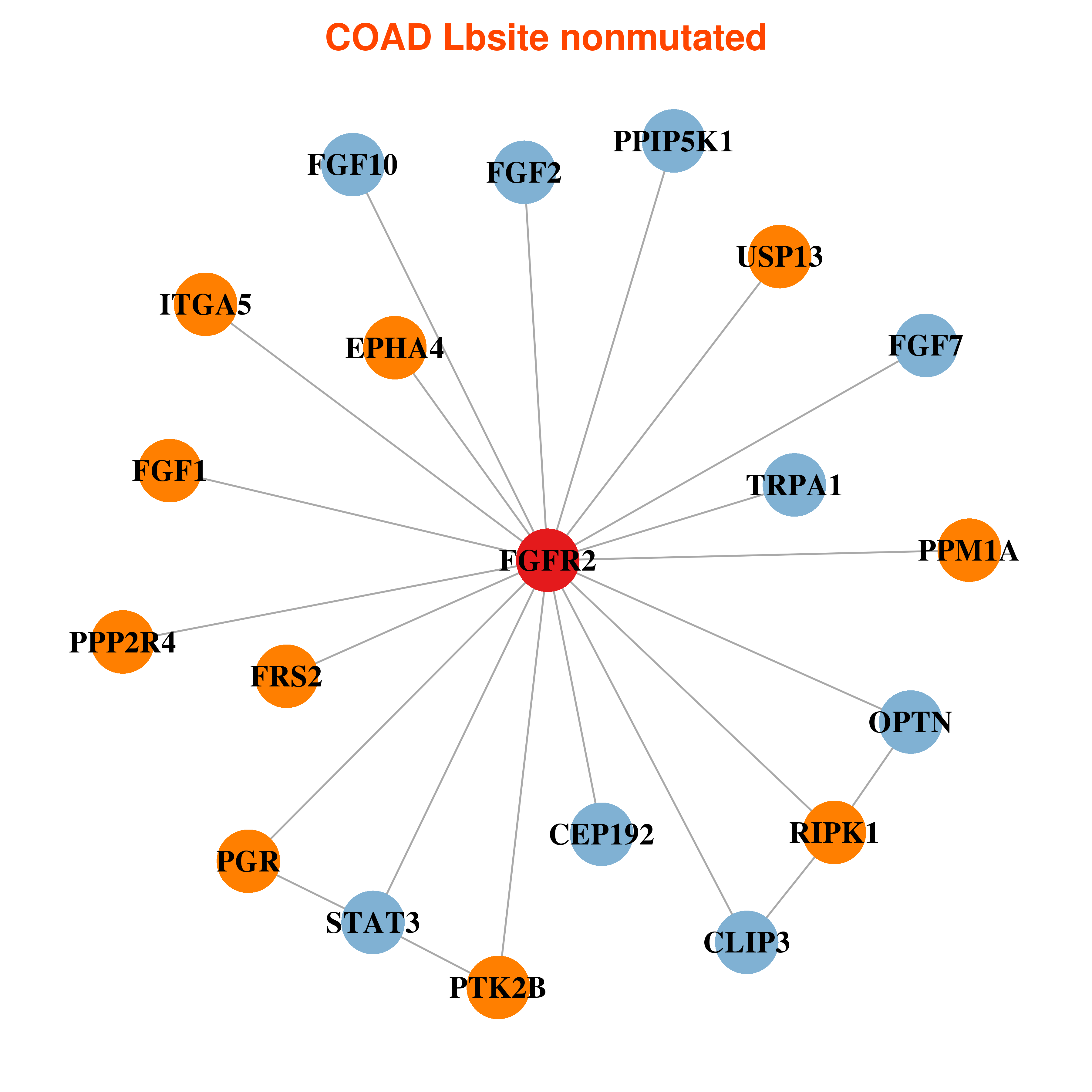
 Differential co-expressed gene network based on protein-protein interaction data (CePIN) Differential co-expressed gene network based on protein-protein interaction data (CePIN) |
| * In COAD | |
 |
|
| Top |
| Top |
| Phenotype information for FGFR2 |
 Gene level disease information (DisGeNet) Gene level disease information (DisGeNet) |
| Disease ID | Disease name | # PubMed | Association type |
| umls:C0010278 | Craniosynostoses | 78 | Biomarker, GeneticVariation |
| umls:C0001193 | apert syndrome | 51 | Biomarker, GeneticVariation |
| umls:C0010273 | Craniofacial Dysostosis | 41 | Biomarker, GeneticVariation |
| umls:C0220658 | PFEIFFER SYNDROME | 31 | Biomarker, GeneticVariation |
| umls:C1458155 | Breast Neoplasms | 20 | AlteredExpression, Biomarker, GeneticVariation |
| umls:C1510455 | Acrocephalosyndactylia | 12 | Biomarker, GeneticVariation |
| umls:C2936791 | antley-bixler syndrome | 7 | Biomarker, GeneticVariation |
| umls:C1852406 | Cutis Gyrata Syndrome of Beare And Stevenson | 6 | Biomarker, GeneticVariation |
| umls:C0376634 | Craniofacial Abnormalities | 5 | AlteredExpression, Biomarker |
| umls:C0795998 | Jackson-Weiss syndrome | 4 | Biomarker, GeneticVariation |
| umls:C0038356 | Stomach Neoplasms | 4 | AlteredExpression, Biomarker, PostTranslationalModification |
| umls:C0008925 | Cleft Palate | 4 | Biomarker |
| umls:C0175699 | Saethre-Chotzen Syndrome | 3 | Biomarker, GeneticVariation |
| umls:C0023890 | Liver Cirrhosis | 2 | AlteredExpression, Biomarker |
| umls:C3714756 | Intellectual Disability | 2 | Biomarker |
| umls:C0000772 | Abnormalities, Multiple | 1 | Biomarker |
| umls:C0003090 | Ankylosis | 1 | Biomarker |
| umls:C2350233 | Antley-Bixler Syndrome Phenotype | 1 | Biomarker |
| umls:C0006435 | Burns, Chemical | 1 | Biomarker |
| umls:C0008924 | Cleft Lip | 1 | Biomarker |
| umls:C0014170 | Endometrial Neoplasms | 1 | Biomarker |
| umls:C0018553 | Hamartoma Syndrome, Multiple | 1 | Biomarker |
| umls:C0206762 | Limb Deformities, Congenital | 1 | Biomarker |
| umls:C0024121 | Lung Neoplasms | 1 | Biomarker |
| umls:C0026613 | Motor Skills Disorders | 1 | Biomarker |
| umls:C1450010 | Plagiocephaly, Nonsynostotic | 1 | Biomarker |
| umls:C0037268 | Skin Abnormalities | 1 | Biomarker |
| umls:C0037274 | Skin Diseases | 1 | Biomarker |
| umls:C0080178 | Spinal Dysraphism | 1 | Biomarker |
| umls:C0040427 | Tooth Abnormalities | 1 | Biomarker |
 Mutation level pathogenic information (ClinVar annotation) Mutation level pathogenic information (ClinVar annotation) |
| Allele ID | AA change | Clinical significance | Origin | Phenotype IDs |
| 28311 | S252W | Pathogenic | Germline;somatic;unknown | GeneReviews:NBK1455 MedGen:C0001193 OMIM:101200 Orphanet:ORPHA87 SNOMED CT:205258009 MedGen:C0476089 OMIM:608089 SNOMED CT:254878006 |
| 28335 | A648T | Pathogenic | Germline | - |
| Top |
| Pharmacological information for FGFR2 |
 Gene expression profile of anticancer drug treated cell-lines (CCLE) Gene expression profile of anticancer drug treated cell-lines (CCLE)Heatmap showing the correlation between gene expression and drug response across all the cell-lines. We chose the top 20 among 138 drugs.We used Pearson's correlation coefficient. |
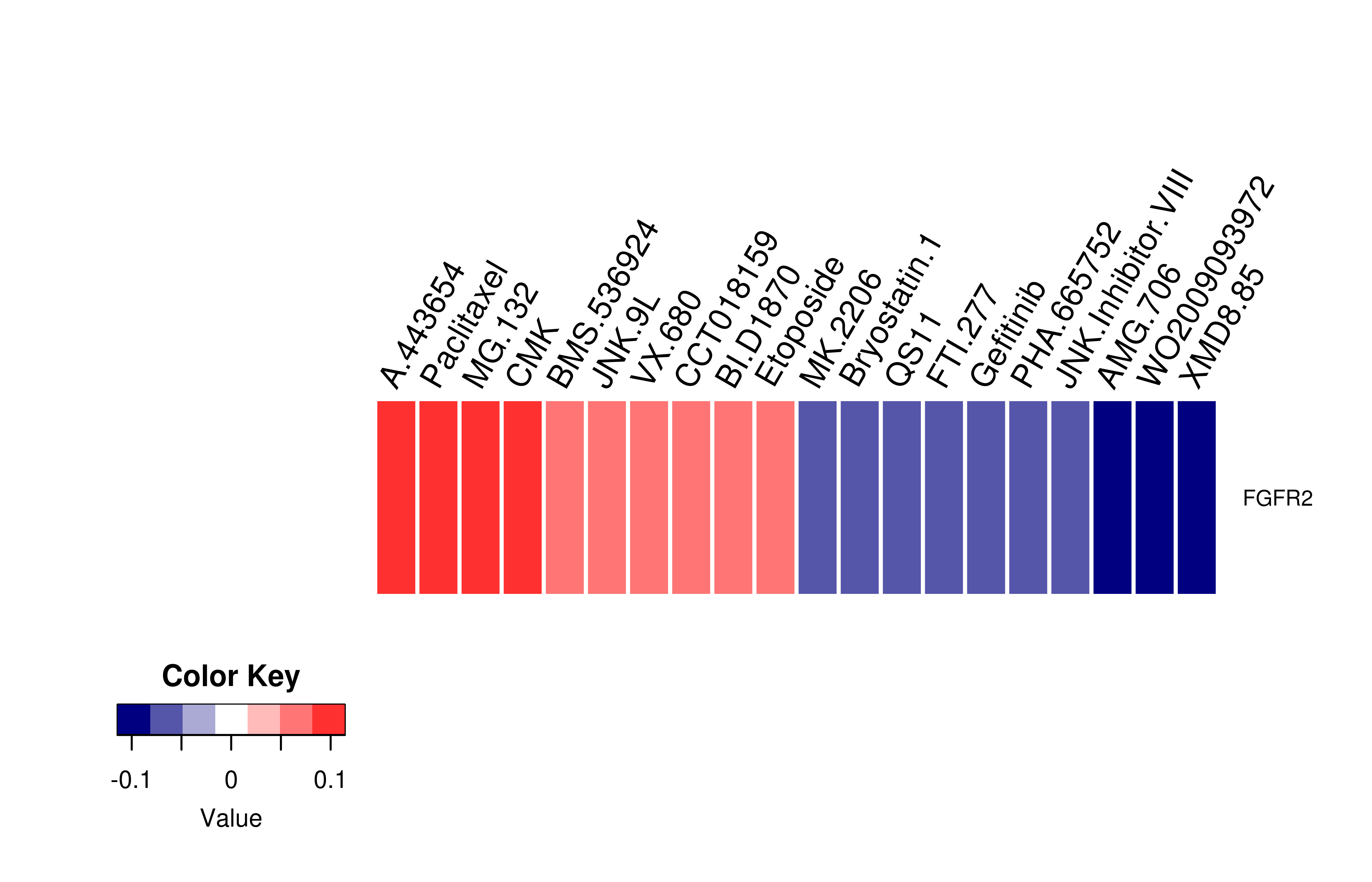 |
 Gene-centered drug-gene interaction network Gene-centered drug-gene interaction network |
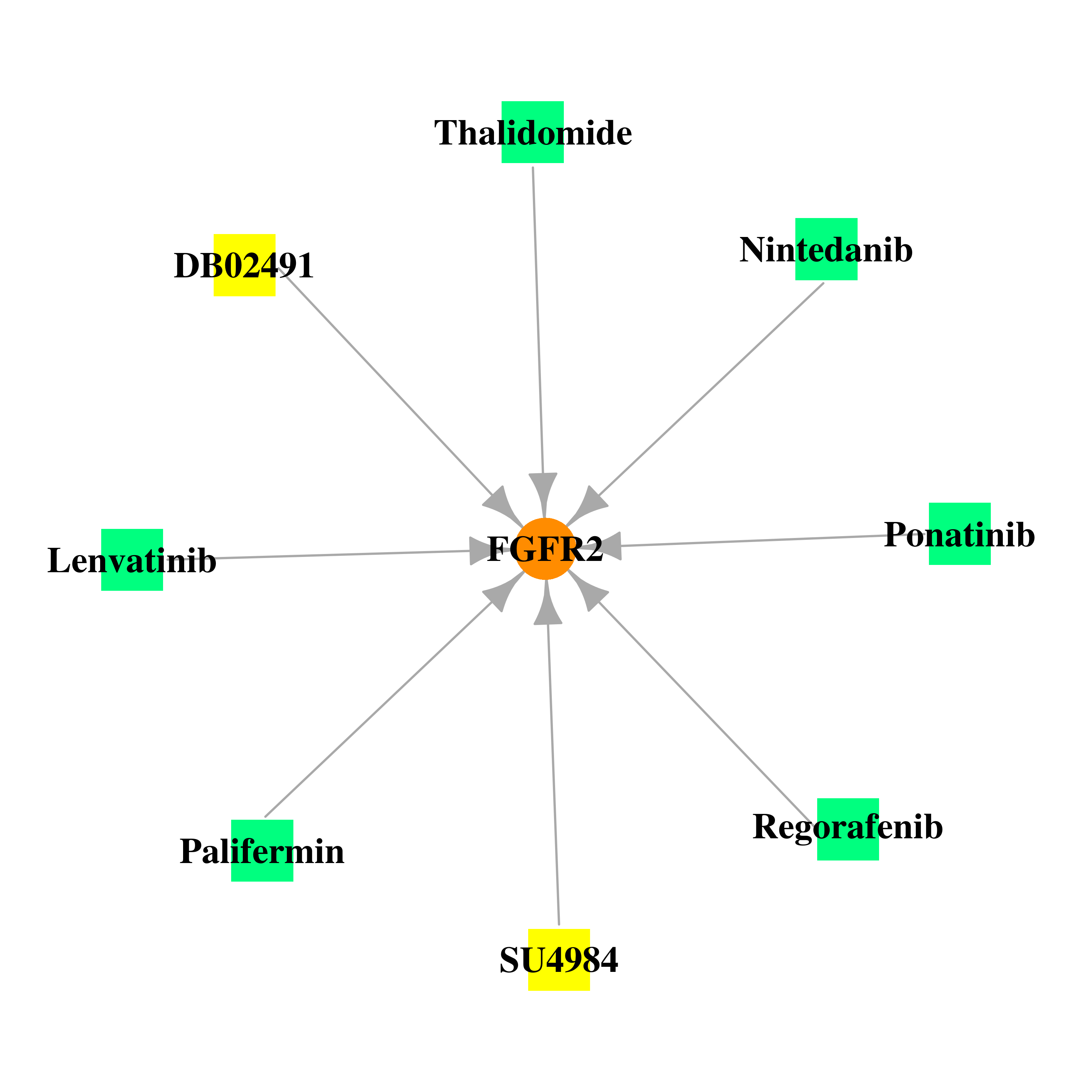 |
 Drug information targeting mutLBSgene (Approved drugs only) Drug information targeting mutLBSgene (Approved drugs only) |
| Drug status | DrugBank ID | Name | Type | Drug structure |
| Approved | DB00039 | Palifermin | Biotech |  |
| Approved|investigational|withdrawn | DB01041 | Thalidomide | Small molecule |  |
| Experimental | DB02058 | SU4984 | Small molecule |  |
| Experimental | DB02491 | 4-[4-(1-Amino-1-Methylethyl)Phenyl]-5-Chloro-N-[4-(2-Morpholin-4-Ylethyl)Phenyl]Pyrimidin-2-Amine | Small molecule | 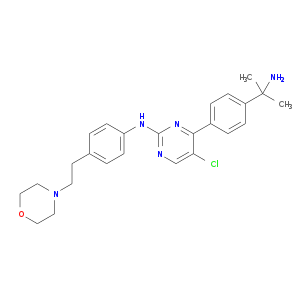 |
| Approved | DB08896 | Regorafenib | Small molecule | 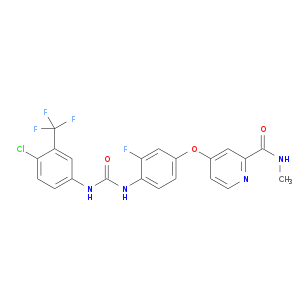 |
| Approved | DB08901 | Ponatinib | Small molecule | 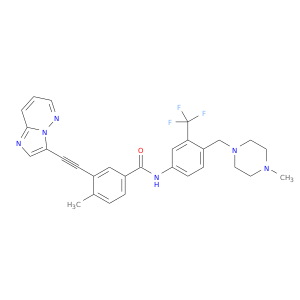 |
| Approved | DB09078 | Lenvatinib | Small molecule | 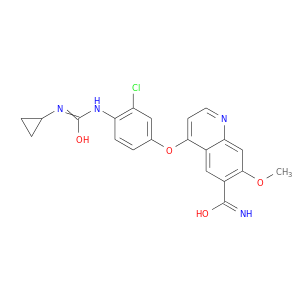 |
| Approved | DB09079 | Nintedanib | Small molecule |  |
 Gene-centered ligand-gene interaction network Gene-centered ligand-gene interaction network |
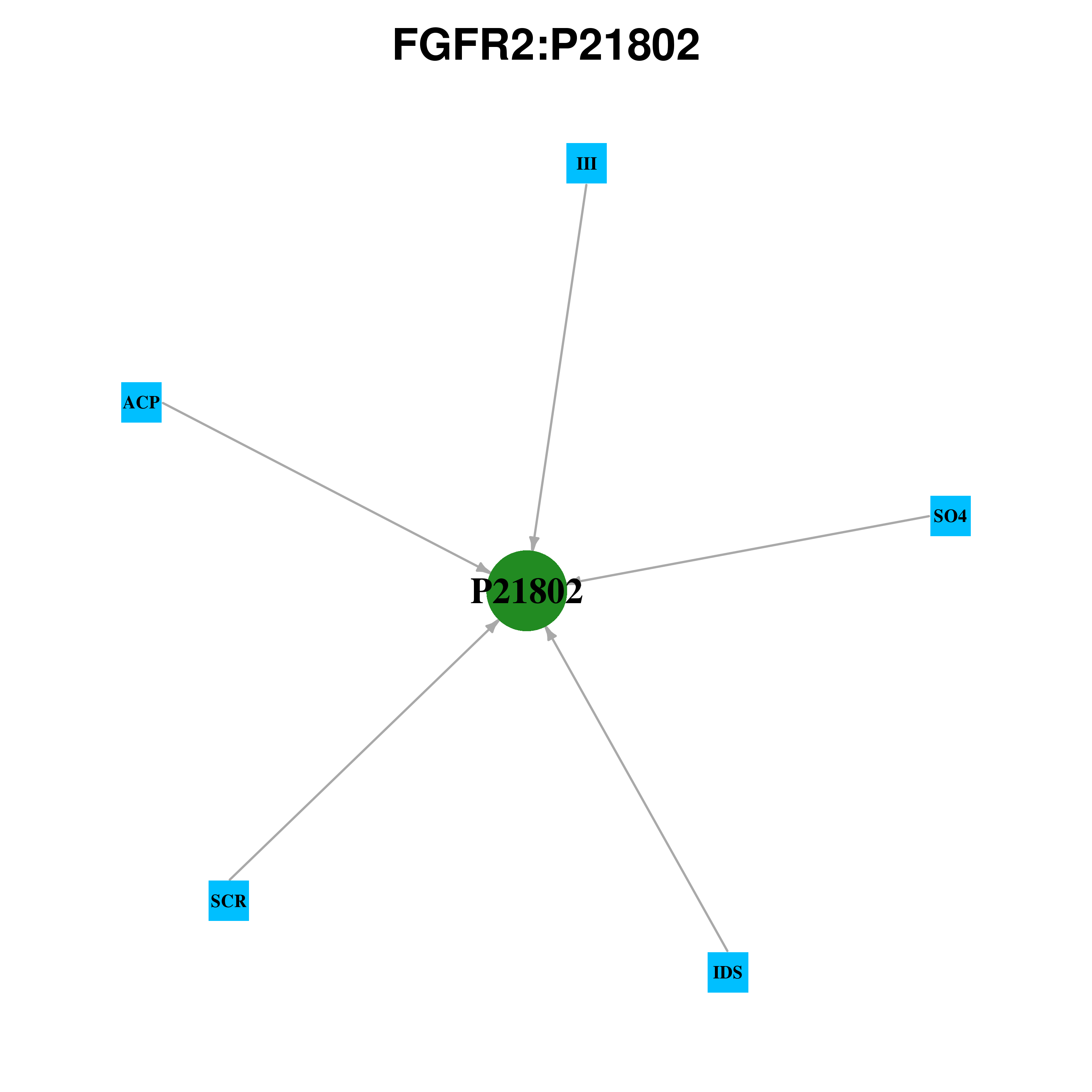 |
 Ligands binding to mutated ligand binding site of FGFR2 go to BioLip Ligands binding to mutated ligand binding site of FGFR2 go to BioLip |
| Ligand ID | Ligand short name | Ligand long name | PDB ID | PDB name | mutLBS | III | Peptide ligand (GLU,GLU,TYR,LEU) | 2pvf | A | D626 R664 V709 | MG | MAGNESIUM(2+) | 4j96 | B | D644 | MG | MAGNESIUM(2+) | 4j99 | D | D644 G646 | SO4 | SULFATE | 1djs | A | H254 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j98 | B | I548 G570 D626 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j96 | B | I548 L633 D644 | SGN | N,O6-DISULFO-GLUCOSAMINE | 1e0o | B | K164 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pvy | A | L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pvy | C | L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pvy | D | L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pzr | B | L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2q0b | B | L633 | M33 | 5'-O-[(S)-HYDROXY{[(S)-HYDROXY(METHYL)PHOSPHORYL]OXY}PHOSPHORYL]ADENOSINE | 3b2t | A | L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j97 | A | L633 | AA2 | 4-[4-(1-AMINO-1-METHYLETHYL)PHENYL]-5-CHLORO-N-[4-(2-MORPHOLIN-4-YLETHYL)PHENYL]PYRIMIDIN-2-AMINE | 1oec | A | L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j97 | D | L633 D644 | MG | MAGNESIUM(2+) | 2pvf | A | N631 D644 | MG | MAGNESIUM(2+) | 2py3 | A | N631 D644 | MG | MAGNESIUM(2+) | 2py3 | B | N631 D644 | MG | MAGNESIUM(2+) | 2pz5 | A | N631 D644 | MG | MAGNESIUM(2+) | 2pz5 | B | N631 D644 | MG | MAGNESIUM(2+) | 2pzp | A | N631 D644 | MG | MAGNESIUM(2+) | 2pzp | B | N631 D644 | MG | MAGNESIUM(2+) | 4j99 | C | N631 D644 | MG | MAGNESIUM(2+) | 4j99 | D | N631 D644 | SCR | SUCROSE OCTASULFATE | 3cu1 | C | R210 H213 | SCR | SUCROSE OCTASULFATE | 3cu1 | A | R210 H213 I217 | SO4 | SULFATE | 1djs | A | R210 I217 | SO4 | SULFATE | 1e0o | D | R251 H254 | IDS | 2-O-SULFO-ALPHA-L-IDOPYRANURONIC ACID | 1e0o | B | T174 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j97 | B | V495 G570 N631 L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j99 | A | V495 I548 D626 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j99 | B | V495 I548 D626 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2q0b | A | V495 I548 L633 | M33 | 5'-O-[(S)-HYDROXY{[(S)-HYDROXY(METHYL)PHOSPHORYL]OXY}PHOSPHORYL]ADENOSINE | 3b2t | B | V495 I548 N631 L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j99 | D | V495 I548 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j99 | C | V495 I548 N631 L633 D644 G646 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pvy | B | V495 L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pzr | A | V495 L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pwl | A | V495 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pwl | B | V495 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j95 | C | V495 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j95 | D | V495 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j97 | C | V495 L633 D644 | 3RH | (6S)-6-PHENYL-5,6-DIHYDROBENZO[H]QUINAZOLIN-2-AMINE | 3ri1 | A | V495 M538 I548 L633 | 3RH | (6S)-6-PHENYL-5,6-DIHYDROBENZO[H]QUINAZOLIN-2-AMINE | 3ri1 | B | V495 M538 L633 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pvf | A | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2py3 | A | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2py3 | B | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pz5 | A | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pz5 | B | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pzp | A | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 2pzp | B | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j95 | A | V495 N631 L633 D644 | ACP | ADENOSINE 5'-[BETA,GAMMA-METHYLENE]TRIPHOSPHATE | 4j95 | B | V495 N631 L633 D644 |
| Top |
| Conservation information for LBS of FGFR2 |
 Multiple alignments for P21802 in multiple species Multiple alignments for P21802 in multiple species |
| LBS | AA sequence | # species | Species | A355 | ISFHSAWLTV- | 2 | Homo sapiens, Mus musculus | A355 | RSNSSVYLRVV | 1 | Drosophila melanogaster | A355 | ISFHTAWLTV- | 1 | Gallus gallus | A515 | EAVTVAVKMLK | 3 | Homo sapiens, Gallus gallus, Mus musculus | A515 | AETIVAVKMVK | 1 | Drosophila melanogaster | A567 | VIVEYASKGNL | 3 | Homo sapiens, Gallus gallus, Mus musculus | A567 | VIVEYAPHGNL | 1 | Drosophila melanogaster | D626 | KCIHRDLAARN | 3 | Homo sapiens, Gallus gallus, Mus musculus | D626 | RCIHRDLAARN | 1 | Drosophila melanogaster | D644 | VMKIADFGLAR | 4 | Homo sapiens, Drosophila melanogaster, Gallus gallus, Mus musculus | E339 | FEDAGEYTCLA | 3 | Homo sapiens, Gallus gallus, Mus musculus | E339 | FDQEGWYTCLA | 1 | Drosophila melanogaster | E489 | GKPLGEGCFGQ | 3 | Homo sapiens, Gallus gallus, Mus musculus | E489 | GSILGEGAFGR | 1 | Drosophila melanogaster | E534 | SDLVSEMEMMK | 3 | Homo sapiens, Gallus gallus, Mus musculus | E534 | ASLVREMEVMK | 1 | Drosophila melanogaster | E565 | LYVIVEYASKG | 3 | Homo sapiens, Gallus gallus, Mus musculus | E565 | LWVIVEYAPHG | 1 | Drosophila melanogaster | F492 | LGEGCFGQVVM | 3 | Homo sapiens, Gallus gallus, Mus musculus | F492 | LGEGAFGRVVM | 1 | Drosophila melanogaster | G488 | LGKPLGEGCFG | 3 | Homo sapiens, Gallus gallus, Mus musculus | G488 | LGSILGEGAFG | 1 | Drosophila melanogaster | G490 | KPLGEGCFGQV | 3 | Homo sapiens, Gallus gallus, Mus musculus | G490 | SILGEGAFGRV | 1 | Drosophila melanogaster | G493 | GEGCFGQVVMA | 3 | Homo sapiens, Gallus gallus, Mus musculus | G493 | GEGAFGRVVMA | 1 | Drosophila melanogaster | G570 | EYASKGNLREY | 3 | Homo sapiens, Gallus gallus, Mus musculus | G570 | EYAPHGNLKDF | 1 | Drosophila melanogaster | G646 | KIADFGLARDI | 4 | Homo sapiens, Drosophila melanogaster, Gallus gallus, Mus musculus | H167 | MEKRLHAVPAA | 2 | Homo sapiens, Gallus gallus | H167 | ELKRLQHSLSG | 1 | Drosophila melanogaster | H167 | MEKRLHACPAA | 1 | Mus musculus | H213 | KVRNQHWSLIM | 3 | Homo sapiens, Gallus gallus, Mus musculus | H213 | VQKN--WTLRF | 1 | Drosophila melanogaster | H254 | VERSPHRPILQ | 3 | Homo sapiens, Gallus gallus, Mus musculus | H254 | NDRTRSAPIIV | 1 | Drosophila melanogaster | I217 | QHWSLIMESVV | 3 | Homo sapiens, Gallus gallus, Mus musculus | I217 | --WTLRFVEAT | 1 | Drosophila melanogaster | I348 | LAGNSIGISFH | 3 | Homo sapiens, Gallus gallus, Mus musculus | I348 | LASSGLGRSNS | 1 | Drosophila melanogaster | I548 | KHKNIINLLGA | 3 | Homo sapiens, Gallus gallus, Mus musculus | I548 | KHINIINLLGC | 1 | Drosophila melanogaster | K161 | WTNTEKMEKRL | 2 | Homo sapiens, Mus musculus | K161 | FR---KELKRL | 1 | Drosophila melanogaster | K161 | WTHTDKMEKRL | 1 | Gallus gallus | K164 | TEKMEKRLHAV | 1 | Homo sapiens | K164 | --KELKRLQHS | 1 | Drosophila melanogaster | K164 | TDKMEKRLHAV | 1 | Gallus gallus | K164 | TEKMEKRLHAC | 1 | Mus musculus | K176 | AANTVKFRCPA | 3 | Homo sapiens, Gallus gallus, Mus musculus | K176 | SGNTVNLACPV | 1 | Drosophila melanogaster | K517 | VTVAVKMLKDD | 3 | Homo sapiens, Gallus gallus, Mus musculus | K517 | TIVAVKMVKEE | 1 | Drosophila melanogaster | L487 | TLGKPLGEGCF | 3 | Homo sapiens, Gallus gallus, Mus musculus | L487 | SLGSILGEGAF | 1 | Drosophila melanogaster | L633 | AARNVLVTENN | 3 | Homo sapiens, Gallus gallus, Mus musculus | L633 | AARNVLVSDGY | 1 | Drosophila melanogaster | L665 | TTNGRLPVKWM | 3 | Homo sapiens, Gallus gallus, Mus musculus | L665 | NTNGRLPIKWM | 1 | Drosophila melanogaster | M538 | SEMEMMKMIGK | 3 | Homo sapiens, Gallus gallus, Mus musculus | M538 | REMEVMKMIGK | 1 | Drosophila melanogaster | N571 | YASKGNLREYL | 3 | Homo sapiens, Gallus gallus, Mus musculus | N571 | YAPHGNLKDFL | 1 | Drosophila melanogaster | N631 | DLAARNVLVTE | 3 | Homo sapiens, Gallus gallus, Mus musculus | N631 | DLAARNVLVSD | 1 | Drosophila melanogaster | P666 | TNGRLPVKWMA | 3 | Homo sapiens, Gallus gallus, Mus musculus | P666 | TNGRLPIKWMA | 1 | Drosophila melanogaster | Q494 | EGCFGQVVMAE | 3 | Homo sapiens, Gallus gallus, Mus musculus | Q494 | EGAFGRVVMAE | 1 | Drosophila melanogaster | R178 | NTVKFRCPAGG | 2 | Homo sapiens, Mus musculus | R178 | NTVNLACPVYG | 1 | Drosophila melanogaster | R178 | NTVKFRCPAMG | 1 | Gallus gallus | R210 | GGYKVRNQHWS | 3 | Homo sapiens, Gallus gallus, Mus musculus | R210 | GVYVQKN--WT | 1 | Drosophila melanogaster | R251 | LDVVERSPHRP | 3 | Homo sapiens, Gallus gallus, Mus musculus | R251 | VQINDRTRSAP | 1 | Drosophila melanogaster | R255 | ERSPHRPILQA | 3 | Homo sapiens, Gallus gallus, Mus musculus | R255 | DRTRSAPIIV- | 1 | Drosophila melanogaster | R630 | RDLAARNVLVT | 3 | Homo sapiens, Gallus gallus, Mus musculus | R630 | RDLAARNVLVS | 1 | Drosophila melanogaster | R664 | KTTNGRLPVKW | 3 | Homo sapiens, Gallus gallus, Mus musculus | R664 | KNTNGRLPIKW | 1 | Drosophila melanogaster | S215 | RNQHWSLIMES | 3 | Homo sapiens, Gallus gallus, Mus musculus | S215 | KN--WTLRFVE | 1 | Drosophila melanogaster | S354 | GISFHSAWLTV | 2 | Homo sapiens, Mus musculus | S354 | GRSNSSVYLRV | 1 | Drosophila melanogaster | S354 | GISFHTAWLTV | 1 | Gallus gallus | T174 | VPAANTVKFRC | 2 | Homo sapiens, Gallus gallus | T174 | SLSGNTVNLAC | 1 | Drosophila melanogaster | T174 | CPAANTVKFRC | 1 | Mus musculus | V175 | PAANTVKFRCP | 3 | Homo sapiens, Gallus gallus, Mus musculus | V175 | LSGNTVNLACP | 1 | Drosophila melanogaster | V495 | GCFGQVVMAEA | 3 | Homo sapiens, Gallus gallus, Mus musculus | V495 | GAFGRVVMAEA | 1 | Drosophila melanogaster | V564 | PLYVIVEYASK | 3 | Homo sapiens, Gallus gallus, Mus musculus | V564 | PLWVIVEYAPH | 1 | Drosophila melanogaster | V667 | NGRLPVKWMAP | 3 | Homo sapiens, Gallus gallus, Mus musculus | V667 | NGRLPIKWMAP | 1 | Drosophila melanogaster | V709 | PGI-PVEELFK | 3 | Homo sapiens, Gallus gallus, Mus musculus | V709 | PHILSAEELYS | 1 | Drosophila melanogaster | W356 | SFHSAWLTV-- | 2 | Homo sapiens, Mus musculus | W356 | SNSSVYLRVVS | 1 | Drosophila melanogaster | W356 | SFHTAWLTV-- | 1 | Gallus gallus | W669 | RLPVKWMAPEA | 3 | Homo sapiens, Gallus gallus, Mus musculus | W669 | RLPIKWMAPES | 1 | Drosophila melanogaster | Y566 | YVIVEYASKGN | 3 | Homo sapiens, Gallus gallus, Mus musculus | Y566 | WVIVEYAPHGN | 1 | Drosophila melanogaster |
 |
Copyright © 2016-Present - The University of Texas Health Science Center at Houston |